harm
128 posts

harm
@harm__w
Scientific co-founder and head of research at Neptune Bio






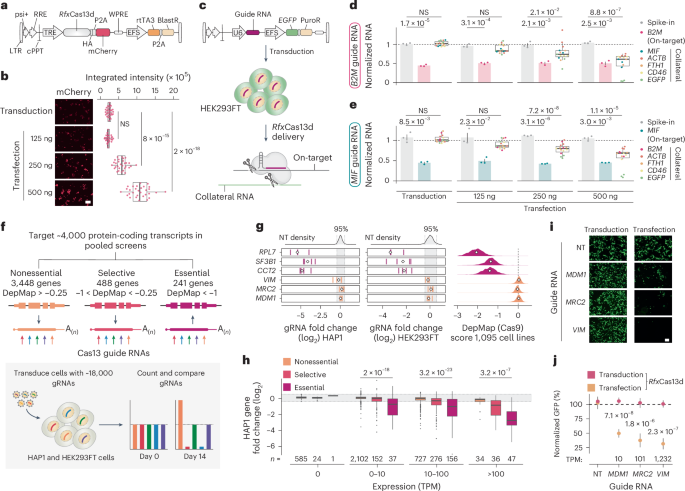
🚨🚨🚨 Thrilled to share our newest study — identifying functional long noncoding RNAs using transcriptome-scale RNA-targeting CRISPR screens. 🔎🔎🔎 We hope this scalable approach will be helpful for folks who want to connect noncoding RNAs to phenotypes & diseases.


Interested in 3' UTR changes in scRNA-seq? Check out our: * CPA-Perturb-seq paper * UCSC Perturb-seq tracks (see who regulates your favorite 3’ UTR!) * Seurat extension for polyA site analysis (PASTA) * Deep neural perturbation network (APARENT-Perturb) cell.com/cell/fulltext/…

Excited to share our work on targeting Cas13 to splice junctions for isoform-specific RNA knockdown. We learned a lot about the experimental design, assessment and downstream analysis of this approach. Phenomenal effort from all involved! biorxiv.org/content/10.110…








Using Cas13d guide design to control gene dosage enables exiting new applications and represent an alternative to recent microRNA (@YaleMichaels), dCas9 (@MarcoJost_) or degron approaches (@GemmaNoviello). 10/13