Lynn
1.2K posts


Lynn
@Lynn_evodevo
Postodoc @darwintreelife @sangerinstitute Working with @bat1kgenomes Affiliated @TrinCollCam|Previous postdoc @Chema_MD lab @QMUL|EvoDevo & Genomics Biologist
Cambridge, England Katılım Temmuz 2017
4.3K Takip Edilen403 Takipçiler
Lynn retweetledi

Thrilled to share our new paper in @emboreports, now online! We explored transcriptomic evolution during early embryogenesis of two spiralian annelids, to understand how early cell fate modes shape development before the gastrulation. #EvoDevo #Embryogenesis #RegulatoryEvolution
English

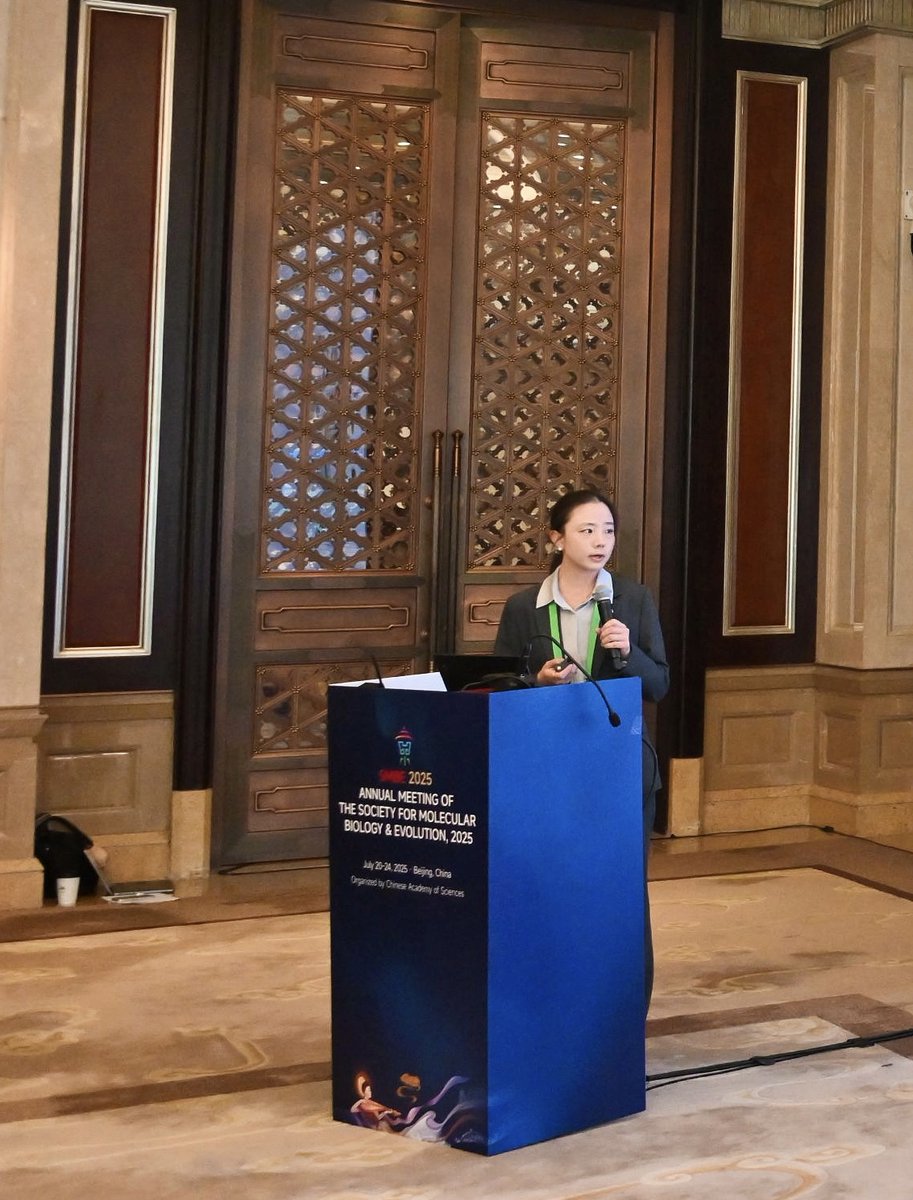
Thrilled to attend my first SMBE annual meeting in Beijing at the end of July, I am honoured to present our work on Bat #Phylogenomics, and (unexpectedly) to be the last speaker of our session on the very last day.
Thanks to the organisers and everyone who joined! #Bat1K #Sanger




English

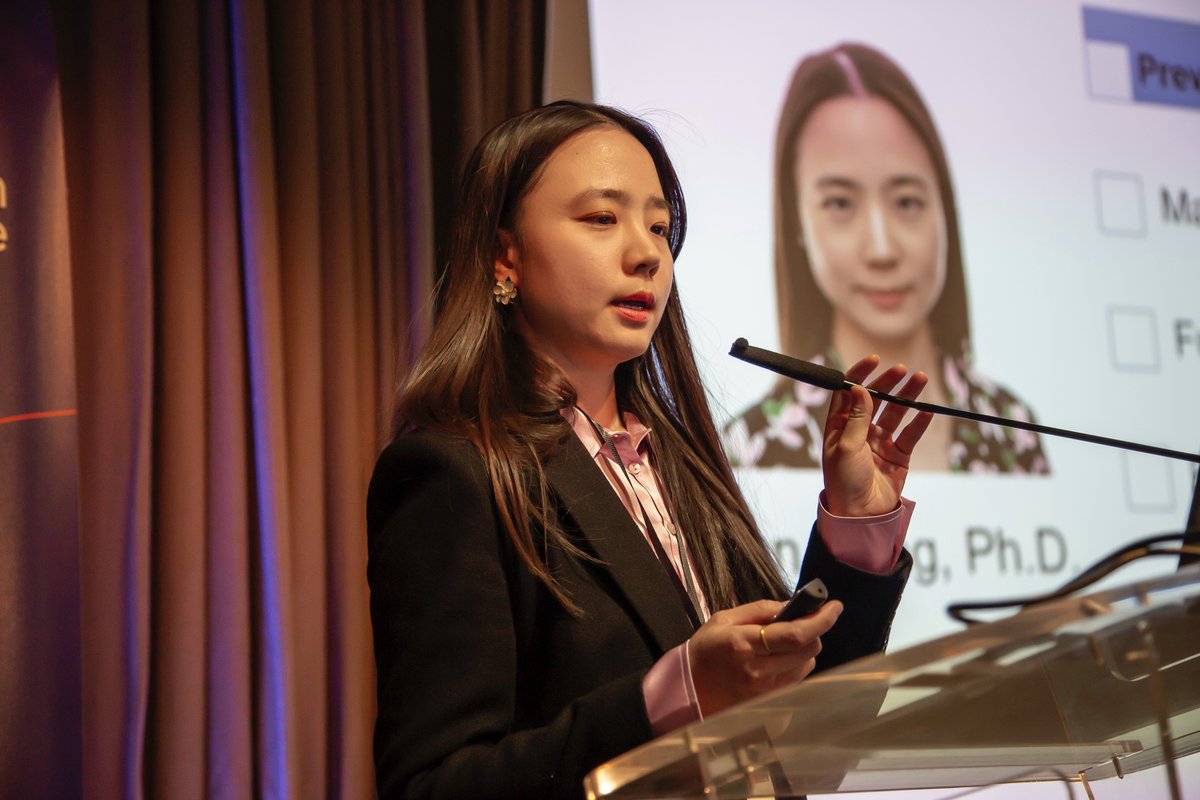
I gave a 3-minute flash talk @sangerinstitute #ScienceDay last week, sharing my work with @bat1kgenomes consortium on bat phylogeny.
Grateful to the organisers and community!
So glad the open ending resonated with many in the genomics crowd!
#Phylogenomics #WomenInSTEM 🦇

English

Truly honoured to have attended the EMBO #EvoDevoTempo workshop last month in Paris.
It was such a pleasure to present the works with @Chema_MD lab.
Grateful for the inspiring talks and the chance to connect with so many brilliant scientists.
#EvoDevo #AcademicLife #WomenInSTEM


English
Lynn retweetledi

🚀 Our perspective is out in @Nature!
We present a roadmap for Multimodal Foundation Models (MFMs) — large AI models pretrained across multi-omics and multi-timepoint data — to serve as the computational backbone for building virtual cells.
Read the full paper in Nature: nature.com/articles/s4158…
🔍 Why MFMs?
Biology is inherently multimodal, and molecular layers are deeply interconnected and context-specific. MFMs aims to integrate these layers to uncover shared biological principles that govern diverse cell states, offering a unified substrate for downstream inference.
🧠 What’s new?
💡 From hypothesis-driven to data-centric workflows: MFMs shift biology’s paradigm. Instead of crafting bespoke models for narrow tasks, we can now pretrain over massive datasets, distill foundational knowledge, and refine insights through lab-in-the-loop experimentation—where models guide experiments, and experiments update models.
🧬 Conditional gene regulation: MFMs go beyond static models. By training across multiple omics layers (e.g., chromatin accessibility, transcriptomics), they can learn context-specific gene functions and regulatory programs—key to understanding development and disease.
🧪 In silico perturbation: Biology’s combinatorial complexity is immense—thousands of genes, millions of interactions. MFMs provide a framework to simulate perturbations before wet-lab execution. Trained on CRISPR perturb-seq data, they can predict molecular responses across cell types, tissues, and time—enabling programmable biology at scale.
⚙️ What makes MFMs possible?
Envisioned techniques include:
- Unified tokenization from nucleotides to pathways
- Hybrid attention across intra- and inter-modal interactions
- Prompt-driven multitasking for temporal prediction, conditional generation, and modality translation
- Human knowledge integration from curated databases and biomedical literature
These design principles translate the architecture of foundation models into the molecular domain.
⚠️ What are the challenges?
MFMs aren’t just about scale—they demand accessibility, reliability, and transparency.
- Low-resource learning techniques (e.g., LoRA, adapters) are vital for democratizing training
- Human-agnostic benchmarks are needed, as conventional labels may punish models that uncover novel biology
- Uncertainty modeling is essential to mitigate hallucinations and increase scientific trust
Interpretability and ethical stewardship must be foundational in this emerging ecosystem.
Kudos to all co-authors for the collective effort and vision: @HOATIANCUI1, @Alejandro__TL, @mariabrbic, @JulioSaezRod, @simocristea, @genophoria, @mo_lotfollahi, @fabian_theis.
Let’s build the future of virtual cells together.




English

Happy New Year! 🎉 Excited to start 2025 by sharing our latest study from @Chema_MD lab! 🚨 #EvoDevo #EarlyEmbryogenesis
Cell fate specification modes shape transcriptome evolution in the highly conserved spiral cleavage biorxiv.org/content/10.110…
English

In Memoriam: Professor Wen-Ying Yin (1922–2023) | Zootaxa mapress.com/zt/article/vie…
English
Lynn retweetledi

BREAKING NEWS
The 2024 #NobelPrize in Physiology or Medicine has been awarded to Victor Ambros and Gary Ruvkun for the discovery of microRNA and its role in post-transcriptional gene regulation.

English

Revisiting the four Hexapoda classes: Protura as the sister group to all other hexapods | PNAS pnas.org/doi/10.1073/pn…
English

How we learned to build a gliding mammal thenode.biologists.com/how-we-learned… via @the_Node
English

@Ella_Maru Yes, my current model is very speculative based on our results; many more explorations are needed in the future.
English

@Lynn_Yan_Liang Thanks for the response, Lynn! It's definitely an interesting area to explore. Hopefully, new genetic tools will make it possible in the future.
English

The paper from my first PhD project on “The functional evolution of collembolan Ubx on the regulation of abdominal appendage formation” is now online! #EvoDevo #Collembola
Read it here: link.springer.com/article/10.100…
English

@Ella_Maru Thanks for your interest! It is a pity we cannot investigate the post translational regulation of Ubx “during late embryogenesis” due to the lack of genetic tools; but I agree it could be worthy to explore whether those regulation would alter Ubx’s function temporally.
English

@Lynn_Yan_Liang Congratulations! Have you investigated post-transcriptional regulation of Ubx and Dll during late embryogenesis?
English