David Gilmore
737 posts

David Gilmore
@DavidGilmore3
Just a grumpy guy wit a twitter account. I've not picked up my guitar in years.
Katılım Aralık 2010
519 Takip Edilen175 Takipçiler

@ThinMatrix Who would have thought the old Equilinox movement math would be resurrected for the new beast!
English


@DavidGilmore3 Totally wrong. Global science leadership has more power than publishers - they just choose not to use it.
English

It's an absolute scandal and tragedy that India is forced to spend $250m/year to provide access to the latest science to its people. A complete and total failure of global science leadership and of science writ large.
Niko McCarty.@NikoMcCarty
Indian gov't is buying a subscription to 13,000 academic journals, and then making them all available to "18 million students, faculty, and researchers" for free. The cost is $715 million over 3 years. It includes Elsevier, Nature, and AAAS. Have any other countries done this?
English

St. Jude welcomes Georgios Skiniotis, PhD, to the Department of Structural Biology. He’ll also lead the new Center of Excellence for Structural Cell Biology, advancing technologies like cryo-ET and vEM to study cellular mechanisms. ow.ly/u4h750UcizY

English

@brianwut @peterrhague @BBCNews It's what Data says (from time to time) in Star Trek TNG - that's where SpaceX got it.
English

@peterrhague @BBCNews “Nominal” is American Spaceman English for “smashing”
English


@AdanBecerraPhD Why? Does history search backward work at last?
English

@WW3finalboss @Maks_NAFO_FELLA Comrade Stalin had the solution to this sort of problem #redarmyblues
English

@scorpio8675309 @PhysInHistory Haha - yes.
My response to the original post was going to be "no, it doesn't."
English

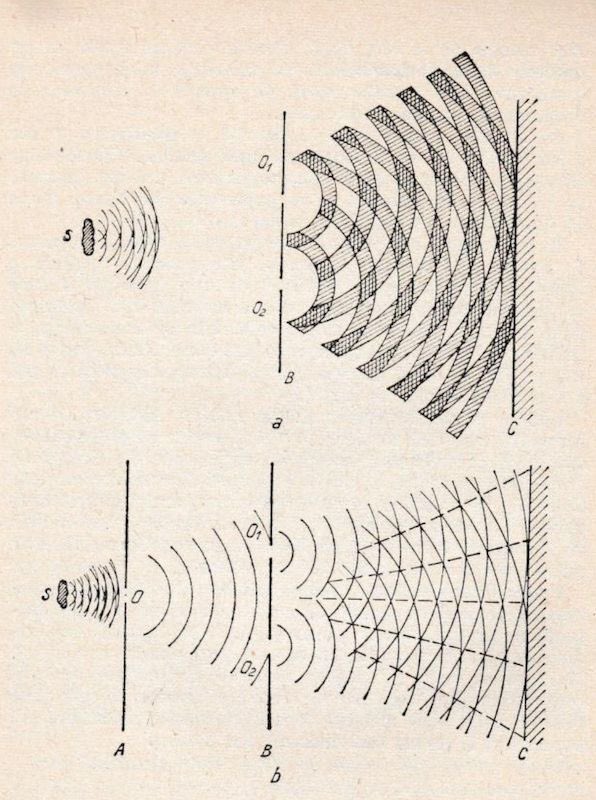
@PhysInHistory The double slit experiment demonstrates how a quantum field interaction produces a probability distribution that can be described by a mathematical wave function. The notion there is a wave-particle duality is as archaic as the geocentric model of the solar system.
English

@buildmodels Don't you just, like, wave a stick at it and pronounce "It Is So!"?
English

Interested in learning more about plans to extend PDB IDs to 12 characters?
Register for the November 7 Virtual Office Hour on Supporting Extended PDB IDs
rcsb.org/news/feature/6…
English

@dugu9sword You probably shouldn't call the results of cryo-EM "density maps" - that phrase can be used in casual conversation, but is not appropriate for a publication.
English

1/ 🔍 Wonder about the answer?
cryo-EM ✖️ Foundation Model 🟰 ❓
cryo-EM ✖️ Flow Matching 🟰 ❓
cryo-EM ✖️ Diffusion Transformer 🟰 ❓
Excited to introduce our new work--cryoFM, the first cryo-EM foundation model for protein densities with flow matching, which generalizes to four tasks without finetuning.
📃 Paper: arxiv.org/abs/2410.08631
🏠️ Project: bytedance.github.io/cryostar/cryof…
Joint work with Yilai Li @li_yilai , Jing Yuan @eugenejyuan , Quanquan Gu @QuanquanGu
English

@ELittlechild @TrekCulture Even more hateful than Gul Dukat.
English

Deep Space Nine update:
I REALLY REALLY REALLY DISLIKE KAI WINN!
#StarTrek #StarTrekDS9 @TrekCulture
GIF
English

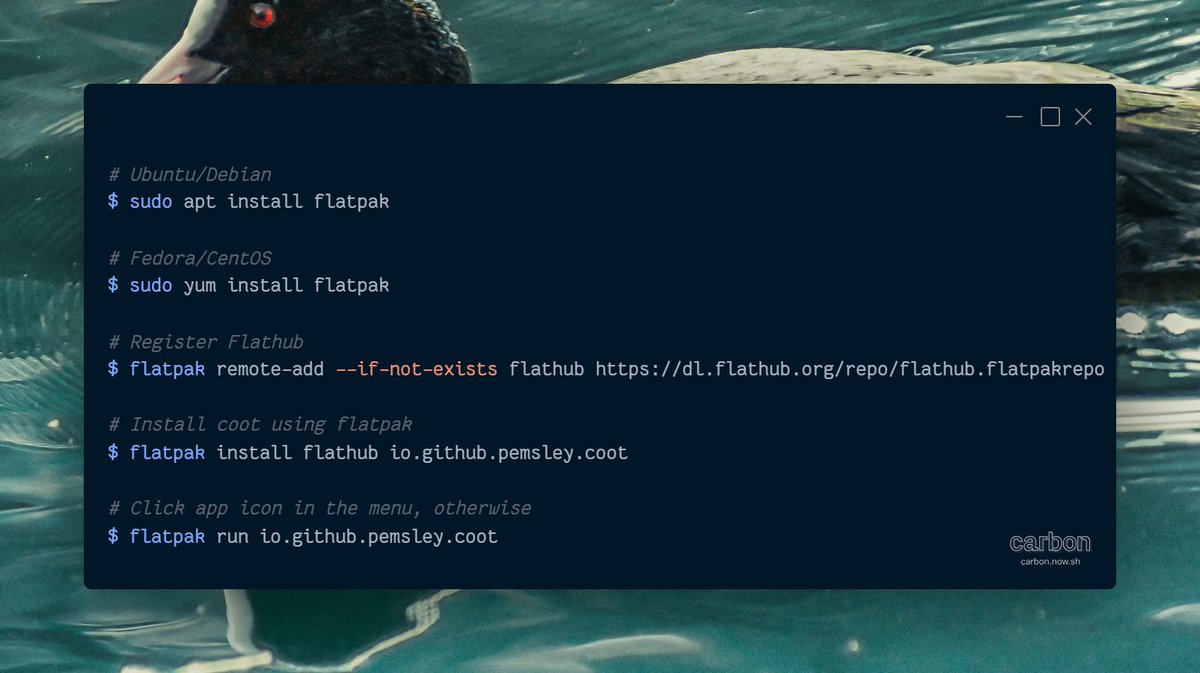
🦆 Coot can now be easily installed on Linux using Flatpak!
📦 Install and run it using the following command (It needs admin privileges to install Flatpak)
#coot
#bioinformatics
#flatpak
#linux
Yoshitaka Moriwaki@Ag_smith
🎉 New formula Coot 1.1.09 in Brewsci/bio for Linux and macOS! ℹ️ Crystallographic Object-Oriented Toolkit 🍺 brew install brewsci/bio/coot #Bioinformatics
English

@stargazermellu @McMasterCarr Have you seen an uptick in visitors?
English

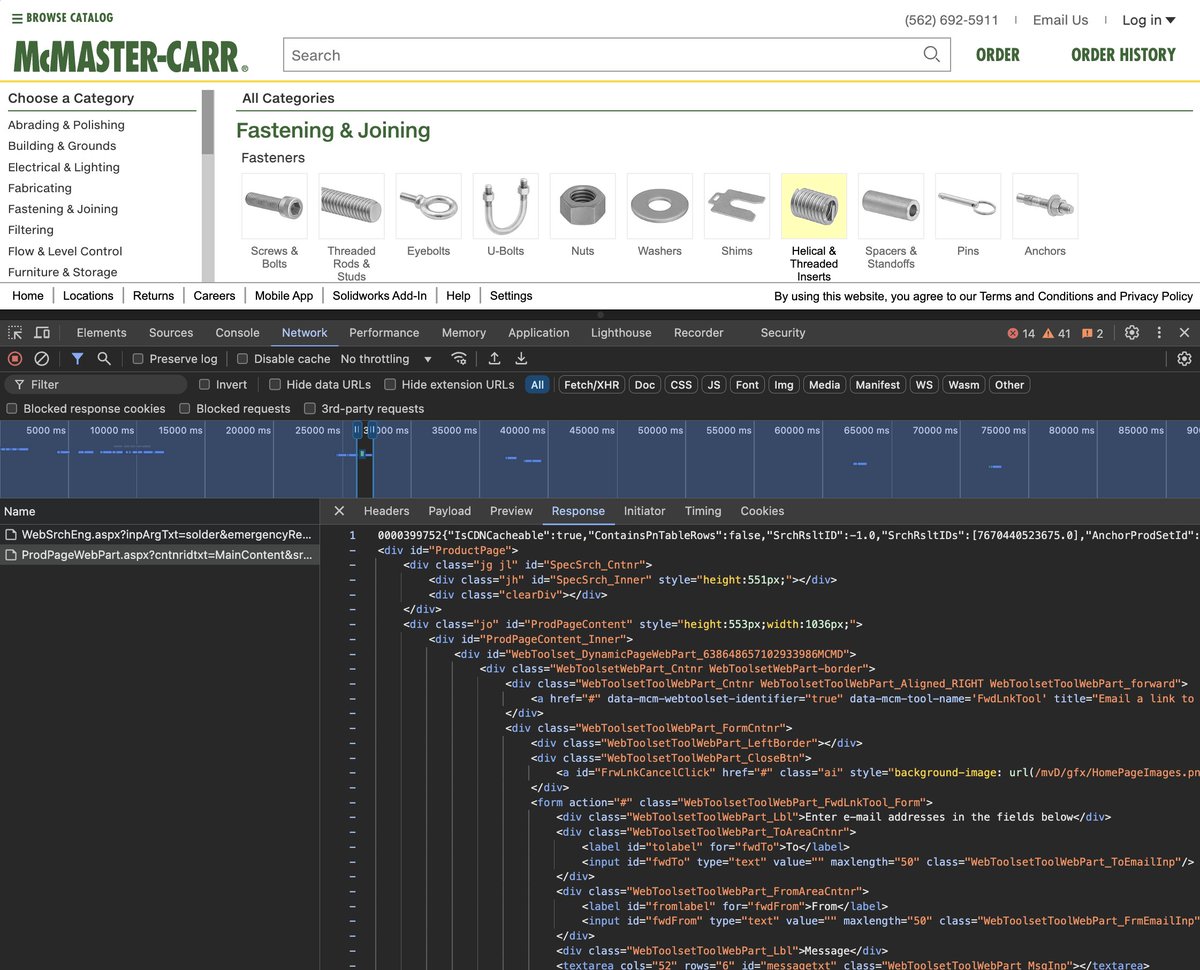
very cool stuff happening behind the scenes here! every time you hover over an link long enough (~1.5/10th of a second) it preemptively loads that page's code in the background. (raw HTML ready to go is also far less computationally intensive than say, json or graphql too!)
Kenneth Cassel@KennethCassel
how come a company founded over 100 years ago has the fastest site on the internet?
English

@Nature because he saw in solving the structure of hemoglobin the resemblance to myoglobin. He reasoned that if this was the case for these structures, it is likely for others too and to deny previous structures to aid elucidation of new ones would set the field back decades.
English

Much of the success of breakthroughs, such as AlphaFold, is thanks to a database of protein structures dreamed up in the 1960s by Helen Berman go.nature.com/4hp4UCP
English

@Nature Helen Berman didn't dream up a database. The PDB was started in 1971 by Walter Hamilton and team. Helen Berman became the director of the PDB in 1998 - 27 years later. It was Max Perutz who first pushed for a public archive of protein structures...
English

@teampulao @LKrauss1 It is not real. It is computer generated. It is based on some real observations.
English

Reality is so much more amazing than fiction.
Black Hole@konstructivizm
Europa & Io moons orbiting Jupiter, captured by the Cassini space probe
English