
Naail Kashif-Khan
532 posts


@NKhan212
Making proteins in the lab and with AI 🔬💻 Wielder of petri dish and keyboard 🧫⌨️ Love heavy metal and the Arsenal 🎸🔴








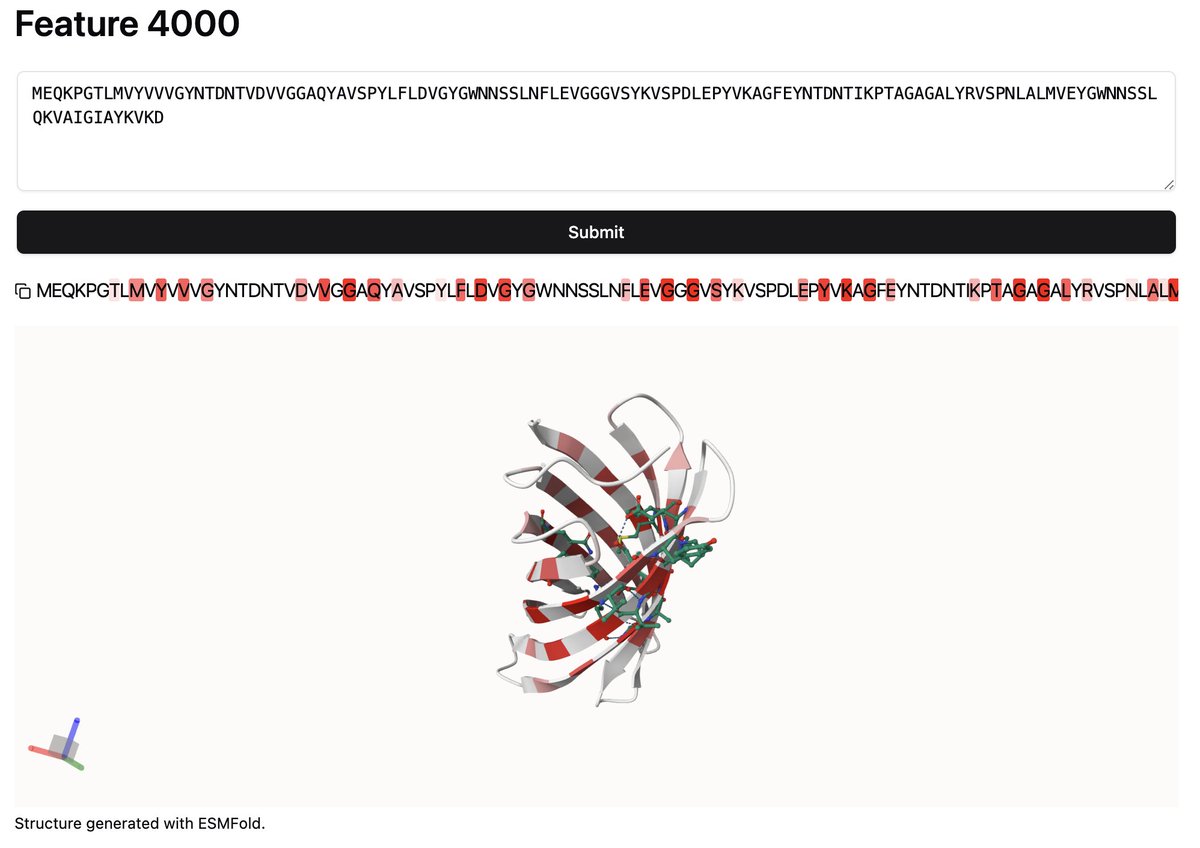

Ever wondered how a protein language model sees your favorite protein? Checkout out our SAE visualizer where you can now search any sequence for activating features.


ProtSCAPE: Mapping the landscape of protein conformations in molecular dynamics 1. ProtSCAPE is a deep learning architecture designed to map protein conformations from molecular dynamics (MD) simulations, using a novel combination of learnable geometric scattering with dual attention mechanisms. 2. The model employs geometric scattering to capture both local and global protein structures, representing proteins as graphs, which are then processed by a transformer with dual attention—focusing on residues and amino acids. 3. ProtSCAPE’s latent representations are temporally coherent, allowing it to capture conformational transitions, such as phase changes between open and closed states, and stochastic switching between meta-stable conformations. 4. Unlike conventional MD trajectory analysis that often misses complex transitions, ProtSCAPE excels in generating detailed low-dimensional representations that retain structural and temporal context, enabling enhanced visualization and downstream analysis of protein dynamics. 5. ProtSCAPE effectively generalizes from short to long trajectories and from wild-type to mutant proteins, offering insights into how mutations can affect the protein conformational landscape. 6. The model can interpolate between states to reconstruct intermediate conformations, validated with case studies on proteins like MurD, which showed hinge-like transitions consistent with experimental data. 7. ProtSCAPE outperformed traditional graph-based methods (GNNs) in predicting pairwise distances and dihedral angles, demonstrating its superior ability to decode the dynamics of protein conformations. 8. This tool holds significant promise for studying complex protein functions, such as allostery, binding, and enzymatic catalysis, by providing a comprehensive view of protein motion across various temporal scales. @egbertcastro @dbhaskar92 @KrishnaswamyLab @Siddharth2814 💻Code: github.com/KrishnaswamyLa… 📜Paper: arxiv.org/abs/2410.20317 #ProteinDynamics #DeepLearning #Bioinformatics #MolecularDynamics #Transformer #ProteinConformation #MachineLearning #ComputationalBiology



@Ginkgo’s protein language model, built on @googlecloud technology, offers unprecedented insights for researchers, accelerating the development of life-saving medicines. #DrugDevelopment #ProteinLLM #AIinBiotech #GinkgoBioworks #GoogleCloud loom.ly/RGh0ORc

AlphaFold3 reproduced and params/code released. 🤩












