

HeilemannLab (👉🏻 heilemannlab.bsky.social)
503 posts


@HeilemannLab
Single-molecule biophysics lab at Goethe University Frankfurt. Super-resolution microscopy. We moved 👉🏻 https://t.co/hdHe9Trgyb








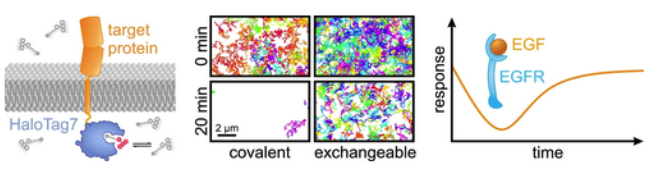
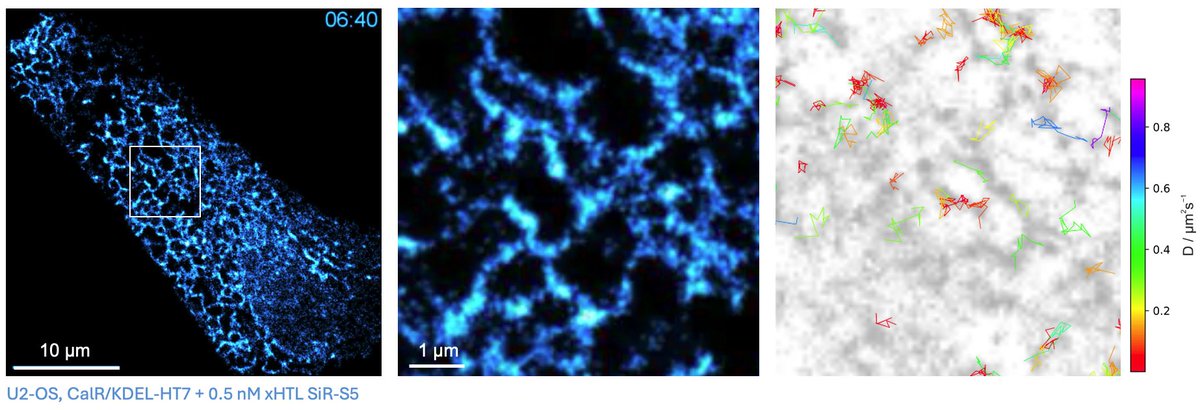
Long-Term Single-Molecule Tracking in Living Cells using Weak-Affinity Protein Labeling (Mike Heilemann and co-workers) @HeilemannLab #openaccess onlinelibrary.wiley.com/doi/10.1002/an…


Check out how we used simulations and single molecule FRET to determine how a invasion bacterial protein activates a plasma membrane receptor to enter our cells. Kudos to @serena_arghittu and @gabriel_hella, and great collaboration with @HeilemannLab nature.com/articles/s4146…


If robots could dream of microtubules, how would they look like? An amazing story by Alon Saguy, @ShechtmanLab and colleagues. Proud we could contribute to it onlinelibrary.wiley.com/doi/10.1002/sm…





Out today from the Vandenberg and Dedecker labs! PSF splitting with the ‘Circulator’, which encodes the fluorophore emission band into the PSF, improves the information content of fluorescence microscopy and enables improved super-resolution imaging and single-particle tracking.

Preprint alert🚨Did you ever experience issues in labeling kinetics and fluorescence brightness, when working with SNAP-tag in live cells? Our newly developed SNAP-tag2 system might be the solution! 🚀Check out our latest preprint on #bioRxiv: 🔗biorxiv.org/content/10.110… (1/4)




