Tim Aikin
926 posts

Tim Aikin
@TheDynamicTim
New Ventures @ForesiteLabs applying emerging technology to biotech and therapeutics. Former @NSFGRFP fellow studying cancer & signaling dynamics @JohnsHopkins


While humans acted as GPT-5’s hands for carrying out the protocols, we also piloted an autonomous robot. It was built to execute arbitrary Gibson cloning protocols from natural language, with human supervision for safety.

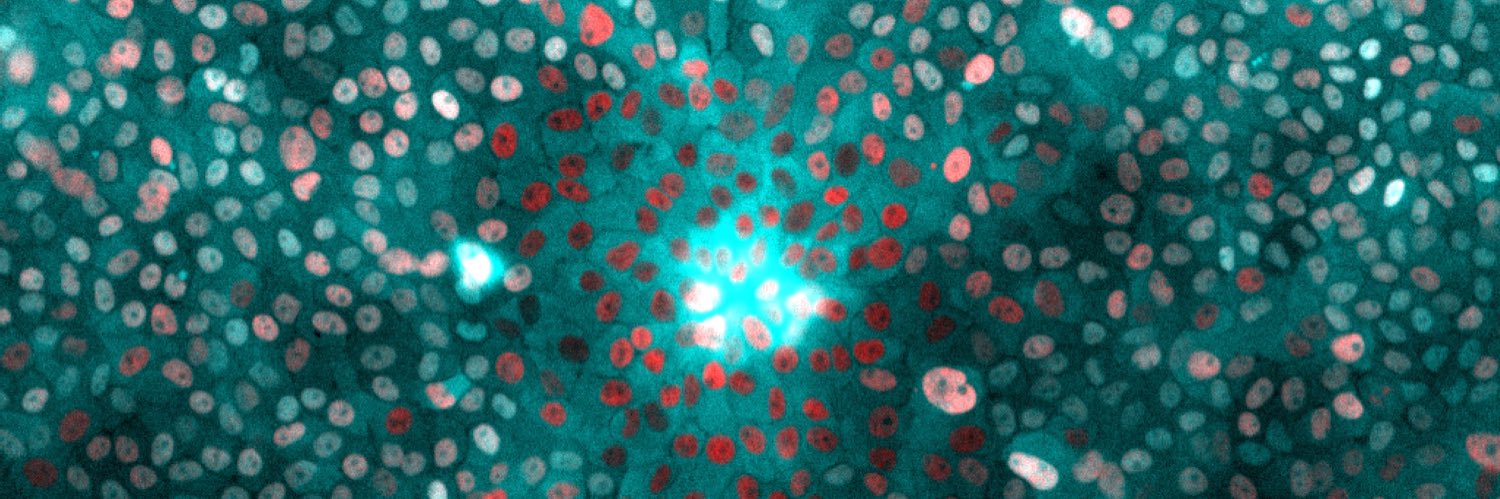
[Research] Freeze-framing the cellular world to capture a fleeting moment of cellular activity dlvr.it/TMgKNV #UOsaka #research




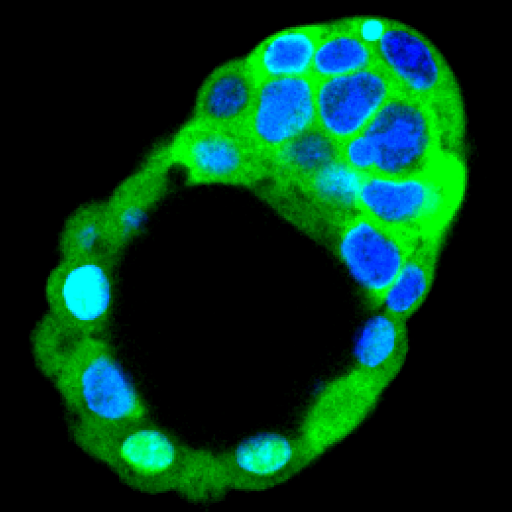
Exciting new inroads into human biology and therapy development: Human intestinal organoids with an autologous tissue-resident immune compartment! #Gjorevski #Cabon #IHB @Nature @IHB_Research #MOESM4" target="_blank" rel="nofollow noopener">nature.com/articles/s4158…




We are delighted to share the #tyrosine @KinaseLibrary, now in @Nature: nature.com/articles/s4158… Tyrosine predictions and enrichment analysis are now available: kinase-library.phosphosite.org #TheKinaseLibrary #KL #part2 🧵 1/9



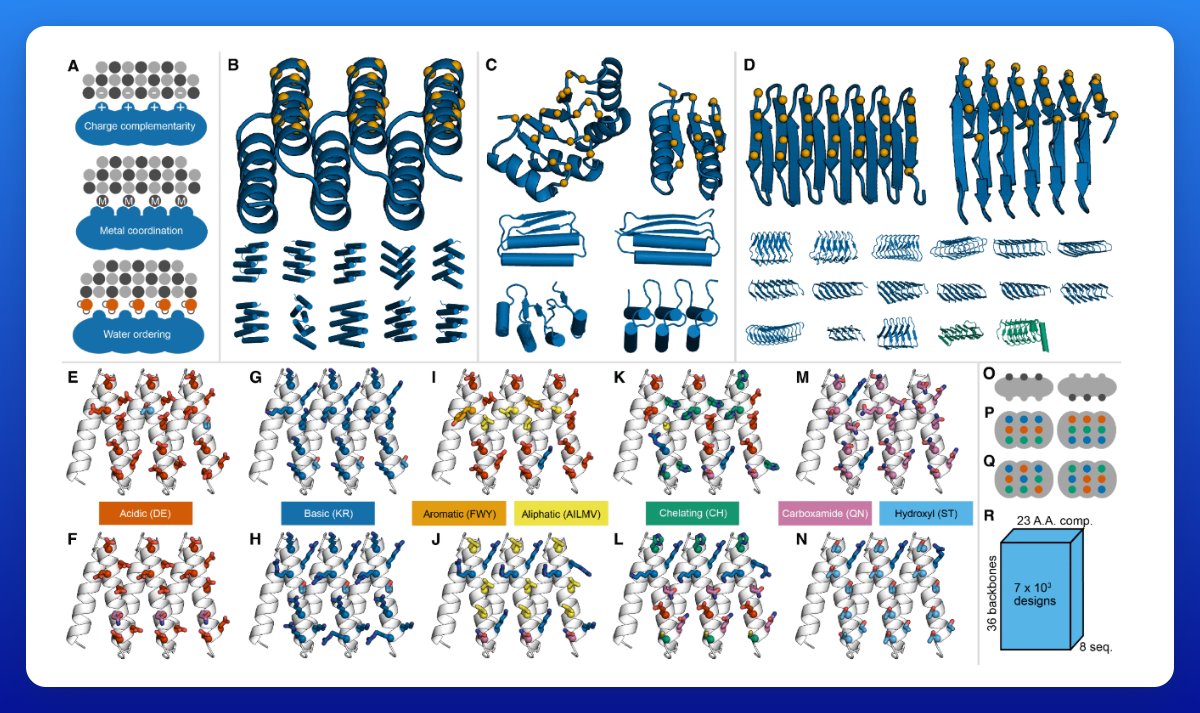
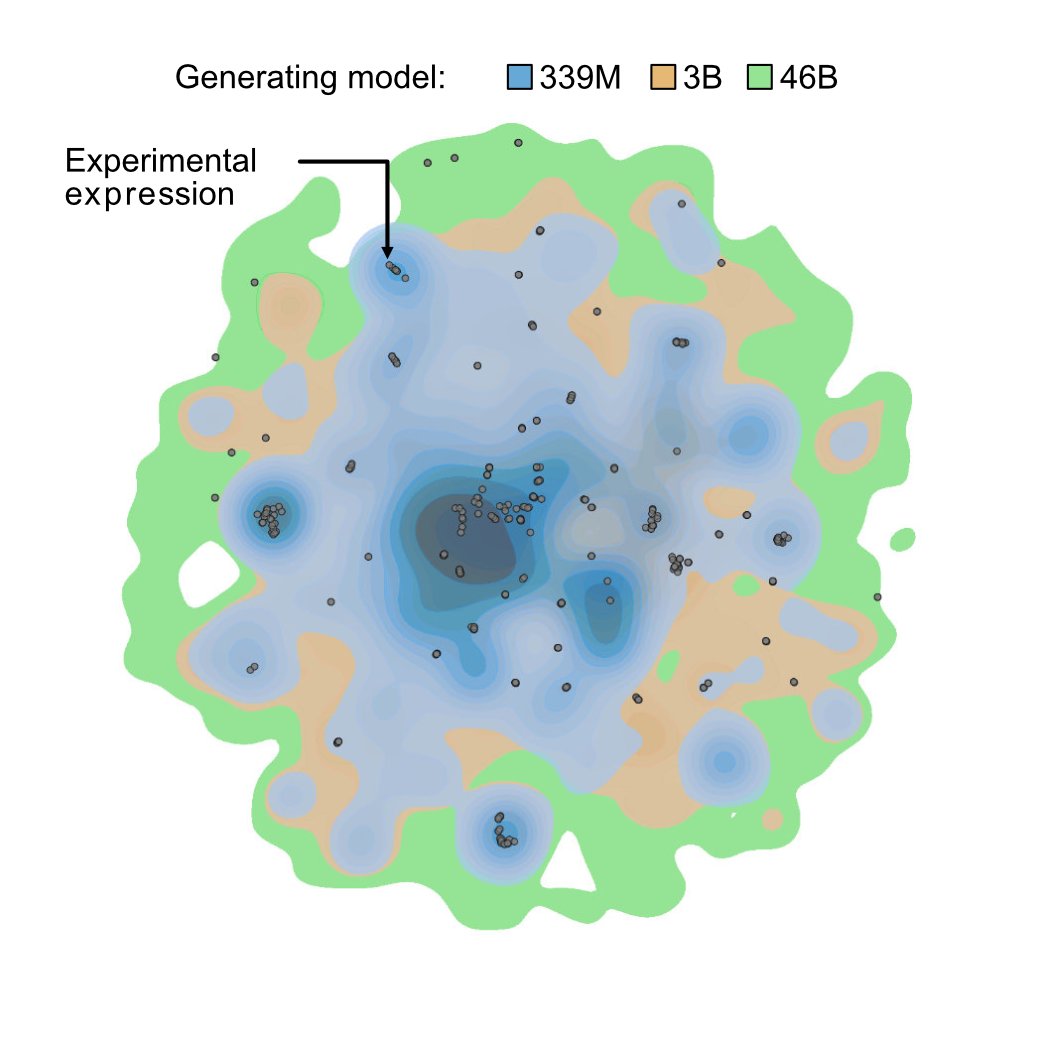
One of the first projects we took on at @ProfluentBio was designing novel gene editing proteins with language models. This grew into an initiative called OpenCRISPR, and today I’m excited to share that work (including the sequences we designed)! biorxiv.org/content/10.110…

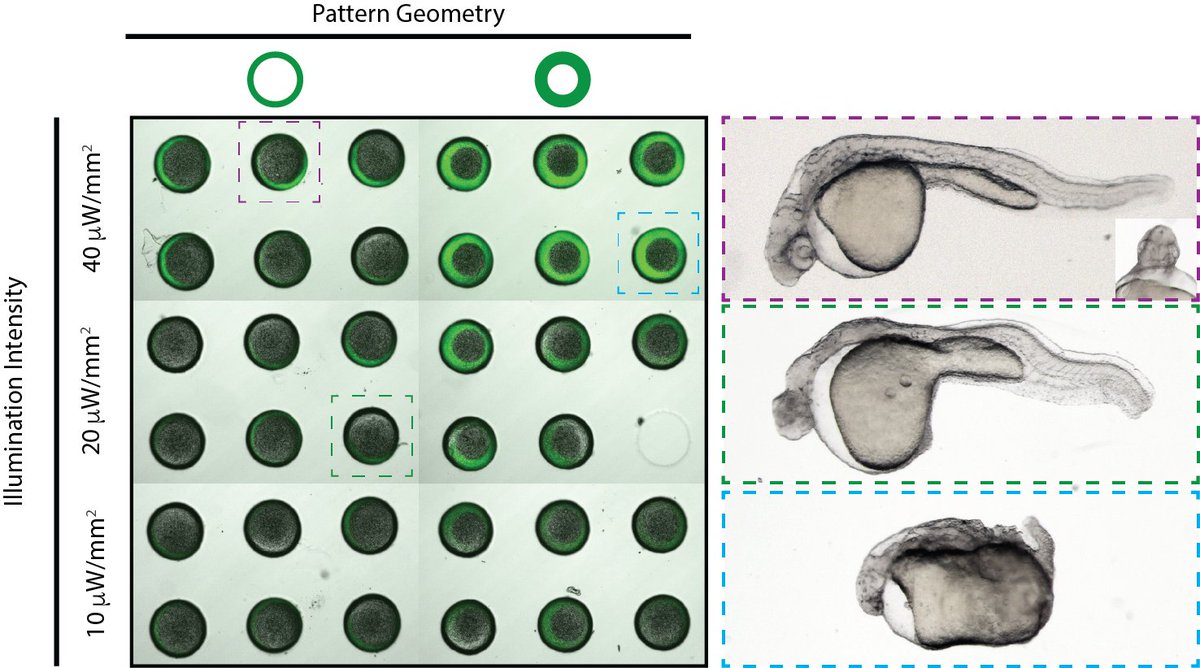
We’re very excited to share the first preprint from my lab along with @h4rrymcnamara, @BillZJia, @AdamEzraCohen, and Alex Schier. We describe a set of methods for exerting spatial control over Nodal signaling in zebrafish development. biorxiv.org/content/10.110…