algobaker
4.2K posts


algobaker
@algobaker
ML PhD student Previous: research at LLM hyperscaler, AI in pharma.



Today we're launching Latent-Y: the world's first autonomous agent for drug design, lab-validated end to end. Give it a research goal. Latent-Y reasons, designs, iterates, and delivers lab-ready antibodies, autonomously or collaboratively, with the biological reasoning of a PhD protein design expert. Technical report: tinyurl.com/latent-y-techr… Blog post: latentlabs.com/latent-y Apply for access: platform.latentlabs.com








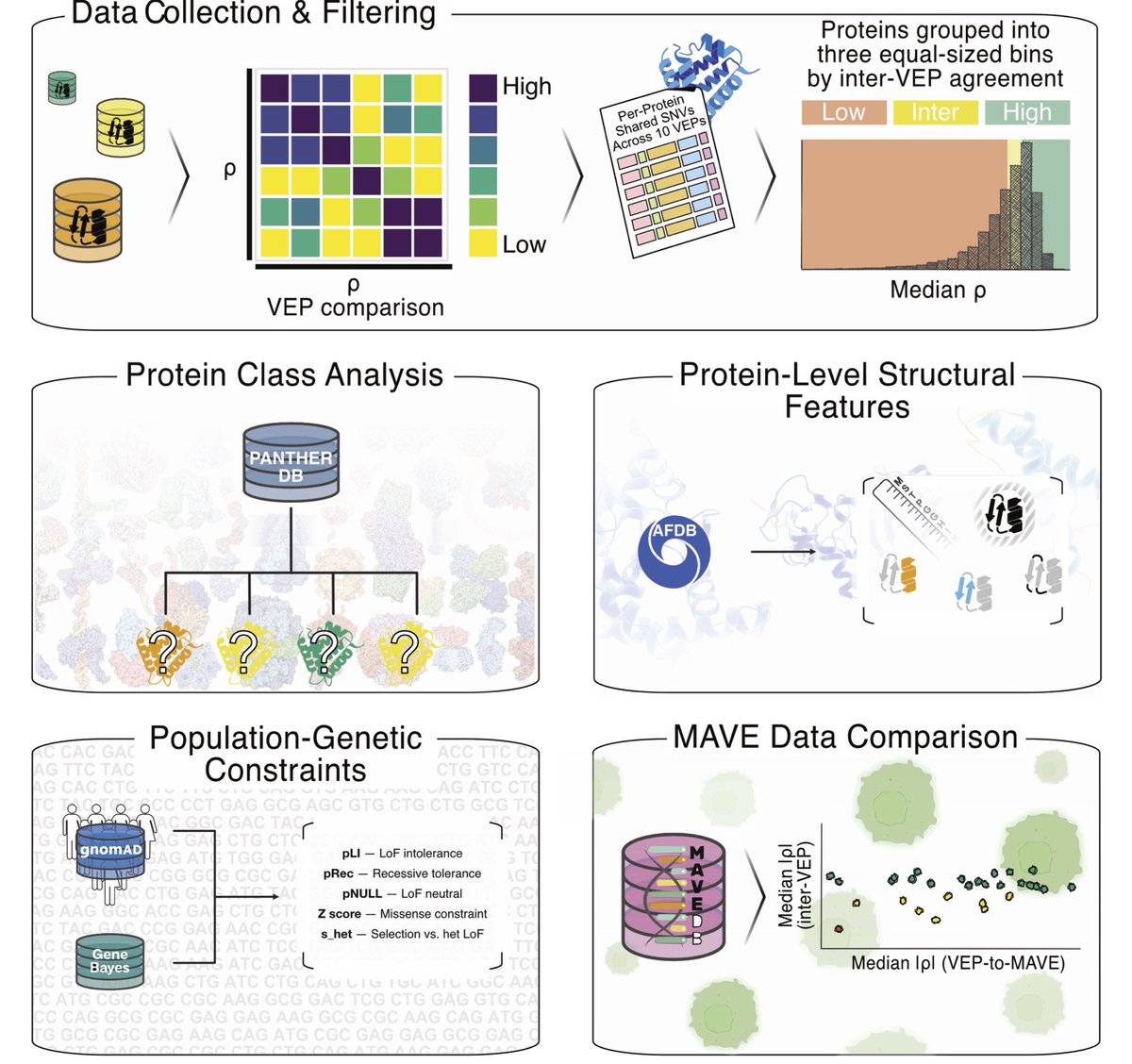
Disagreement among variant effect predictors guides experimental prioritization of target proteins biorxiv.org/content/10.648… #biorxiv_bioinfo


Few know this, but I (George) was the only person in history to get a perfect score in CMU compilers, which is likely the best compilers course in the world. Combine that with crazy low level knowledge of hardware from 10 years of hacking. Then add a team of people who are talented enough to push back on my dumb ideas and clean up the implementations of the good ones. The team who keeps this whole operation running, software, infrastructure, and product. I love how there's no hype in deep learning compilers. It was one of the most annoying things about self driving cars, all the noobs who burned through billions on crap that was obviously dumb, and the companies who deserved to go bankrupt years ago if not for government bailouts (Tesla and China will devour them all). In this space, the competition is @jimkxa at Tenstorrent, @clattner_llvm at Modular, and @JeffDean at Google. Three of the living legends of computer science. And companies like @nvidia and @AMD, who are definitely live players, making single chips that have more power than the whole Internet two decades ago. This space is so fun to play in. If you haven't, read the tinygrad spec. It's all coming together beautifully.







Extremely excited to announce LigandForge 🧬⚡ Generate high-quality peptides at over 10,000x - 1M the speed of state-of-the-art methods like Bindcraft and Boltzgen. Predict binding affinity with 83% correlation to experimental binding data. 150 protein targets benchmarked.



Do people like this? We don't do this for codex because it exists to help you and it's important that you remain the owner and accountable for your work without AI taking credit. At the same time it does mean that you can't trace how popular codex is among repos.




There's a new generation of empirical deep learning researchers, hacking away at whatever seems trendy, blowing with the wind... no accumulation of real understanding, or foundations. No real passion or depth, just light amusement and career advancement. I'm hoping it's a phase.