Genevieve Roberts
42 posts

Genevieve Roberts
@GenRoberts_PhD
Salt Lake City, UT Katılım Şubat 2019
163 Takip Edilen102 Takipçiler
Genevieve Roberts retweetledi

Our latest in @Nature:
Convergence of coronary artery disease genes on endothelial cell programs
nature.com/articles/s4158…
Incredible work led by Gavin Schnitzler and @kanghelenyihua — a long-standing partnership between our lab and @Dr_RajatGupta
My comments below:
1/
English

@willboneferroni Congratulations Will! Couldn't agree more with your colleagues' account of working with you!
English
Genevieve Roberts retweetledi

This sounds like a not-great aspect of using CRISPR in humans - what do the gene editing folks think about this finding, in terms of disease therapy?
Chris Gibson@RecursionChris
Others have previously reported evidence of somewhat similar bias, but on a smaller scale in specific cell types. This is the first time we’ve seen whole genome evidence in both cancer and somatic cells, suggesting this is a general feature of CRISPR-Cas9 editing.
English
Genevieve Roberts retweetledi

Tweetorial time! We @RecursionPharma mapped consequences of #CRISPR screening of >17K human genes, found a systematic bias confounding all CRISPR screens, traced its molecular cause, and propose a debiasing algorithm.
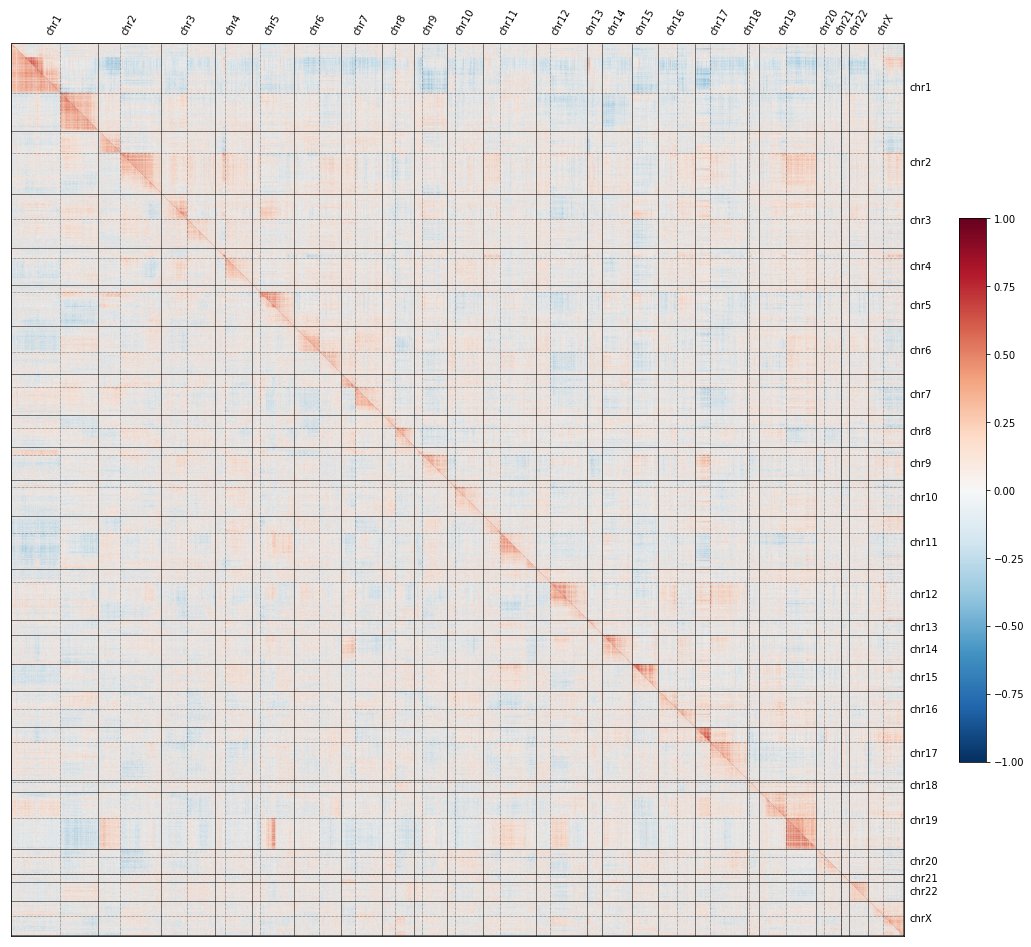
English

@MarieCoignet Thank you for all of your amazing work here...especially on the surveys!! Couldn't have pulled this off without you @MarieCoignet !!
English

Extremely proud that our AncestryDNA COVID-19 paper is out in Nature Genetics! So grateful to the entire team of people that got this across the finish line, with special thanks to Kristin Rand, Eurie Hong, Brooke Rhead and Raghav…lnkd.in/gXfKAxmQ lnkd.in/gB_Ty2-i
English
Genevieve Roberts retweetledi

Congrats to our @Ancestry collaborators for new findings in @NatureGenet on genetic association with COVID-19 susceptibility & severity. Our collaboration has enabled RGC & Ancestry to share vital learnings with the global scientific community. go.nature.com/3M0BHgI
English

Extremely proud that our AncestryDNA paper is out in @NatureGenet!
This COVID-19 GWAS paper explores the "typical" COVID phenotypes (e.g. hospitalization) + 4 phenotypes that are hard to collect in BioBanks (eg. symptom severity) with self-reported data
nature.com/articles/s4158…
English
Genevieve Roberts retweetledi

Online now: Expanded COVID-19 phenotype definitions reveal distinct patterns of genetic association and protective effects (Roberts et al.) go.nature.com/3vbYTBQ
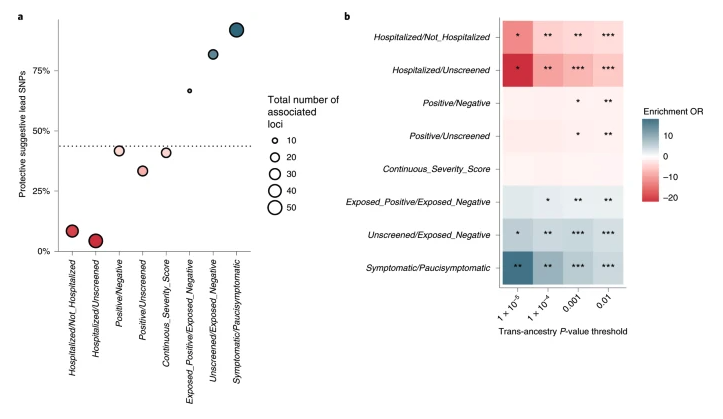
English
Genevieve Roberts retweetledi

Join us tomorrow for our @lmrl_bio talk at #NeurIPS2021, where we’ll share how our Recursion OS platform and maps of human biology are rapidly accelerating drug discovery. Interested in joining our team? We’re hiring! Apply today: boards.greenhouse.io/recursionpharm…

English

Check out our NeurIPS 2021 talk for an in depth look on what we're doing at Recursion to decode biology and radically improve lives. lnkd.in/gAZ_iDcf
English

@jgschraiber *sigh* The clotting disorder is highly unusual, so it makes sense to figure out why it happens so it can be clinically managed appropriately (e.g. no heparin), but unfortunate that it's gotten so much media attention and will unnecessarily create more hesitancy.
English

@ShaiCarmi Interesting! I looked at the LD patterns (pic). The arrows show the SNPs we report and the rest are from the paper you linked. One of our signals tags the one from that paper, but our other two signals appear pretty independent.
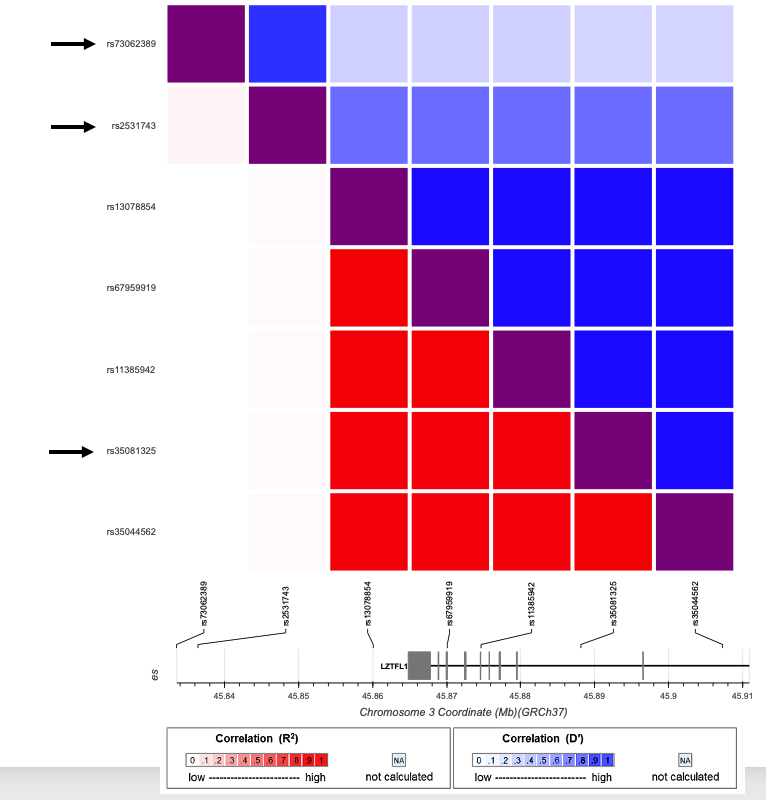
English

@GenRoberts_PhD By the way, there was some analysis of the chr3 haplotypes here: biorxiv.org/content/10.110…
It would be really interesting to dig further into the haplotype structure (though probably beyond the current paper I guess).
English

Very excited to announce that the @Ancestry Science Team just uploaded a new COVID-19 GWAS pre-print with larger sample sizes and an expanded set of 8 phenotype definitions.
medrxiv.org/content/10.110…
English

@ShaiCarmi 2) The 3 SNPs in the chr3 locus are not in LD (r2<0.05), so we didn't consider a haplotype analysis, but I will look again at D'. Thanks again!
English

@GenRoberts_PhD Very nice work! Two quick comments if I may.
1) In the replication analysis of the known SNPs - I think maybe better to apply Bonferroni correction (ie 0.05/13).
2) The 3 associated SNPs on chr3 - are they linked? How many risk/protective haplotypes are there?
Thanks.
English

@ShaiCarmi 1) We considered Bonferroni, but it's a bit conservative & many replication studies use a more relaxed 0.05 even for multiple SNPs. Our major conclusions hold if you apply BF, but we may add a * to show which associations surpass P<0.05/13 in Fig 2. Thanks for the suggestion!
English

@genandgenes I certainly agree that loci discovered with population controls require increased scrutiny to be sure they are not due to SES confounding, but many of the loci discovered with very large population control GWAS do replicate in follow-up studies that do not use population controls
English

It's imp to be clear re: what we are testing with pop controls when not everyone has the same exposure levels. What is the counterfactual?
When I see these results, it's a red flag that we're measuring who was vs wasn't able to stay home. Not nec biology specific to #Covid_19.
Abdel Abdellaoui@dr_appie
The estimated shared genetic effects between several Covid-19 outcomes and a wide range of heritable traits. (the estimated genetic effects for the Covid-19 outcomes come from the January 2021 release of @covid19_hgi)
English

@MGuru2020 @Ancestry We found that our novel phenotype definitions overwhelmingly identified suggestive *protective* associations, but we remain cautious about which of these will replicate as sample sizes grow larger.
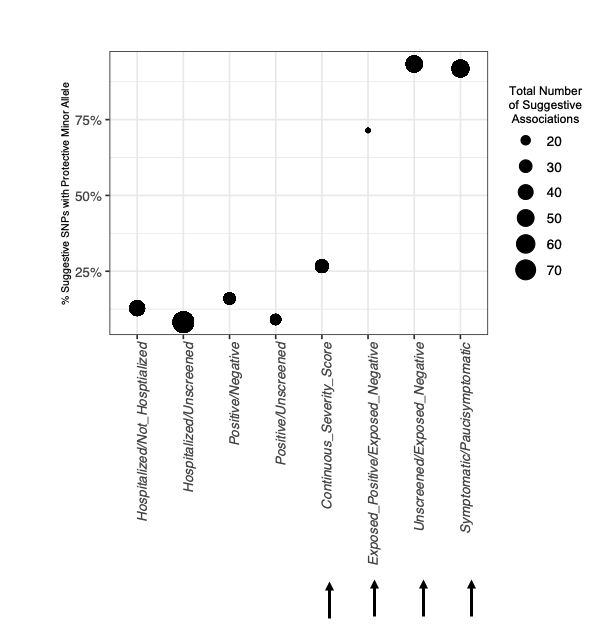
English


