Eric J. M. Lang
85 posts


Eric J. M. Lang
@Eric_jm_lang
Computational chemist at @BristolChem with interests in protein dynamics, protein design, enzyme catalysis, allosteric regulation and drug disc. & devel.
Bristol, UK Katılım Şubat 2018
170 Takip Edilen116 Takipçiler

Excited to launch my group at @ETH_BSSE where we create photocatalytic enzymes☀️by computational design and directed evolution.🧬💻
Thanks, @snsf_ch for funding my #ambizione fellowship and @BPL_ethz for hosting me.👩🔬👨🔬
📢#PhD: Ad coming soon
📢#PostDoc: bristol.ac.uk/jobs/find/deta…

English
Eric J. M. Lang retweetledi

#HighlightOfTheWeek Peptide conformations for one force field and different generalized Born models for implicit solvation.
@AdrianMulholla1
#compchem
pubs.acs.org/doi/10.1021/ac…
English

@dgr8ashoka @Nature Congratulations Prasun, that's amazing!
English

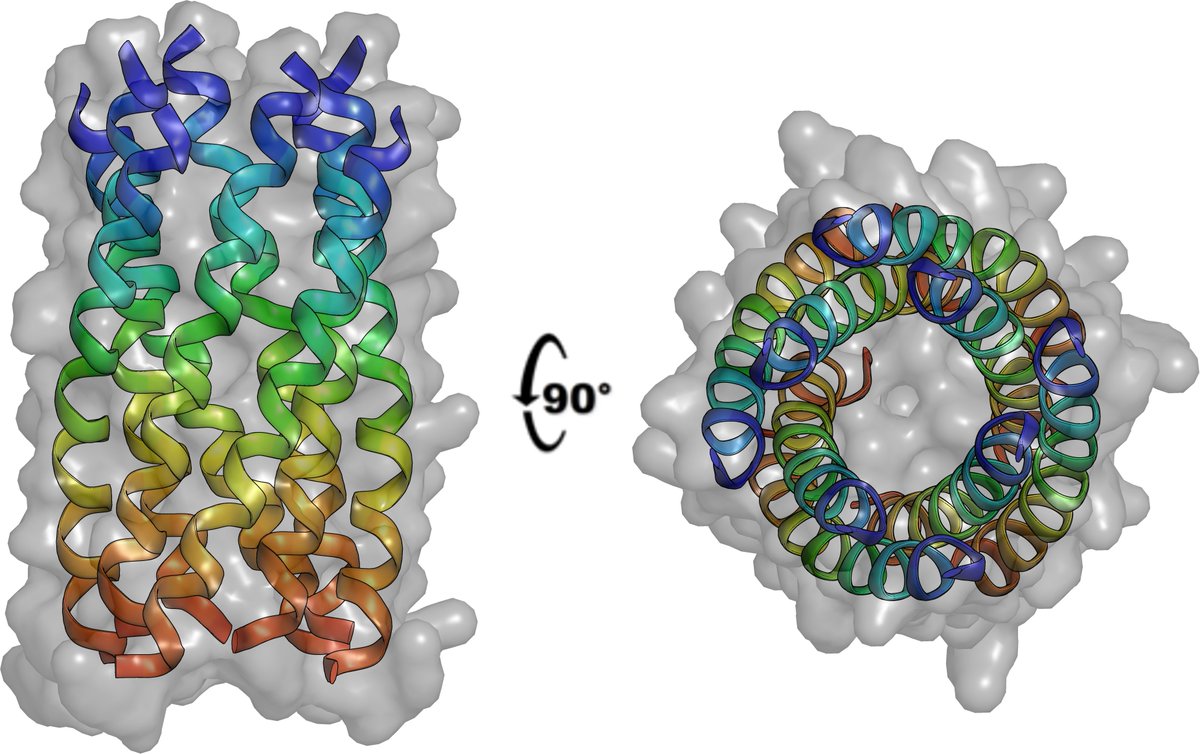
Truly a dark matter of protein structure space. Longest by far, and first assembly in solution and in crystal structure of 310-helices is out in @Nature.
nature.com/articles/s4158…
English

@Blendenfleck You can also pair it the the Hacker's Keyboard app to have access to all the keys you need in a standard layout 😉
English

@JCIM_ACS @sabahahmad_IN Reminds me of one of the swimming Aliens in Alien Resurection 🧐😅
English

#HighlightOfTheWeek Binding of a glycerophospholipid to the catalytic site of the phospholipase A1 PlaF using steered molecular dynamics simulations. Work by @sabahahmad_IN.
#compchem
pubs.acs.org/doi/10.1021/ac…
English
Eric J. M. Lang retweetledi

Check out this interesting study performed by @Eric_jm_lang and @AdrianMulholla1 with our collaboration 👇
Adrian Mulholland@AdrianMulholla1
Implicit solvent models fail to reproduce secondary structures of de novo designed peptides. 65 implicit solvent model/forcefields combinations; test set of designed α-helical #peptides >800 µs #MD @Eric_jm_lang Emily Baker @WoolfsonLab chemrxiv.org/engage/chemrxi…
English
Eric J. M. Lang retweetledi
The data show that implicit solvent models generally fail to reproduce the experimentally observed secondary structure content, and none performs well for all 5 peptides. The results show that these models are not usefully predictive. #CompChem #MD #ImplicitSolvent #Protein
English
Eric J. M. Lang retweetledi

Implicit solvent models fail to reproduce secondary structures of de novo designed peptides. 65 implicit solvent model/forcefields combinations; test set of designed α-helical #peptides >800 µs #MD @Eric_jm_lang Emily Baker @WoolfsonLab chemrxiv.org/engage/chemrxi…
Català
Eric J. M. Lang retweetledi
Eric J. M. Lang retweetledi

Congratulations to @TwidaleRebecca on her PhD viva! Thanks @matteodp & @marcvanderkamp for being examiners. Some of her work: Crystallography & QM/MM Identify Binding of Hydrolyzed Carbapenem & Penem Antibiotics to L1 Metallo-β-Lactamase in the Imine Form pubs.acs.org/doi/10.1021/ac…
English
Eric J. M. Lang retweetledi

Check out 👀 our #COVIDisAirborne preprint w/ tech aspects of our billion-atom simulation of Delta #SARSCoV2 in a respiratory aerosol, multiscale sims of spike opening @ltchong @arvindr_, pH @AdrianMulholla1 & calcium effects @tfmiller3 @EntosAI
tinyurl.com/c0v1d1sa1rb0rn3
English
Eric J. M. Lang retweetledi

An announcement I’ve been aching to make! After much sweat, we’ve built a trainable version of AlphaFold2, implemented in PyTorch, which we’re calling OpenFold.
GitHub: github.com/aqlaboratory/o…
Colab: colab.research.google.com/github/aqlabor…
Why a trainable version of AlphaFold2 you ask? ⬇️
English
Eric J. M. Lang retweetledi

Great stuff on #compchem #qmmm modelling workflow for zinc #enzymes from Zongfan, @TwidaleRebecca, @reynier_s, @CharlieColenso, @Eric_jm_lang @Spencer_D60, @AdrianMulholla1
@bcompb 💪
pubs.acs.org/doi/abs/10.102…
English
Eric J. M. Lang retweetledi

Evolution never ceases to amaze me! In our latest work, we show how directed evolution – tasked solely with improving activity – innovates a dynamical network organizing the whole protein in a reactive state primed for catalysis. nature.com/articles/s4155… 1/5

English
Eric J. M. Lang retweetledi

Check out our latest paper published in @NatureChemistry on our multi-step approach to design de novo peptides forming ion channels!
nature.com/articles/s4155…
English
Eric J. M. Lang retweetledi

Mechanism of covalent binding of ibrutinib to Bruton's tyrosine kinase revealed by QM/MM calculations - work of Angus Voice & @TwidaleRebecca @BristolChem @gtresadern @hermanvvlijmen @JanssenEMEA in Chemical Science #BTK #TCI #EGFR #QMMM #CovalentInhibitor pubs.rsc.org/en/content/art…
English
Eric J. M. Lang retweetledi

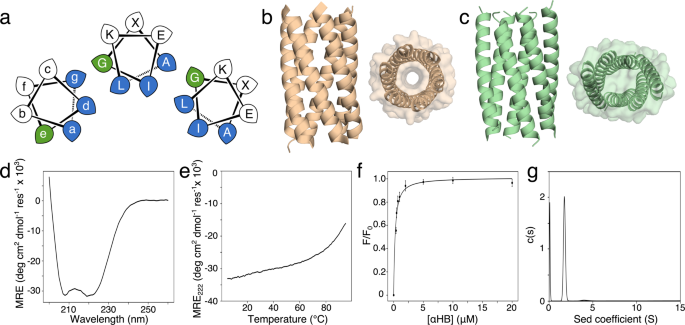
New work from Dek Woolfson’s lab: Structural resolution of switchable states of a de novo peptide assembly WoolfsonLab @BristolChem @BrisBioDesign @BristolBiochem @Will_M_Dawson @Eric_jm_lang @Guto_Rhys @AdrianMulholla1
go.nature.com/3t5eG2H
English

@Eric_jm_lang @NatureComms @WoolfsonLab @AdrianMulholla1 @BristolChem @BristolBiochem Congratulations Eric, very nice work!!!
English

Our latest paper: “Structural resolution of switchable states of a de novo peptide assembly” is in @NatureComms: nature.com/articles/s4146…. Collaboration between @WoolfsonLab and @AdrianMulholla1's group. @BristolChem @BristolBiochem #compchem.
English

Multiple MD simulations were run on the GPUs of @BristolUni_ACRC (the HPC systems of @BristolUni), using @ambermdprog. The resulting 138 μs of simulations were then analyzed using the InfleCS program, developed by Annie Westerlund @DelemotteLab.
English

Sharing co-first authorship with the amazing @Will_M_Dawson, @guto_rhys and Kathryn Shelley!
English
