

OMA Database
172 posts
@OMABrowser
Orthologs among 2000+ complete genomes across all of life. Follow us to stay informed of new releases and features. We welcome feedback and suggestions!







After trying many different ideas over years, we're happy to have found a scalable solution without sacrificing accuracy. It works by clustering homologs using k-mer based placement, followed by a highly parallelised taxonomy-guided processing of families using subsampling.(3/4)





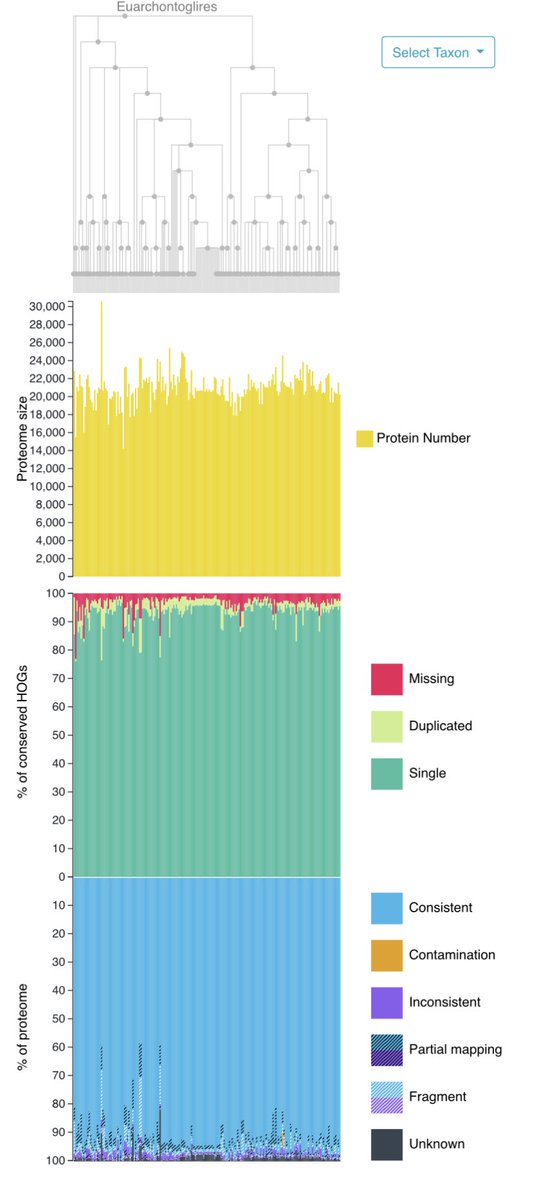







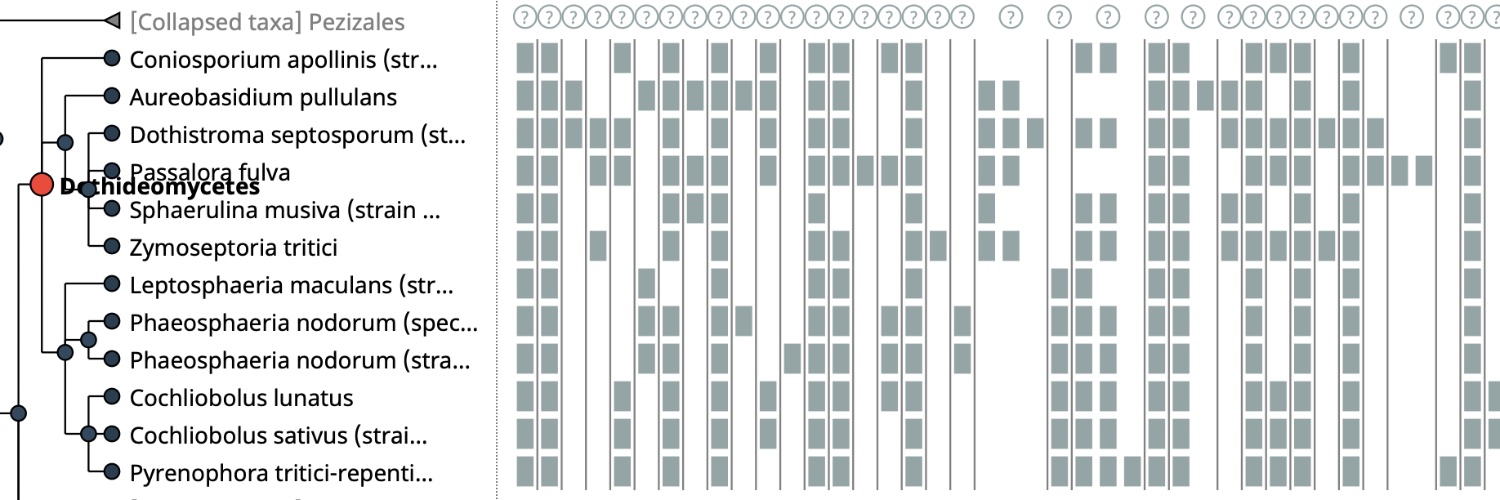
@KamilSJaron @rmwaterhouse @nuala_oleary @giulio_formenti @CibeleCaio @MahUliano @Why_NeverS @SinaMajidian @CorreardSolenne @Toby_Baril_Bio @EBPgenome @SangerVGP @SangerToL @genomeark @erga_biodiv @BioGenEurope @CatBiogenoma @DAISEA_AfricaBP New topics include: ✅ Starting a comparative genome study from CNGBdb: datasets, analysis platform, and spatial transcriptomics database ✅ Genome profiling using Genomescope ✅ OMA and OMArk for homology exploration and gene annotation quality control