KJWON_LAB ری ٹویٹ کیا

🔬Can whole-slide Hematoxylin and eosin (H&E) stained histology images predict gene expression profiles in single-cell spatial omics? #spatial #SingleCell
Introduce our latest paper, BLEEP (Bi-modaL Embedding for Expression Prediction): a state-of-the-art tool predicting spatial gene expression profiles purely from histology images.
👉 Paper: arxiv.org/abs/2306.01859
👉 Code: github.com/bowang-lab/BLE…
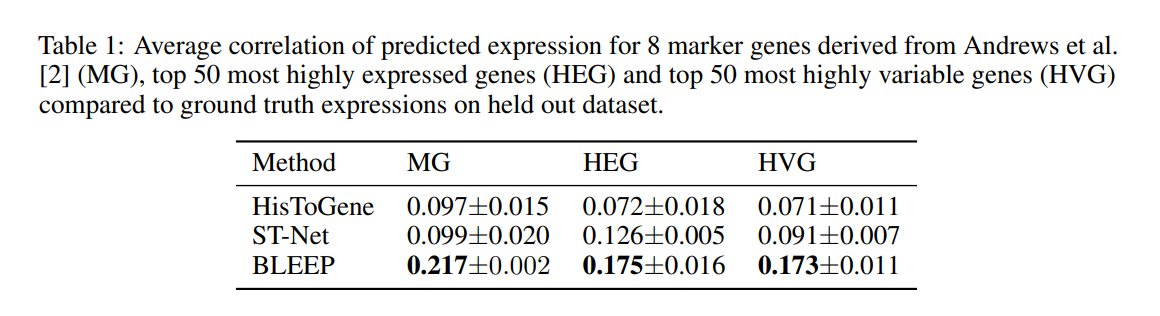
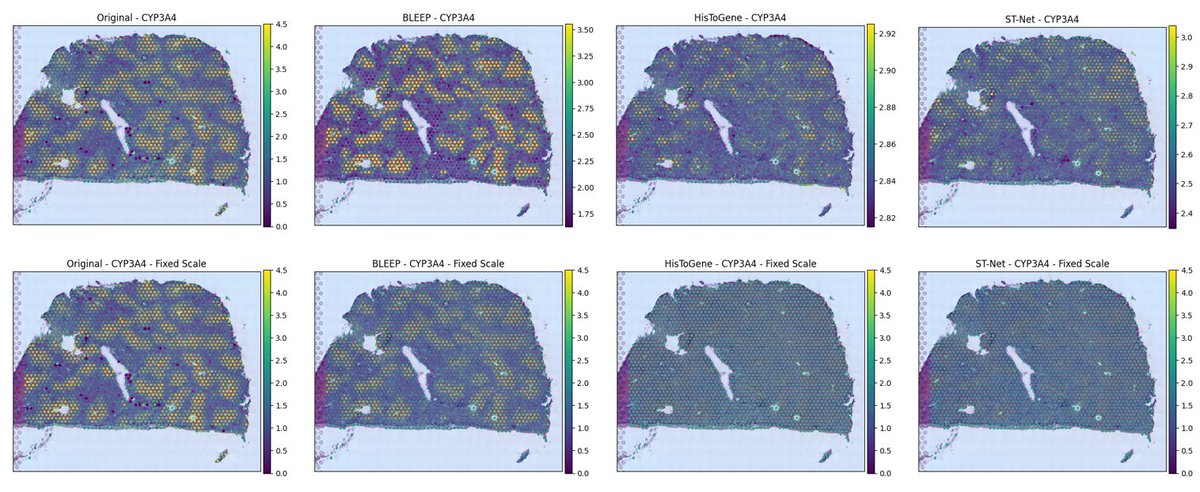
Compared to alternatives, it offers superior correlation & captures the variance of original expression profiles more effectively.
Understanding molecular mechanisms behind tissue architecture is key to unraveling disease pathways & formulating effective treatments. Yet, gene expression profiling is often a costly, lengthy process.
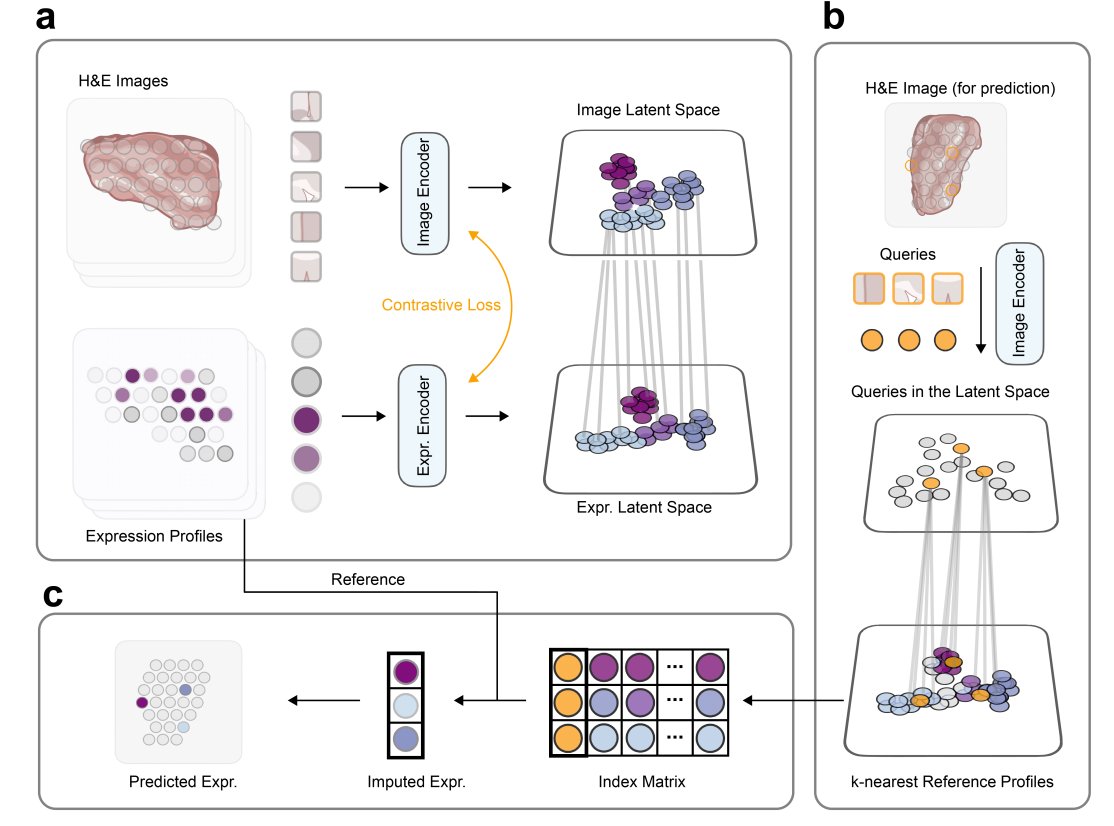
BLEEP is a game-changing bi-modal embedding framework. It uses a contrastive learning approach to create a low-dimensional joint embedding space from paired image & expression profiles at micrometer resolution. This space serves as a reference for predicting expression in unknown image patches from whole-slide H&E stained histology images.
Benchmarked on a human liver tissue dataset from the 10x Visium platform, BLEEP outperforms ST-Net & His2Gene in terms of gene expression correlation & variance.
All the credits go to the amazing PHD student Ronald (@RonaldXie1 ), who is co-supervised by @garybader1 and myself. @pmcc_ai @VectorInst @UofT_TCAIREM @DonnellyCentre @UHN
English