
Today we celebrate @PascalSturmfels on a successful thesis defense! Photo by @Hannahlhan
Pascal Sturmfels
127 posts


@PascalSturmfels
PhD student at @uwcse + @UWproteindesign working on machine learning for protein design, protein language models, etc.

Today we celebrate @PascalSturmfels on a successful thesis defense! Photo by @Hannahlhan


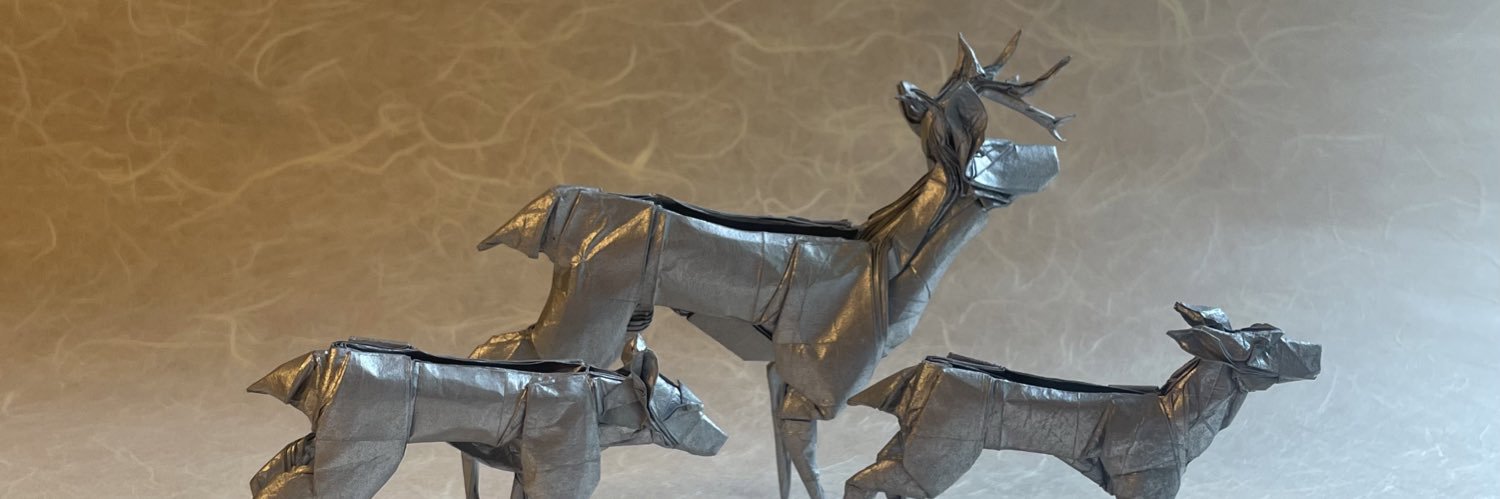
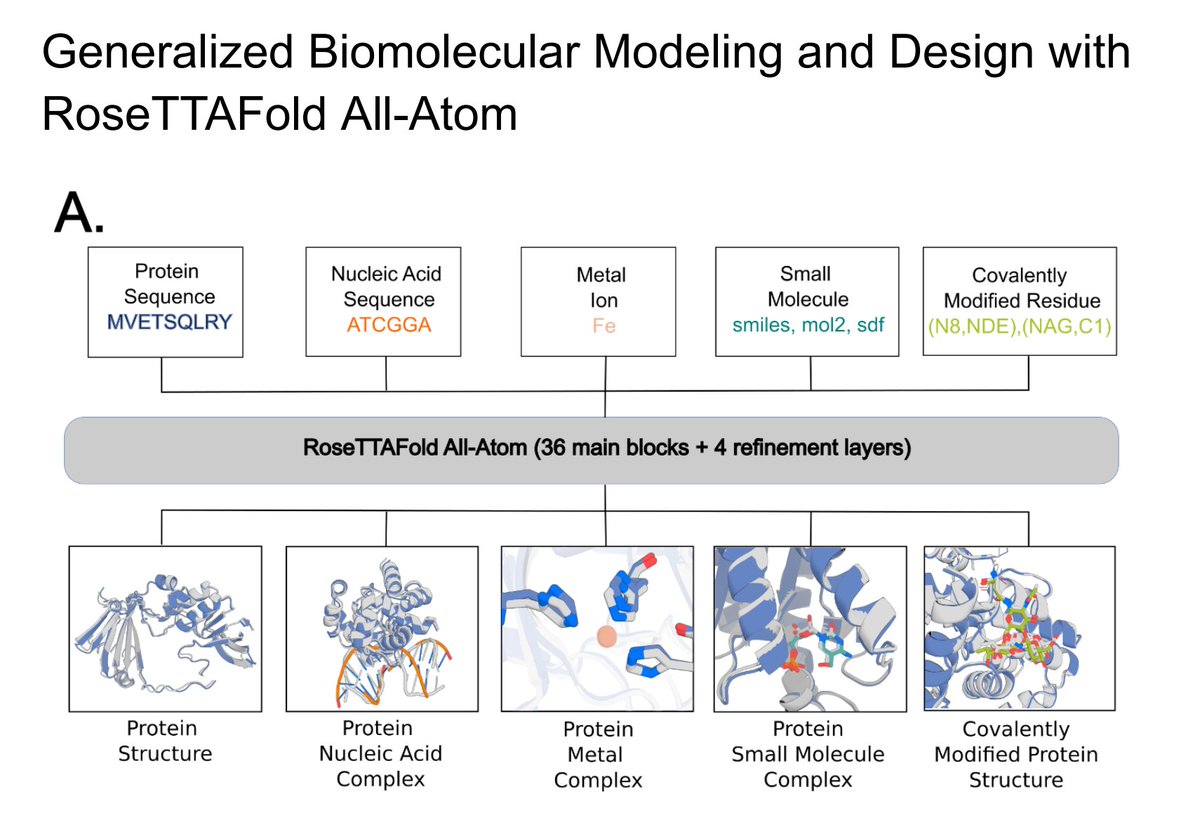
Researchers in Science present a next-generation protein structure prediction and design tool, #RoseTTAFold All-Atom, that can accept a wide range of ligands and covalent amino acid modifications. Learn more in this week's issue: scim.ag/6Ea



Our work on modeling and designing biomolecular assemblies is now in @ScienceMagazine. science.org/doi/10.1126/sc… RFAA Code: github.com/baker-laborato… RFdiffusionAA Code: github.com/baker-laborato…







Come work with me, @samuel_stanton_ , and the fantastic @PrescientDesign @genentech team as a grad student intern this summer in NYC! careers.gene.com/us/en/job/2023… careers.gene.com/us/en/job/2023… 🧵1/




The newest @FredHutch Basic Sciences lab opens its doors! The @SRsrivatsan Lab is developing new sequencing technologies to interrogate structure-function relationships during vertebrate development. The lab is growing and hiring at all levels: srivatsan-lab.com

Very excited to share RoseTTAFold All-Atom and RFdiffusion All-Atom, methods for structure prediction and design of biomolecular assemblies! biorxiv.org/content/10.110… 1/n


