
Eva Maria Novoa
359 posts


Eva Maria Novoa
@EvaMariaNovoa
Group Leader -Epitranscriptomics & RNA Dynamics @CRGenomica. Biochemist and bioinformatician. #RNAmods #nanopore #ribosome. Mom of 3. Piano, beach vball, hiking


Our new paper on reassessing the heritability of human lifespan is out in @ScienceMagazine! 🧬 For decades, the consensus has been that genetics explains just 20–25% of lifespan differences. We found that after accounting for extrinsic mortality, that number jumps to ~50%. A 🧵







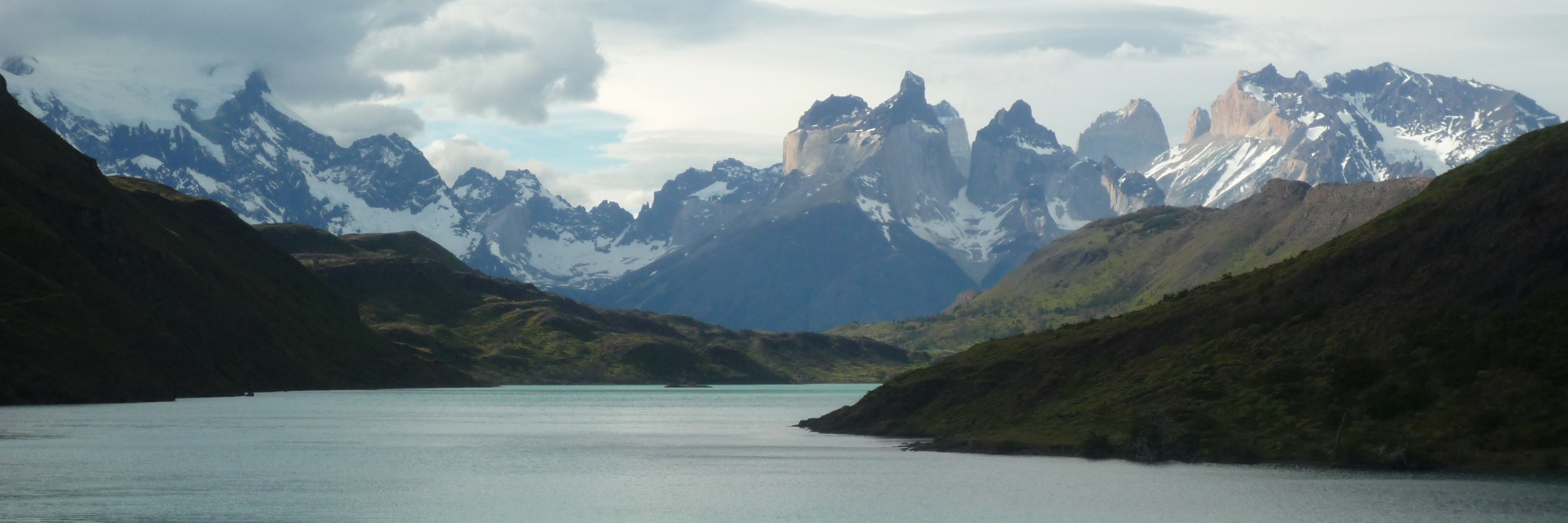
17. University of Washington warns of potential layoffs kuow.org/stories/uw-pre…



📢Nous guanyadors dels Premis Nacionals de Recerca de Catalunya❗ 🎖Núria Sebastián Gallés, Premi Nacional de Recerca 2024 (@UPFBarcelona) 🎖@EvaMariaNovoa, PNR al Talent Jove 2024 ex aequo (@CRGenomica) 🎖@ros_xavi, PNR al Talent Jove 2024 ex aequo (@UniBarcelona) L'enhorabona!







VERY excited to share SeqTagger with the world! If you want to learn more about our super-fast and accurate demultiplexing tool for direct RNA-sequencing (DRS), check out the thread below. bit.ly/3ClVYxG (1/15) 🧵



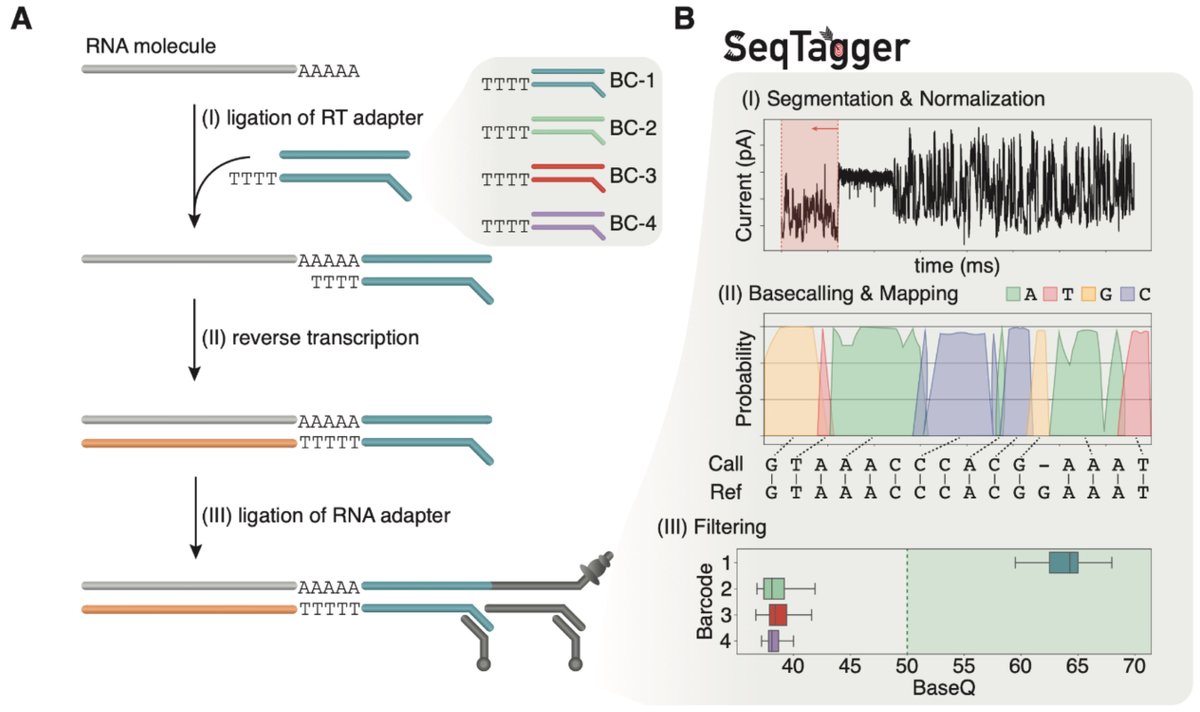

Dynamic rRNA modifications as a source of ribosome heterogeneity dlvr.it/TG1gDF


