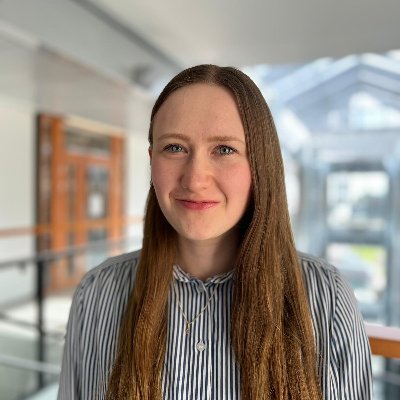
Emily giving an absolutely brilliant presentation @abnconf, reporting the latest findings from @PregnancyMs. Really interesting data coming through - thank you to everyone that has contributed
The ADAMS Study
338 posts


@adams_study
Multiple Sclerosis genetics in diverse ancestries. A study @preventiveneur1. Tweets by @ben_m_jacobs. More info: https://t.co/wJwCPGAZ9h

Emily giving an absolutely brilliant presentation @abnconf, reporting the latest findings from @PregnancyMs. Really interesting data coming through - thank you to everyone that has contributed













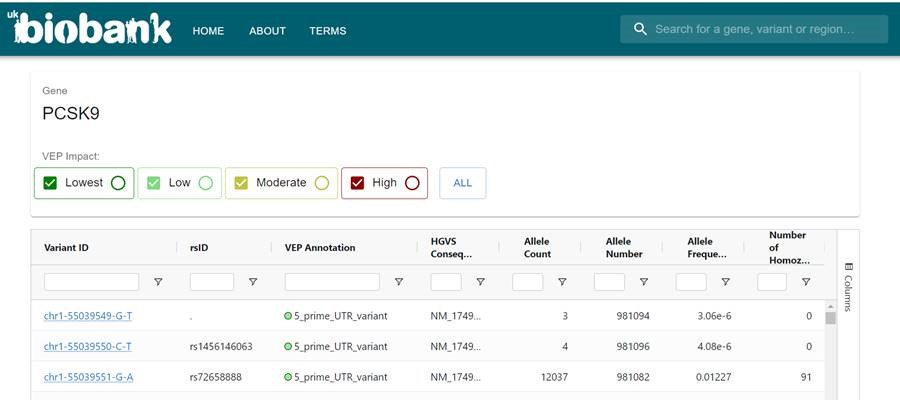








We’re excited to share that the RGC-ME Browser is live! The RGC-ME Browser offers allele frequencies, ancestry estimates, constraint metrics, and transcript annotations based upon a catalog of human protein-coding variations from the exome data of ~1M individuals.

Massive new @gnomad_project release (v4) spanning exome/genome data from *over 800,000 individuals*. Huge congratulations to the entire gnomAD production team for their incredible work getting this over the line! A total game-changer for rare variant interpretation.