Sabitlenmiş Tweet

Want to ask a #CryoEM #TeamTomo or #ElectronMicroscopy question anonymously? Send an email to imagingartifact@gmail.com to have it shared from this account.
English
Imaging_Artifact
3.3K posts


@ImagingArtifact
Sharing the good and the bad of electron microscopy and related ❄️🔬Ice crystal hall of fame 🧊🏆 #ElectronMicroscopy #CryoEM #TeamTomo


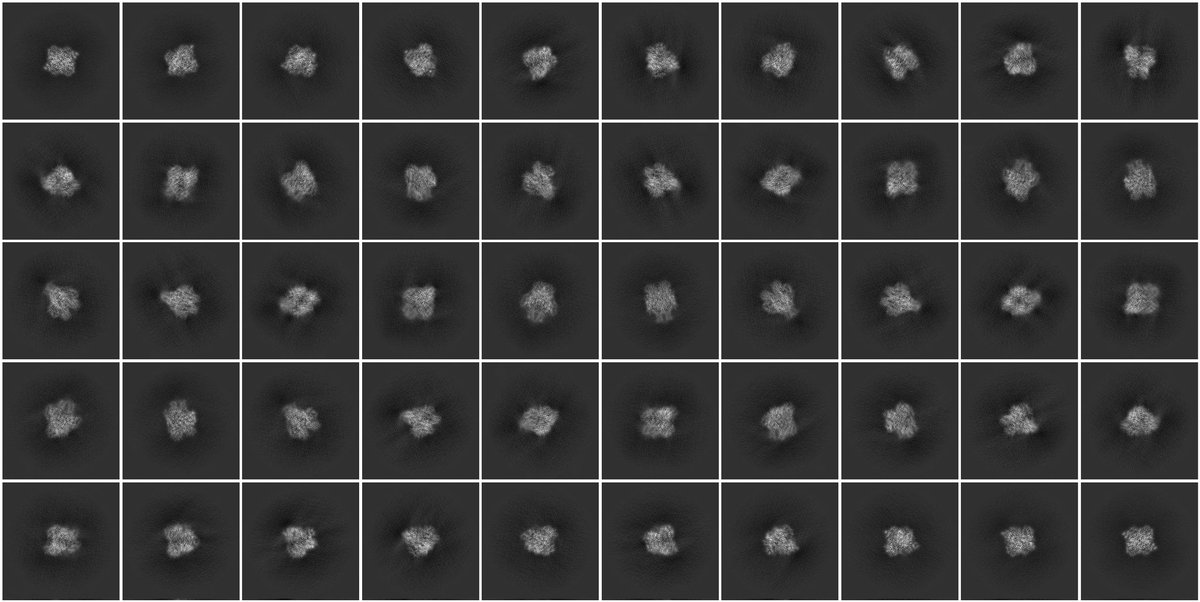










@ChavesSanjuan @ImagingArtifact I also changed the strains of E. coli to get rid of the contaminants but often it still brings other ones (see screenshot) especially if the protein of interest isn’t well behaved and has multiple domain with loops!















