Debbi Hamlin
132.9K posts



It is now possible to directly read and locate precise gene-editing events inside of intact tissue at single-cell resolution.
TL;DR: You can deliver a base editor to the liver and then figure out where it made edits across all the cells in the organ.
Before now, there were two main ways to quantify gene-editing: Bulky and tedious.
For the “bulk” approach, you inject a mouse with an AAV carrying your base-editing payload, say, and then kill that mouse later. You cut out its liver (or brain, or whatever), put it in a blender, and sequence all the DNA therein. This gives the AVERAGE fraction of edited alleles in the organ, but removes spatial information.
The tedious approach is microdissections. Instead of dropping the liver into a blender, you cut it into ultra-thin slices and then use a laser to cut out individual cells. By sequencing just those cells, you can figure out which got the genetic edits. So tedious!
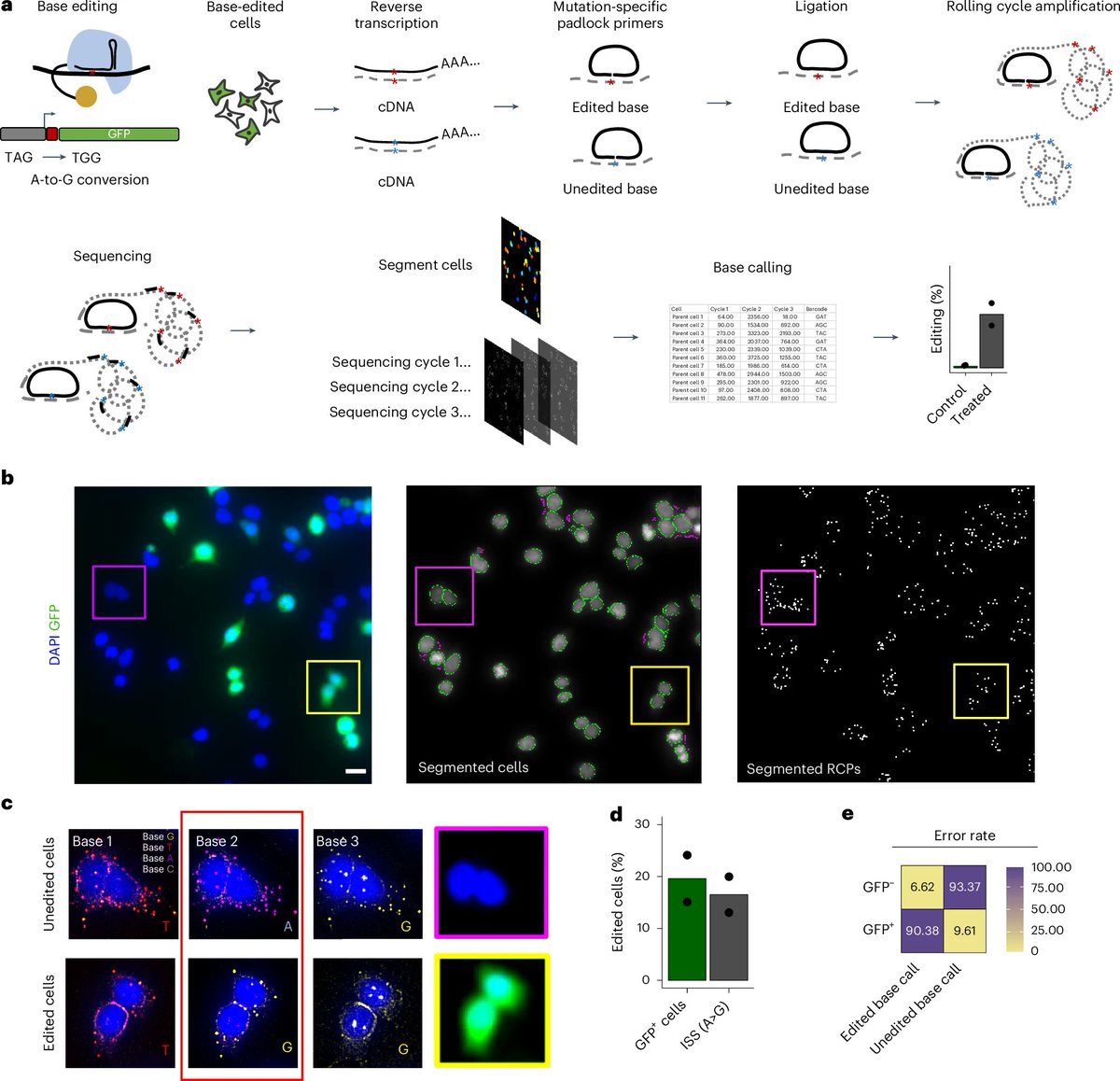
A new paper in Nature Biomedical Engineering solves this problem using a technique called in situ sequencing, or ISS. This is a way to “read” genetic edits in single cells without ever cutting out those cells.
First, they take the tissue and fix it on a slide, so the cells stay in place. Next, they wash these cells with a short strand of DNA, designed to bind exactly where the gene-editing tool cuts. Attached to that DNA strand is a special probe, called a padlock. This padlock is designed, at the molecular level, to CLOSE only if the short DNA sequence exactly matches the gene-editing location. When this probe closes — only in edited cells — it becomes circular.
Next, the scientists “amplify” only the circular DNA (again, INSIDE the cells) and sequence them using a miniature version of sequencing-by-synthesis chemistry. The final result is that you can figure out which alleles got edited in which cells, across the entire organ.
In mouse brains, these scientists saw higher editing in neurons than astrocytes, for example. And when they injected macaques with lipid nanoparticles carrying base editors, they found that editing was surprisingly uniform across the entire organ (a good sign that the technology "could be used to address a wide spectrum of metabolic liver diseases," as the authors write.)
Another important finding (in mice and macaques) is this: "...in experiments where repeated doses of RNA-LNP are administered, the initial dose does not affect the editing efficiency and distribution of the subsequent dose."
This is a nice paper, and well worth reading. It will be good to have more rigorous methods for quantifying gene-editing efficiencies, because these metrics have been so scattered from one paper to the next. It has been really difficult to come up with high-quality benchmarks.

English
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi

Debbi Hamlin retweetledi

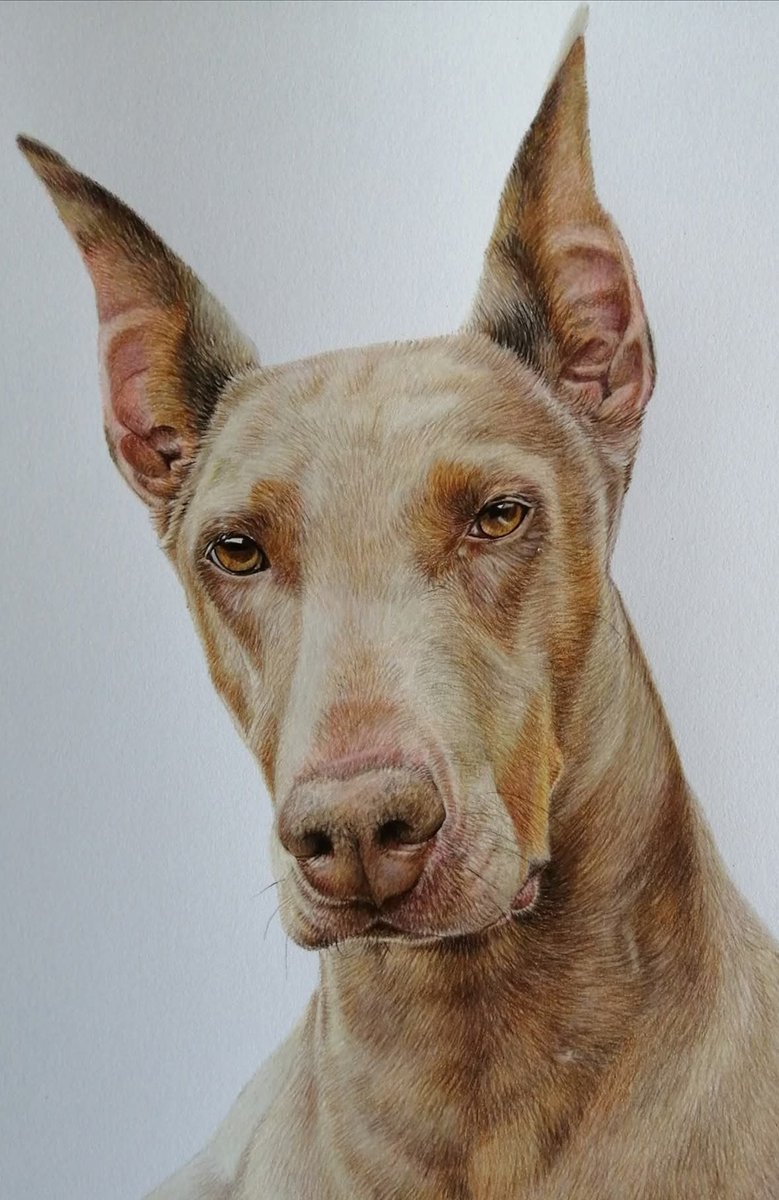
Princess Esmeralda.
Coloured pencil. Size 12x16".
Book a commission at suehewsonartist.weebly.com
#drawing #art #petportraits

English
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi

Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi

@UndisScot I usually stop at the bridge to get some aerial shots but I went a wee bit up the road to get this shot. So glad I did.
It’s a SPECTACULAR place - x.com/weejonny1/stat…
Jonny Campbell 🏴@weejonny1
Kylesku 💙🏴💙 Music - Julia Fowlis -Do Chalum
English
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi

Coyote trotting along the shore as ravens and herons watch him 😄
Please watch my latest YouTube video - youtu.be/FwODLSN8kOY?si…
Thank you, Kate 🥰❤️🇨🇦
Delta BC 🇨🇦 12 Sept 2025

YouTube
English
Debbi Hamlin retweetledi

Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi
Debbi Hamlin retweetledi




















