Arpan Sarkar ری ٹویٹ کیا
Arpan Sarkar
10 posts

Arpan Sarkar
@arpsark
PhD candidate in the Eddy Lab @Harvard interested in protein ML for modeling protein domains and predicting homology
شامل ہوئے Mayıs 2024
243 فالونگ41 فالوورز

Excited to be a part of this team! Check out our work investigating scientific problem solving in LLMs!
Alan Li@alanli2020
1/9 🚀 New paper: Demystifying Scientific Problem-Solving in LLMs — How does reasoning enhancement affect knowledge recall, and do LLMs benefit from external knowledge complimentary to reasoning? Tldr; 📊 SciReas: holistic and efficient evaluation suite for scientific reasoning 🧠 KRUX: a novel framework to study knowledge vs reasoning in LLMs 🔑 Findings: knowledge is a bottleneck; reasoners + in-context knowledge help; long CoT helps knowledge recall/utilization
English
Arpan Sarkar ری ٹویٹ کیا

(1/7) Very excited to share my first PhD preprint on the interactions of two of my favorite mobile genetic elements: phages and group II introns!
biorxiv.org/content/10.110…
English
Arpan Sarkar ری ٹویٹ کیا

Super excited to share my postdoc work investigating how mating and parental behaviors evolve using wild species of mice combined with single nucleus RNA-sequencing of the hypothalamus 🐭🧠🧬!
biorxiv.org/content/10.110…
English

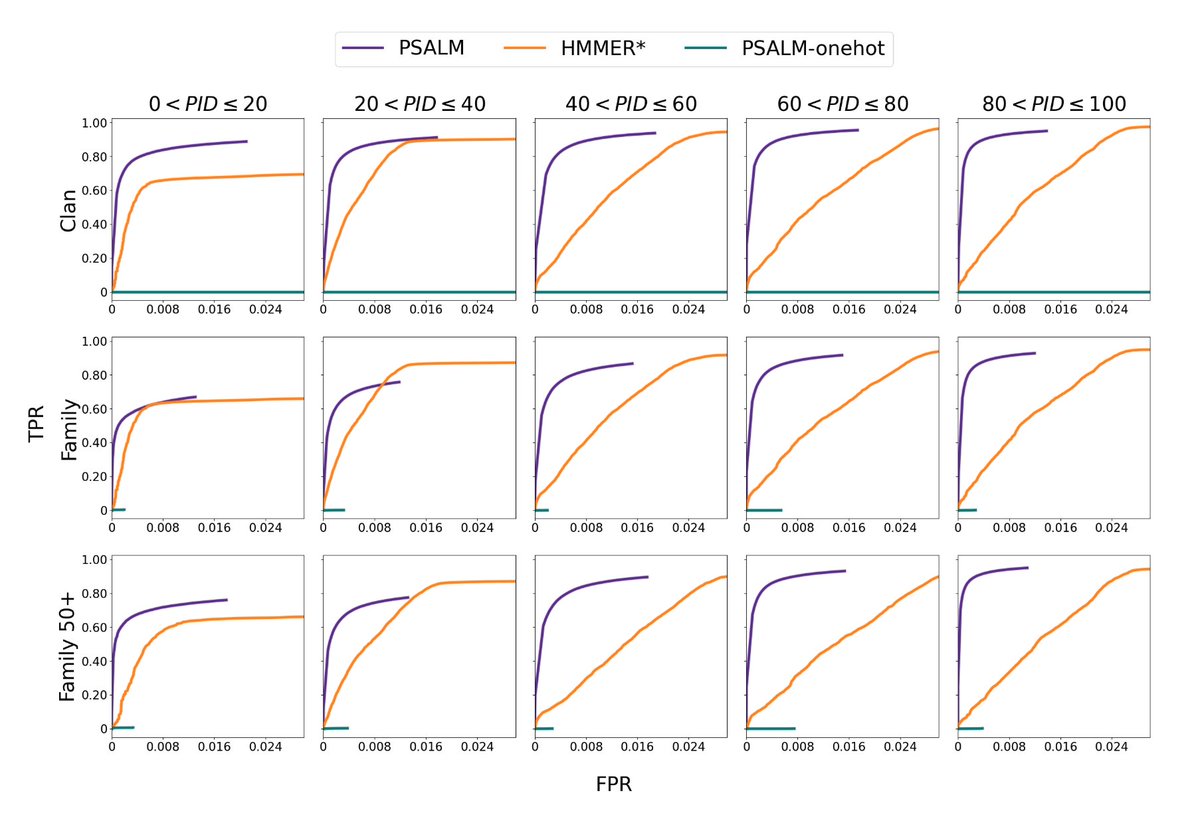
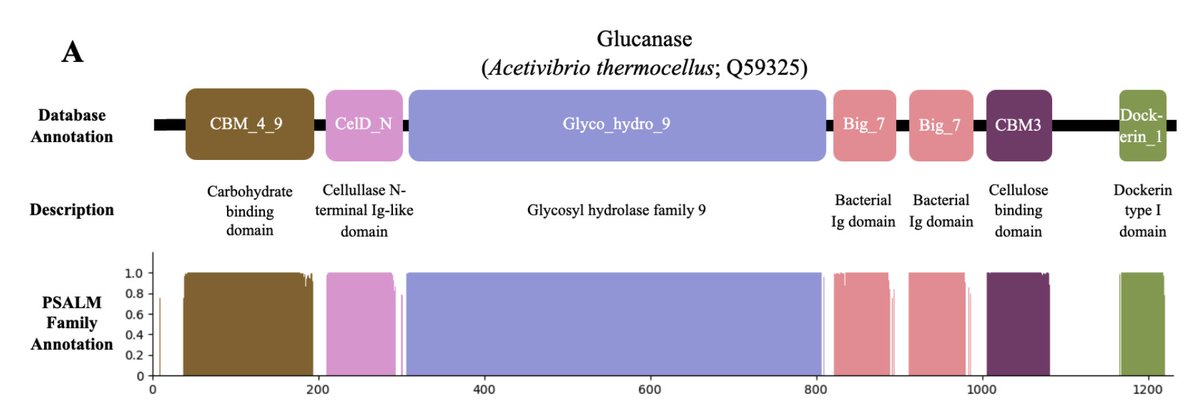
@sokrypton It's also important to look at the homology between any domain in any seq in test to any domain in any seq in training, and I think we do a good job of assessing performance at various domain-level max % identity thresholds in our work on PSALM (biorxiv.org/content/10.110…)
English

Excited to share our new work (with @KumareshSpikes): "Protein Sequence Domain Annotation using Language Models" (PSALM)
biorxiv.org/content/10.110…

English