Sam Kovaka
20 posts


Sam Kovaka
@samkovaka
Computational biology postdoc at JHU







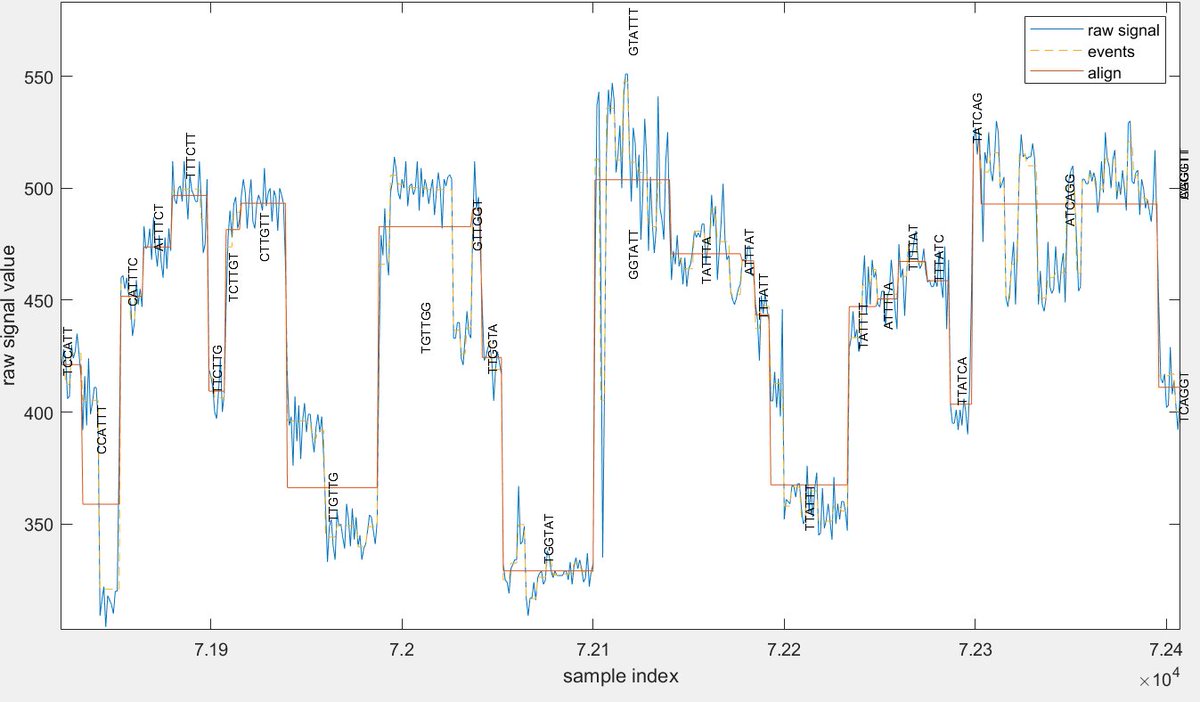
f5c-v1.1 is released and features a new module called resquiggle that aligns raw-signals to basecalled reads [contributed by @hiruna72]. github.com/hasindu2008/f5…

Congratulations to @samkovaka for defending his thesis on Computational Methods for Genomic, Transcriptomic, and Epi-Omic Analysis with Long-Read Sequencing! @timp0, @elapertea and I are so proud! @nanopore @PacBio @JHUCompSci @HopkinsEngineer

At #nanoporeconf, @Samkovaka will present UNCALLED4: an adaptive sampling toolkit for nanopore signal alignment & analysis. UNCALLED4 features interactive alignment visualisations & comparisons, & epigenetic modification detection stats: bit.ly/3Gy20sX









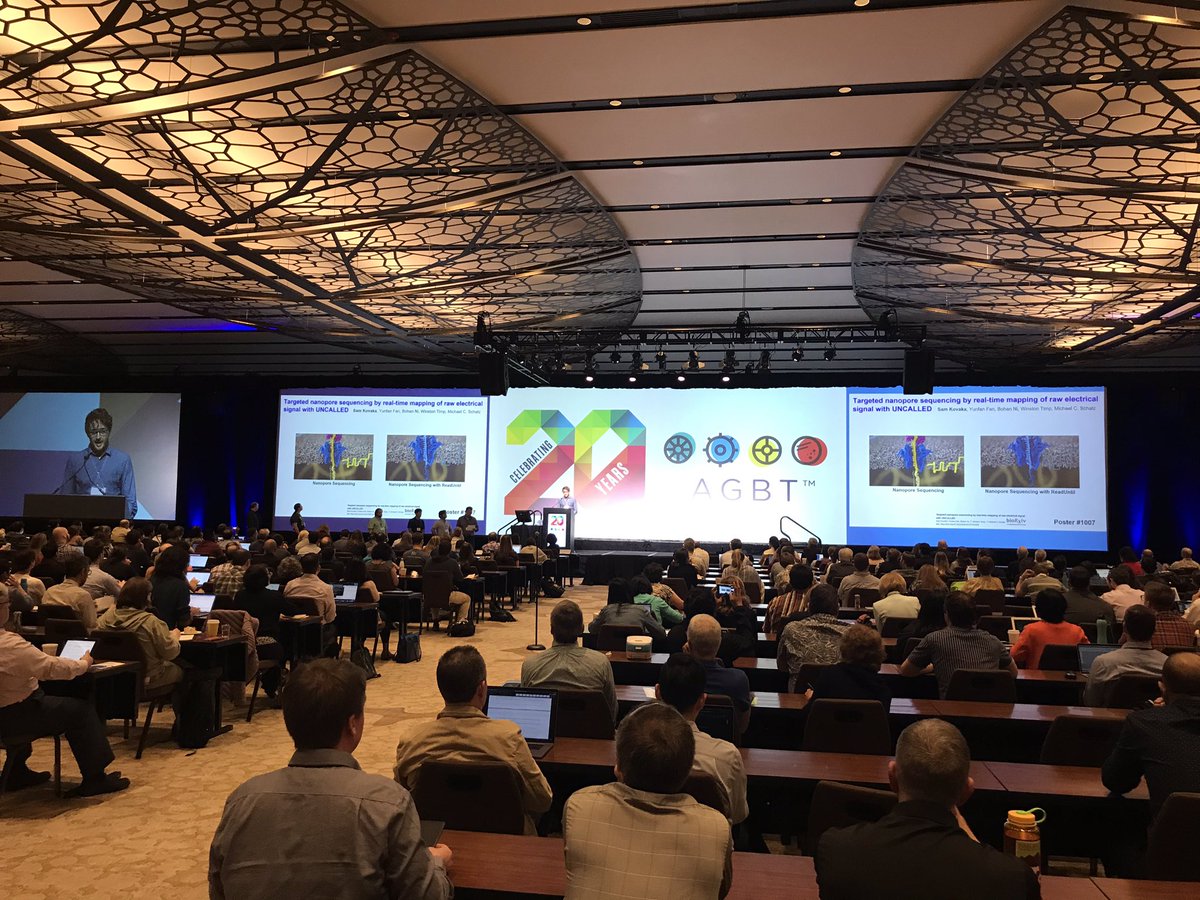
Excited to announce: Targeted nanopore sequencing by real-time mapping of raw electrical signal with UNCALLED. Awesome work with @samkovaka, Yunfan Fan, @bohan_ni & @timp0 that will forever change how sequencing is done. @JHUCompSci @JHUBME @nanopore nature.com/articles/s4158…



Excited to announce: Targeted nanopore sequencing by real-time mapping of raw electrical signal with UNCALLED. Awesome work with @samkovaka, Yunfan Fan, @bohan_ni & @timp0 that will forever change how sequencing is done. @JHUCompSci @JHUBME @nanopore nature.com/articles/s4158…





Excited to announce: Targeted nanopore sequencing by real-time mapping of raw electrical signal with UNCALLED. Awesome work with @samkovaka, Yunfan Fan, @bohan_ni & @timp0 that will forever change how sequencing is done. @JHUCompSci @JHUBME @nanopore nature.com/articles/s4158…