

HMSBioPlex
228 posts

@HMSBioPlex
We are on a quest to map the human interactome via AP-MS. Check back here for updates on our progress! Steve Gygi, Wade Harper & Ed Huttlin HMS Cell Biology





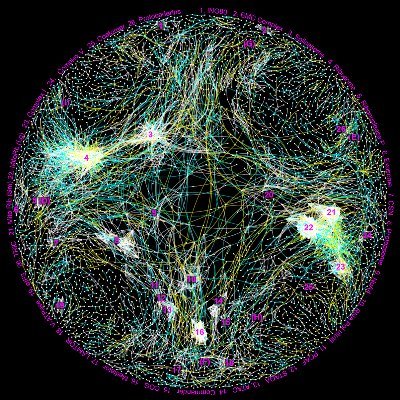
In a Nature article researchers from HPA and UCLA have used a machine learning approach that enables computational integration of protein localization data from IF images and AP-MS protein interaction data to create a MuSIC map of 5,100 proteins in U2-OS. proteinatlas.org/news/2025-05-2…

🚨 New in Nature @Nature! 🚨 Thrilled to be a co-first author on a global cell map built from U2OS protein imaging + interaction data 🧬🌐 📝 Paper: nature.com/articles/s4158… 🗺️ Visualization portal: musicmaps.ai/u2os-cellmap/

🚨 New in Nature @Nature! 🚨 Thrilled to be a co-first author on a global cell map built from U2OS protein imaging + interaction data 🧬🌐 📝 Paper: nature.com/articles/s4158… 🗺️ Visualization portal: musicmaps.ai/u2os-cellmap/





Check out this blog post and paper from Alberto Cruz-Martín, @rhushikeshphadk, and colleagues describing how @HMSBioPlex helped them link the complement pathway to synaptic dysregulation! communities.springernature.com/posts/beyond-p… rdcu.be/dSTkf








@JohnRYatesIII @GygiLab @harvardmed @the_prbb @UPFbiomed @CRGenomica And he made such an amazing talk! Many thanks Steve (@GygiLab) for coming and visiting us. It's been a pleasure to host you in Barcelona! #proteomics #bioplex #AlphaFold @the_prbb @CRGenomica @UPFbiomed @harvardmed

This Friday we are hosting Steve Gygi (@GygiLab) from @harvardmed at @the_prbb Auditorium in #Barcelona. What a multiplexed excitement! 🎉 Come and join us. @UPFbiomed @CRGenomica #proteomics



