
Ex-microbiologist
190 posts



@GenomicsCow How is that different from what they offer now?
English

@kenbwork What makes you think folks have not done this?
“If you were to build a company just to commercialize vanilla single cell at scale…what would tissue dissociation, viability staining, purification, library prep, analysis, etc. actually look like? ”
English

@kenbwork It is likely worth spending timing talking to CROs about their challenges and cost structure. Growing a physical lab to address a rapidly changing market using many assays is a difficult task. It may waste money to “over” optimize when things change quickly.
English

Have been waiting for someone to build a CRO that delivers existing "boring" experiments much faster and cheaper by:
1. Questioning and rethinking each step of the laboratory workflow with modern automation, microfluidics, hardware, software. If you were to build a company just to commercialize vanilla single cell at scale, and you lived and died by the margins you could create over the period of a few years, what would tissue dissociation, viability staining, purification, library prep, analysis, etc. actually look like?
Remember the actual constraints and that there are many ways to combine reagents together that do not involve pipettes, eppendorf tubes, well plates, people (and that includes pipette wielding robots like Hamiltons and Opentrons). Current laboratory workflows are overly anthropomorphized and artifacts of bad tools that have not been thought through much since molecular biology emerged in the 1960s.
2. Rebuilding new tools for each step that are uniquely enabled by large volumes of samples. For example, semiconductors could be perfect for the mass perturbation of charged biomolecules with precision. You can probably massively parallelize most experiments on these things with new, desirable behavior. For instance, no need for lots of copies of DNA in transduction if you can precisely move a single molecule next to the target cell - greater efficiency, throughput, cost. Or you can miniaturize phosphoramidite synthesis into millions of 65 nm squares using voltage induced redox chemistry to serially deprotect your sequence.(ieeexplore.ieee.org/abstract/docum…).
There is some volume of business for a CRO that would justify designing and fabbing fit-for-purpose large node semiconductors for many of these laboratory workflows.
3. Simply executing on operations, setting ambitious timelines for completing projects and delivering data to clients in interactive data portals.
Ripley's Believe It or Not, the bar in this space is 3 months, $20K and a static PDF/spreadsheet to show for a standard bulk RNA-seq experiment.
owl@owl_posting
There aren't enough smart people in biology doing something boring owlposting.com/p/there-arent-… new argument piece i often meet a lot of founders in biology pursuing really crazy, revolutionary ideas. i love them! but i wish there were more people pursuing the boring ideas 🧵
English

@OmicsOmicsBlog @GenomicsCow I was thinking along the lines of the comment from @GenomicsCow about microscopy being commoditized compared to NGS. The imaging tech behind sequencers still amazes me
English

@microbiol_sci @GenomicsCow There’s also the fun bit that many sequencing instruments ARE microscopes at heart, but aren’t used to literally image the DNA sequence directly.
Analogous to how a large number of NGS assays measure biological phenomena that aren’t directly connected to DNA sequence
English

1/3
Sequencing is the new microscope
Credit: ldeming.posthaven.com/sequencing-is-…
But I’d take this catchphrase a slightly different direction . . .
Genomics Cow@GenomicsCow
Proteomics could end up as an “app” for NGS, like we see with scRNA-seq, ChIP-seq, etc. If this happens, things could get interesting with the non-NGS core labs
English

@LGMartelotto @GenomicsCow @mehtamanal @hoheyn @OmicsOmicsBlog @MollyminHe Your approach was incredibly clever, and you released the protocol in a way that others can easily replicate. The field will appreciate it so much!
English

@microbiol_sci @GenomicsCow @mehtamanal @hoheyn @OmicsOmicsBlog @MollyminHe Thanks Anastasia! Very wise words. We worked hard to make it simple and functional for everyone…and we were eager to put the protocol out there so this amazing community can start using it.
English

STAMP protocol from @hoheyn could be the death knell for 10x Genomics $TXG.
“cost per cell is approximately one half of a cent, and the method does not require additional analysis by sequencing”
genomeweb.com/gene-expressio…
@OmicsOmicsBlog @AlbertVilella @LGMartelotto @MollyminHe




English

@GenomicsCow @mehtamanal @hoheyn @OmicsOmicsBlog @LGMartelotto @MollyminHe An assay isn't a product, but you can't have a product without an assay.
This is incredibly exciting work, but it takes a lot of work to turn something that works to something that works for most people, most of the time. That said, I'm excited to see the future for this tech!
English

@mehtamanal @hoheyn @OmicsOmicsBlog @LGMartelotto @MollyminHe Stamp’s promising but remember that *most* researchers use a technology because it’s widely available and easy to that’s in the form of commercial kits and support from your local core, and cores have sequencers not spatial imagers , and are more familiar with sequencing 😎
English

@ToumyGG @GenomicsCow @CatharineAquino Isn't that the definition of "funding" though? These all rely on having sufficient funding to build something that meets rapidly changing market needs AND test its commercial viability.
English

@GenomicsCow @CatharineAquino I don’t think funding is the only solution. A sustainable business model by the company and realistic expectations by customers around the cost of instruments/reagents and services. And also realistic expectations by the investors on ROI.
English

@CatharineAquino Making flexible, innovative, and disruptive tools is a seemingly impossible balance between being innovative, getting to market quickly, and building something robust. The grace to balance these often comes down to funding as @GenomicsCow suggests.
English

@KostChristian @markowenmartin @ATinyGreenCell That assumes there is no growth difference, which is pretty reasonable.
English

@microbiol_sci @markowenmartin @ATinyGreenCell I think this is the easiest explanation. But then the sectors should not appear towards the edge of the growing colonies, right?
English

This happens every year. It may not be mysterious, but I love it. I used E. coli containing plasmids with pigment genes from @ATinyGreenCell . I mix the cultures. The students do a dilution series. And sometimes we see what LOOK like sectored colonies. Or are they? 1/2
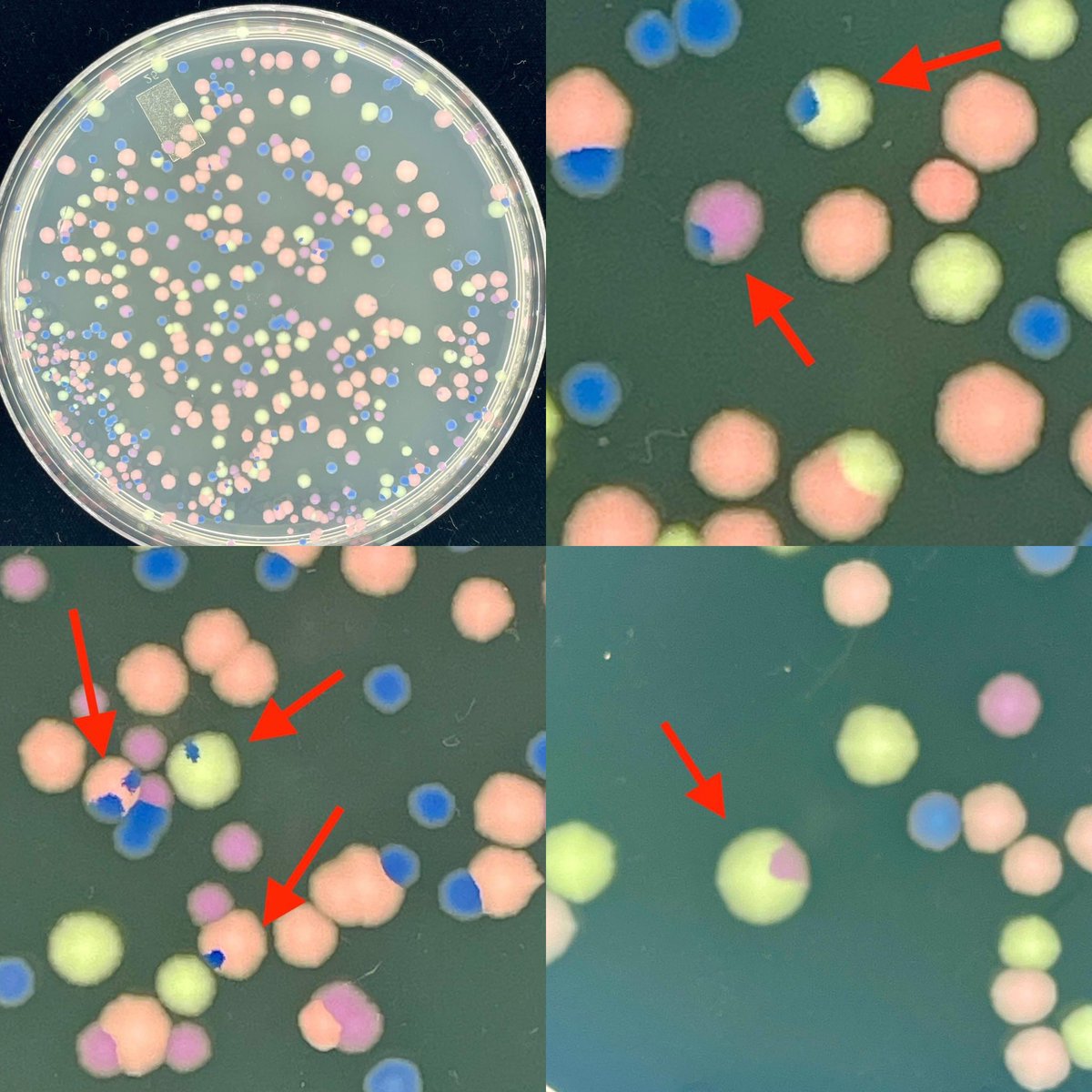
English

There's a lot of speculation about what this is. I have to admit, the teaser is quite effective; it has generated a lot of buzz. It resembles an eclipse. What are they trying to overshadow?
Illumina@illumina
The next big thing in sequencing is coming. And it’s small. Learn more at our virtual event on October 9 at 9 a.m. PT: bit.ly/3NdvvVb
English

@CatharineAquino @RocheSequencing @illumina @agbt They just had a Genome Web article in May saying they’re making “good progress” so maybe AGBT is to soon…
English

Same players but the order is different. Something new from @RocheSequencing ? @illumina wants to be on top of the food chain again? I hope the bag is nice 😂 @agbt sponsors 2025

English

@OmicsOmicsBlog Is it just me or is 10x rolling back their presence at many conferences?
English

@GenomicsCow No shade! They make great products. Their library keep products aren’t always best in class for life sciences folks, but great quality.
English

@microbiol_sci Illumina is pretty proud of their “R” in RnD. But perhaps they will become like other big companies and buy in the tech and focus on the “D” part more. No shame in that! Just gonna have some bummer acquisitions but that’s the same when doing blue skies research
English

The old message “We compete with our clinical customers”
The new message “We compete with our library prep partners”
Keith Robison@OmicsOmicsBlog
Illumina Would Like to Change the Conversation omicsomics.blogspot.com/2024/08/illumi…
English

We need a @dieworkwear for architecture...
Uni of Nomi@Uni_of_Nomi
@ATinyGreenCell Meanwhile in Rotterdam...
English

@bioinformer @illumina They still aren't that cheap though? What do you think the ASP is?
English

someone said on a call today that @Illumina's iSeq are cheap enough that they basically are at the bottom of kids' cereal boxes now. LMAO...
a few minutes later they posted this in the meeting chat...
everyone like💀

English

@lpachter That is true art!
My head knows upset plots are better, but they can be difficult to interpret without careful thought. Not every plot can be perfect i suppose.
English

@microbiol_sci Area proportional Venn diagrams are great for displaying 2^[3] data. And of course Venn art is wonderful. I do find that for 2^[n] where n>3 UpSet plots are more useful.

English

Confession: in graduate school I would occasionally procrastinate by trying to find Venn diagrams. There are dozens of open problems.
Some of the constructions are beautiful, e.g. this symmetric 11-Venn diagram from arxiv.org/abs/1207.6452
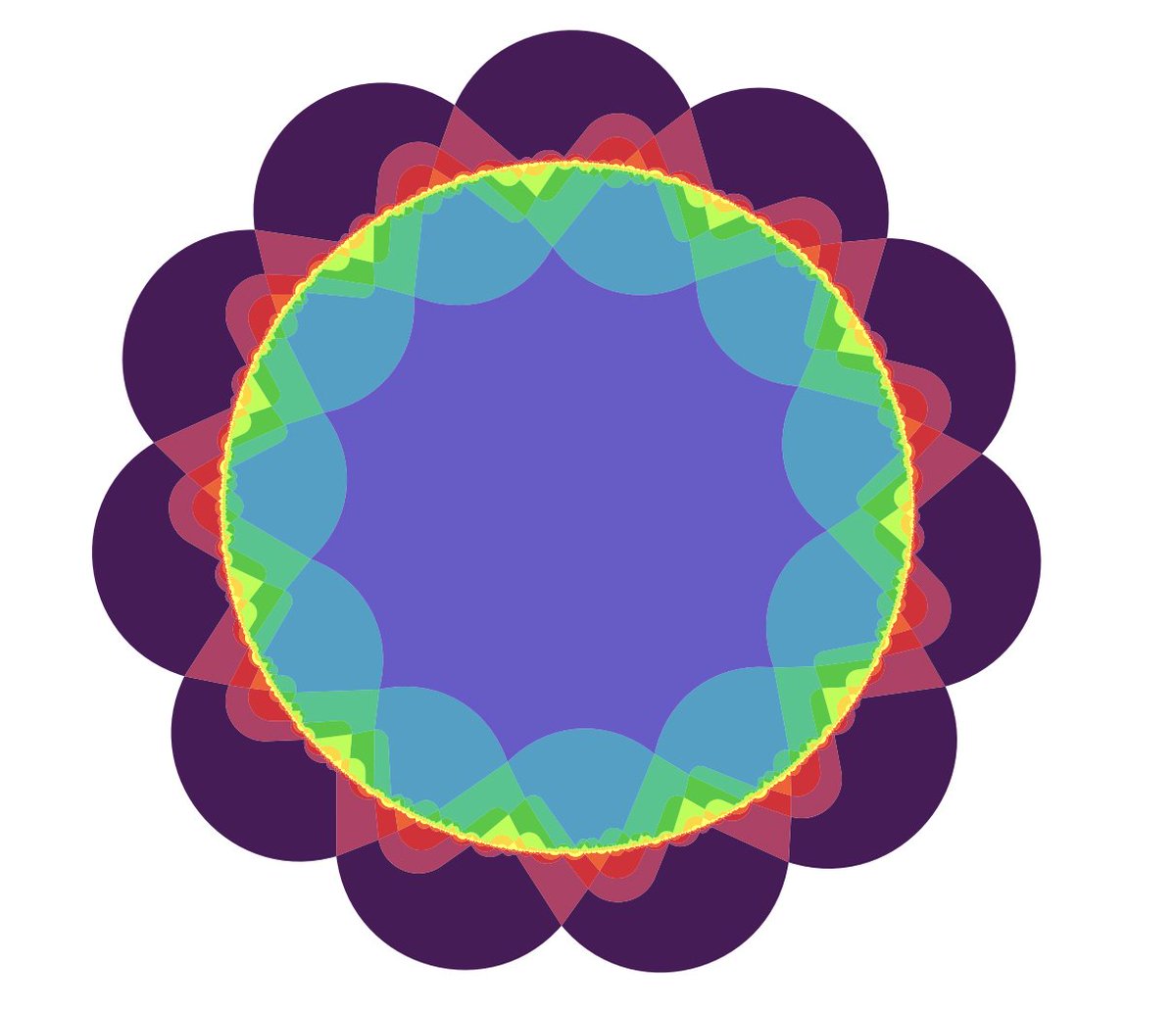

English