Michael Pearce รีทวีตแล้ว
Michael Pearce
88 posts


Michael Pearce
@_MichaelPearce
Interpretability @ Goodfire | Physics | Evolution | Ecology
เข้าร่วม Eylül 2015
625 กำลังติดตาม175 ผู้ติดตาม
Michael Pearce รีทวีตแล้ว
Michael Pearce รีทวีตแล้ว

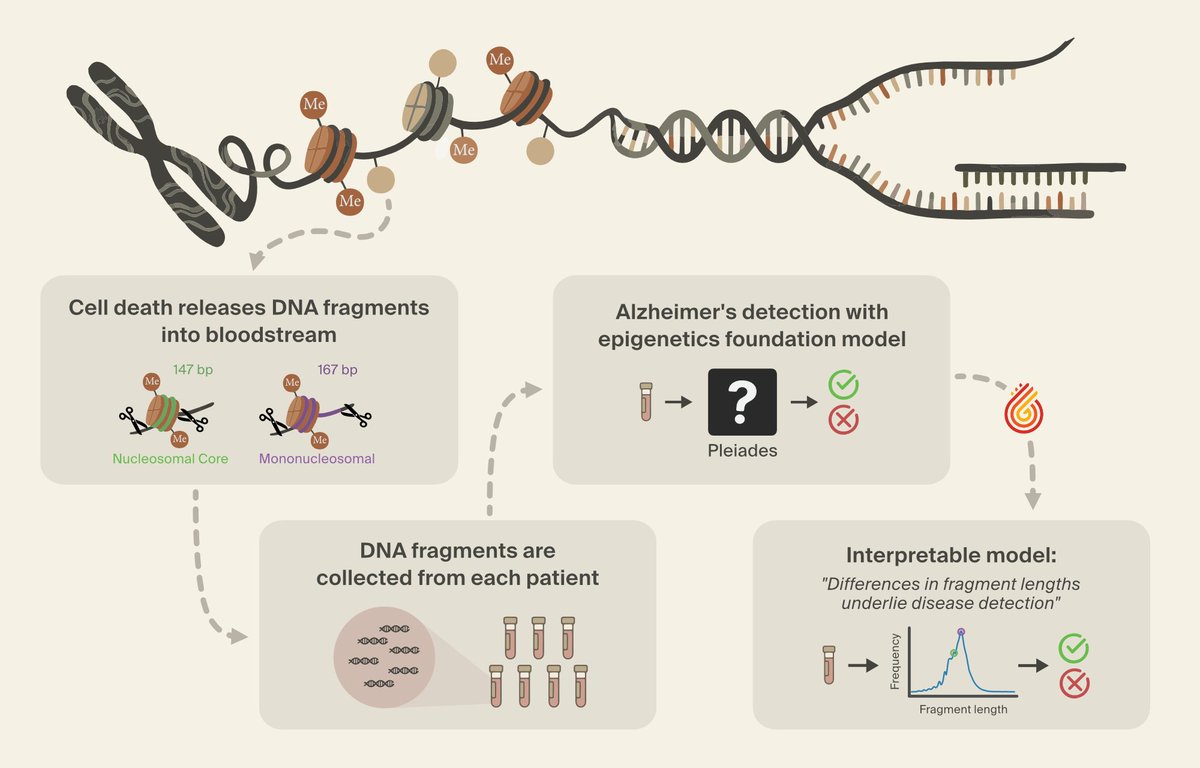
We've identified a novel class of biomarkers for Alzheimer's detection - using interpretability - with @PrimaMente.
How we did it, and how interpretability can power scientific discovery in the age of digital biology: (1/6)
English
Michael Pearce รีทวีตแล้ว

Why use LLM-as-a-judge when you can get the same performance for 15–500x cheaper?
Our new research with @RakutenGroup on PII detection finds that SAE probes:
- transfer from synthetic to real data better than normal probes
- match GPT-5 Mini performance at 1/15 the cost
(1/6)

English
Michael Pearce รีทวีตแล้ว
Michael Pearce รีทวีตแล้ว
Michael Pearce รีทวีตแล้ว

We're excited to announce a collaboration with @MayoClinic!
We're working to improve personalized patient outcomes by extracting richer, more reliable signals from genomic & digital pathology models.
That could mean novel biomarkers, personalized diagnostics, & more.

English
Michael Pearce รีทวีตแล้ว

Does making an SAE bigger let you explain more of your model's features?
New research from @ericjmichaud_ models SAE scaling dynamics, and explores whether SAEs will pack increasingly many latents onto a few multidimensional features, rather than learning more features.

English

@ATinyGreenCell @pdhsu @GoodfireAI Good questions, we used a set of 2400 prokaryote genomes with complete assemblies that are representative genomes in both the GTDB and NCBI databases. No viruses or metagenomic assemblies!
English

@pdhsu @GoodfireAI Im assuming all these data were carefully curated and checked for contamination in the RSA data? Was this refseq only? Was this metagenomics as well? Do you consider MAGs as real? Does this account for virome?
Connections based on all available sequences obv relies on data quali
English

Finding the tree of life within Evo 2 - amazing work by our collaborators @GoodfireAI
Goodfire@GoodfireAI
Arc Institute trained their foundation model Evo 2 on DNA from all domains of life. What has it learned about the natural world? Our new research finds that it represents the tree of life, spanning thousands of species, as a curved manifold in its neuronal activations. (1/8)
English

@PhilEmmanuele @GoodfireAI The flow of gathering activations from random genomic regions and averaging is described in the post. We’re happy to share the resulting data—the species-averaged embeddings and the phylogenetic distances between species—for you to play with. Will let you know when it's available
English

@GoodfireAI Is this data available? I’d love to take a look at it
English

@J33P4 @GoodfireAI We indeed expected it to be there! But what's novel is understanding how the model represents the tree of life (manifold structure and low-dim subspace) and the techniques we developed to isolate the representations, which we plan to extend to more complex bioinformatic questions
English

@GoodfireAI Why is this surprising when it was trained on the tree of life? And that it is well recognised you can detect this from phylogenetic types of comparisons. What’s new about this?
English

@thebasepoint @GoodfireAI Good catch! That's written wrong. Beta is learnable, but alpha is just a hyperparameter. We'll update it
English

@GoodfireAI A tiny question: how is alpha (a loss coefficient) learnable? Seems like if you let it vary the model would send it to zero (or negative).

English

@Sauers_ @GoodfireAI Agree, more would be better! We used 2400+ species but did notice that certain clades were a bit underrepresented by the sampling. Those ones tended to be placed in the center of the umaps.
English

@Sauers_ @GoodfireAI We haven’t looked yet, but it’d be interesting to see how the species representation changes along sequences with different ancestry due to horizontal gene transfer.
English

@Sauers_ @GoodfireAI We're definitely interesting in better interpreting the directions so those types of metrics could be useful there. Most metrics are based on direct sequence comparisons, though, which the model isn’t necessarily doing.
English

Excited to share our work digging into how Evo 2 represents species relatedness or phylogeny. Genetics provides a good quantitative measure of relatedness, so we could use it to probe the model and see if its internal geometry reflects it.
Goodfire@GoodfireAI
Arc Institute trained their foundation model Evo 2 on DNA from all domains of life. What has it learned about the natural world? Our new research finds that it represents the tree of life, spanning thousands of species, as a curved manifold in its neuronal activations. (1/8)
English
Michael Pearce รีทวีตแล้ว

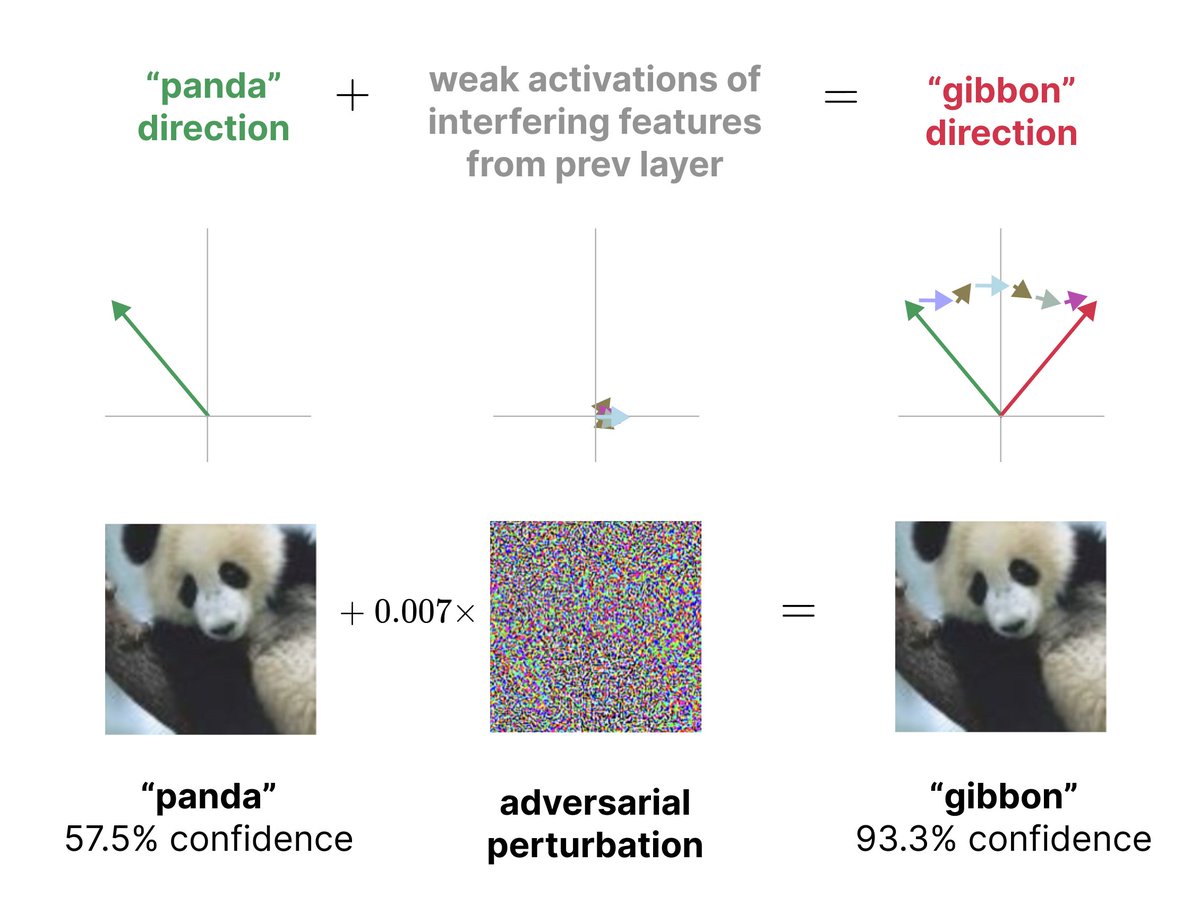
Adversarial examples - a vulnerability of every AI model, and a “mystery” of deep learning - may simply come from models cramming many features into the same neurons!
Less feature interference → more robust models.
New research from @livgorton 🧵 (1/4)
English
Michael Pearce รีทวีตแล้ว
Michael Pearce รีทวีตแล้ว