

Jonathan Weissman's Lab
148 posts
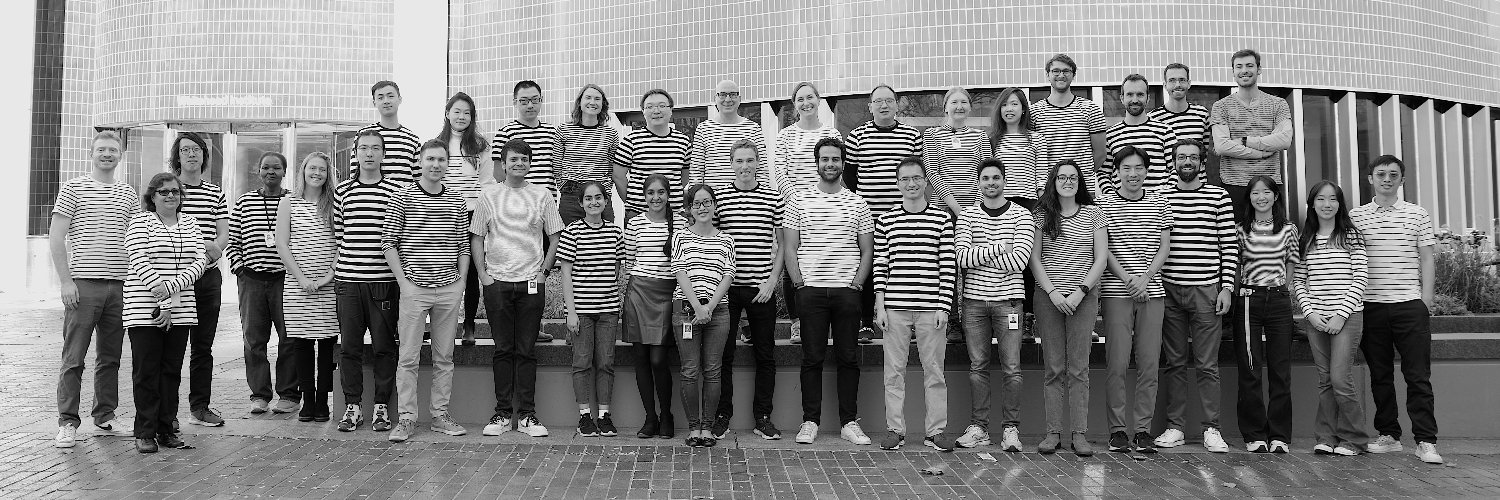


@JswLab
Jonathan S. Weissman's lab at the @WhiteheadInst/@MIT. Account run by lab members.



New preprint on technologies to scale up CRISPR screens. We use them to map 665,856 pairwise genetic perturbations and outline a path to comprehensive interaction mapping in human cells. We also introduce an approach for cloning lentiviral libraries with billions of elements.

Excited to share our work with @DMSabatini & @JswLab. How do lysosomes change with age? We present a metabolic atlas of lysosomal aging, and reveal a lysosomal “aging clock” of metabolites linked to lysosomal storage disorders. Grateful to all co-authors! biorxiv.org/cgi/content/sh…

1/ Can we find more precise and even safer ways to rejuvenate vision? As a step towards this goal, I am excited to share this preprint of my 1st postdoc work from @JswLab with amazing collaborators, esp. @Bruce_Ksander! Using functional genomics, we discovered GSTA4 as a key OSK effector for oxidative resilience—overexpressing it alone rejuvenates RPE & restores vision. A breakdown👇biorxiv.org/content/10.110…

I'm thrilled to share our work from @JswLab (sciencedirect.com/science/articl…). We developed LOCL-TL, an optogenetic approach for monitoring localized translation in mammalian cells. LOCL-TL revealed two mechanistically distinct strategies for mitochondrially localized translation.

1/14 On behalf of the amazing team in @JswLab, we’re excited to share PEtracer (biorxiv.org/content/10.110…) a prime editing-based evolving lineage recorder compatible with both scRNA-seq and high-resolution imaging readouts in intact tissue. By applying PEtracer in a syngeneic mouse model of lung metastasis, we dissect how cell-intrinsic and environmental factors coordinately shape tumor evolution in vivo.


1/12 Excited to share our team's latest work and the first @xaira_thera preprint! Here, we introduce FiCS Perturb-seq, an industrialized platform for generating scaleable, high-quality perturbation data. 📄 Read the preprint: biorxiv.org/content/10.110…

1/14 On behalf of the amazing team in @JswLab, we’re excited to share PEtracer (biorxiv.org/content/10.110…) a prime editing-based evolving lineage recorder compatible with both scRNA-seq and high-resolution imaging readouts in intact tissue. By applying PEtracer in a syngeneic mouse model of lung metastasis, we dissect how cell-intrinsic and environmental factors coordinately shape tumor evolution in vivo.


Congrats @NadigAjay on TRADE out now in @NatureGenet: nature.com/articles/s4158… These statistical metrics enable more meaningful comparisons in Perturb-seq atlases. Also, now find the HepG2 and Jurkat Perturb-seq datasets on GEO GSE264667!

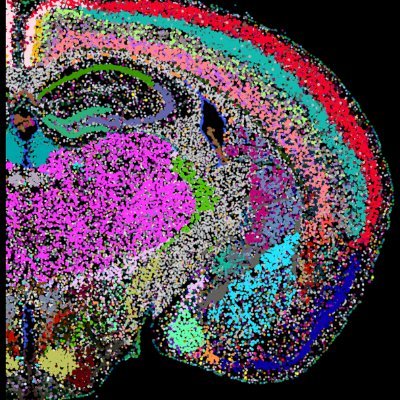
Thrilled to share Perturb-DBiT — spatial unbiased in vivo perturb-seq with genome-scale CRISPR libraries! Hope you enjoy reading it over the Thanksgiving week 🥰 Kudos to @AlevBaysoy, Xiaolong, Feifei, and Paul. & collaboration with @sidichen, Hongbo Chi. biorxiv.org/content/10.110…

Check out our preprint on Perturb-multi (doi.org/10.1101/2024.1…), a multi-year collaborative effort that enabled multi-modal in vivo genotype-phenotype mapping by pooled genetic perturbations and high-dimensionality phenotype readout by imaging and sequencing.



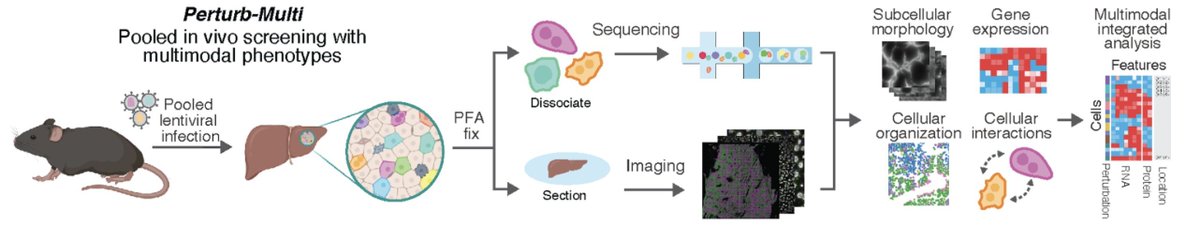
In collaboration with Reuben Saunders, @JswLab, and Xiaowei Zhuang, we are very excited to release Perturb-Multi: a platform for pooled multimodal genetic screens in intact mammalian tissue. Check it out! biorxiv.org/content/10.110…


In collaboration with Reuben Saunders, @JswLab, and Xiaowei Zhuang, we are very excited to release Perturb-Multi: a platform for pooled multimodal genetic screens in intact mammalian tissue. Check it out! biorxiv.org/content/10.110…