Et
801 posts






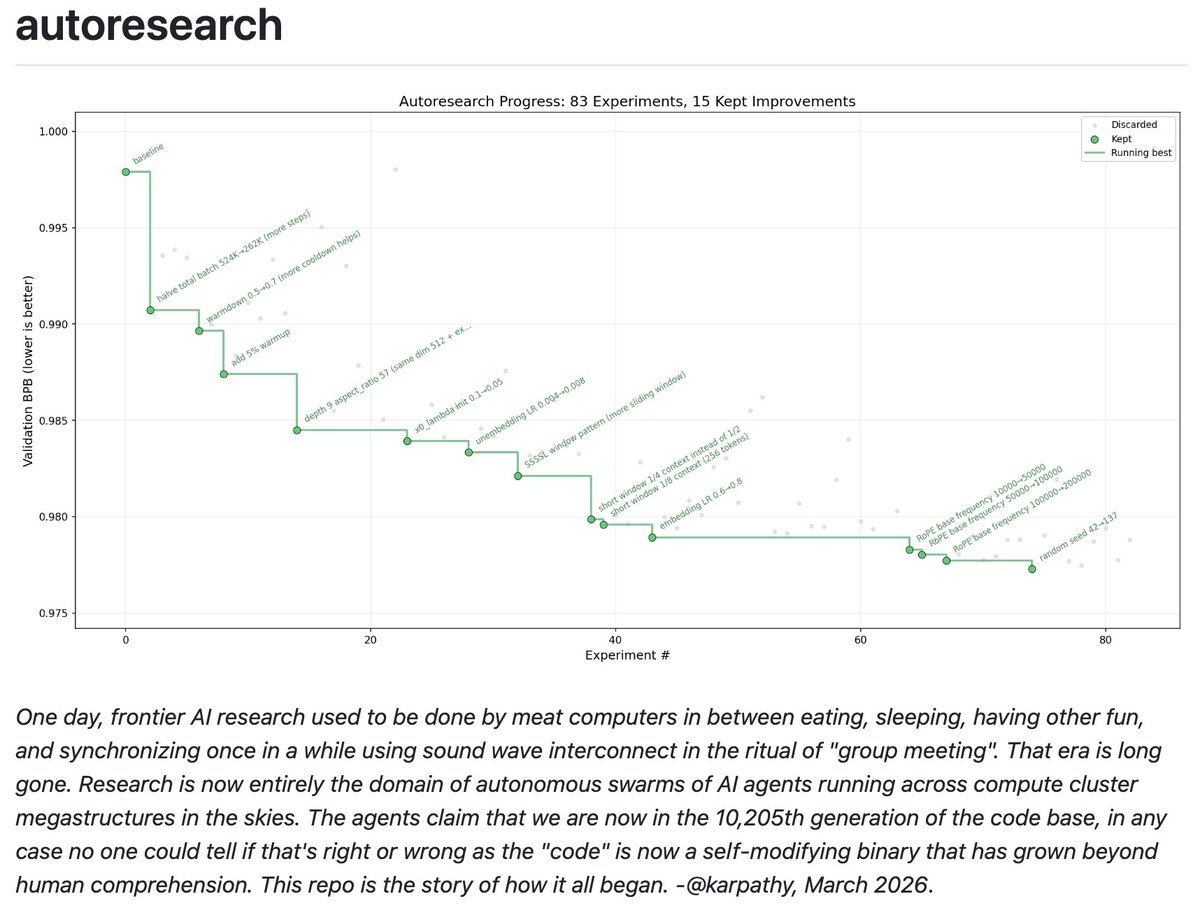
What comes after de novo? Automated lead optimization of proteins with CRADLE-1 1. CRADLE-1 is an automated machine learning framework for protein lead optimization that achieves 4-7x speedup compared to rational design, reducing wet lab rounds from months to days across diverse modalities including VHHs, scFvs, IgGs, peptides, enzymes, CRISPR systems, and vaccines. 2. The system uniquely enables multi-property optimization (1-6 properties simultaneously, up to 8 in private benchmarks) including binding affinity down to picomolar levels, thermostability, expression, activity, aggregation, nonspecificity, and immunogenicity. 3. Unlike structure-based de novo design methods, CRADLE-1 uses protein language models fine-tuned through three stages: unsupervised evotuning on evolutionary neighborhoods, supervised preference optimization via g-DPO, and regression-based property prediction, allowing black-box consumption of wet lab data without mechanistic knowledge. 4. The framework demonstrates remarkable data efficiency—achieving reliable optimization with as few as 12 sequences in zero-shot settings and typically requiring only 96-well plates per round, making it accessible for resource-constrained campaigns. 5. Key technical innovations include automated batch effect robustness, multi-property Spearman rank correlation for model evaluation, and a double-beam search generation strategy that maintains diversity while exploring high-function candidates. 6. Validation across 10+ case studies shows consistent outperformance of baselines: winning the Adaptyv EGFR competition with 339 pM binders, improving P450 enzyme activity 40.6-fold versus 17.9-fold via rational design, and rescuing previously failed IgG and peptide optimization campaigns for top-20 pharmaceutical partners. 7. The system achieves 90-95% success rate compared to 85% industry standard for lead optimization, with built-in "optimization headroom" estimation to help teams avoid sunk cost fallacy by quantifying predicted improvement potential before committing resources. 8. CRADLE-1 operates as a fully automated API or UI service—users input template sequences and assay data, receiving designed sequences within approximately two GPU-days of compute (hours wall-clock with parallelization), without requiring structural data or biochemical expertise. 📜Paper: biorxiv.org/content/10.648… #CRADLE1 #ProteinEngineering #LeadOptimization #ProteinLanguageModels #MachineLearning #DrugDiscovery #AntibodyDesign #EnzymeEngineering #CRISPR #VaccineDesign





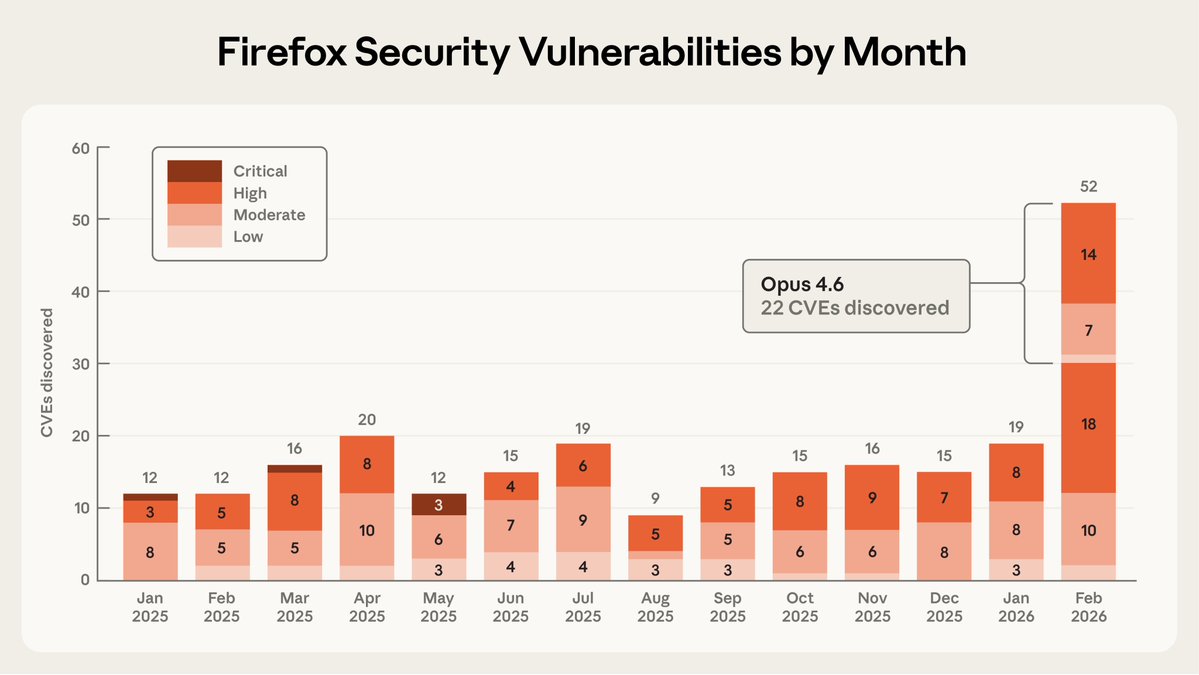
We partnered with Mozilla to test Claude's ability to find security vulnerabilities in Firefox. Opus 4.6 found 22 vulnerabilities in just two weeks. Of these, 14 were high-severity, representing a fifth of all high-severity bugs Mozilla remediated in 2025.