Nirav Patel
2.2K posts


Nirav Patel
@Patel_omics
Science & Technology Advisor @ 10X Genomics, views and opinion are my own. Passionate about Science, Technology, Business and Building Teams








We GoT a Filling…uuuuhoooo 𝗧𝗮𝗿𝗴𝗲𝘁𝗲𝗱 𝘀𝗲𝗾𝘂𝗲𝗻𝗰𝗶𝗻𝗴 𝗼𝗳 𝗺𝘂𝘁𝗮𝘁𝗶𝗼𝗻𝘀 𝘃𝗶𝗮 𝗥𝗡𝗔-𝘁𝗲𝗺𝗽𝗹𝗮𝘁𝗲𝗱 𝗴𝗮𝗽 𝗳𝗶𝗹𝗹𝗶𝗻𝗴 𝗼𝗳 𝗼𝗹𝗶𝗴𝗼𝗻𝘂𝗰𝗹𝗲𝗼𝘁𝗶𝗱𝗲𝘀 𝗳𝗼𝗿 𝘀𝗶𝗻𝗴𝗹𝗲-𝗰𝗲𝗹𝗹 𝗥𝗡𝗔-𝘀𝗲𝗾 Download here: biorxiv.org/content/10.648… 1/ 🧬 New preprint! We developed GoT-Multi-Gap — a method to detect expressed mutations in single cells without needing to know the variant sequence in advance. Built on RNA-templated gap filling. Thread 🧵👇 2/ The problem: detecting mutations in scRNA-seq is hard. Coverage is sparse, and capture methods lose sensitivity the farther a mutation sits from the transcript end. Ligation-based approaches (like our earlier GoT-Multi) require allele-specific probes — limiting scalability. 3/ Our solution: exploit the dual enzymatic activity of Bst FL polymerase. It reverse-transcribes across a gap between flanking probes on mRNA, then uses nick translation to activate the downstream probe for ligation. No allele-specific design needed. 4/ The trick: the downstream probe starts with a 5’-OH (ligation-incompetent). After Bst fills the gap and nicks the terminus, a 5’-phosphate is exposed → SplintR ligase seals the nick. The mutation sequence gets written into the product automatically. 5/ We validated the chemistry in situ with padlock probes + rolling circle amplification on 18S rRNA. Key insight: exonuclease-resistant backbone mods on the downstream probe fine-tune polymerase vs. nick-translation competition, boosting efficiency. 6/ We then built this into 10x Flex (GoT-Multi-Gap). Three cell lines (MCF-7, SK-BR-3, LnCAP), 61 SNVs targeted across a range of gap sizes. Critically — the gap-filling step didn’t perturb Flex performance. UMI counts, gene counts, correlations all preserved. 7/ Results: among targets with ≥10 genotyped cells, mutant calls showed 80-100% specificity to the expected cell line. The main driver of detection sensitivity? Target transcript abundance (Pearson R=0.37, P<0.01). 8/ What didn’t matter much: gap length (4-12 bp), mutation position within the gap, and probe GC content (within the 44-72% window). The assay captures variants across the full gap with comparable sensitivity. 9/ Bottom line: GoT-Multi-Gap enables simultaneous whole-transcriptome + multiplexed mutation profiling from the same single cell, without prior knowledge of variant identity. Scalable, compatible with existing workflows, and ready for complex mutational landscapes. 10/ Huge thanks to all authors and absolute special shout out to @MircaSaurty, @lee_hower and @Kellieiswise and the incredible team @WeillCornell and @scilifelab.







The convergence of AI and biology is not a future possibility. It's underway. Which companies do we see building the future of healthcare? Read @Shea_ARK's The Multiomics–AI Flywheel: Why ARK Believes AI And Biology Are Converging To Reshape Healthcare arkinvestmentmanagement.substack.com/p/the-multiomi…






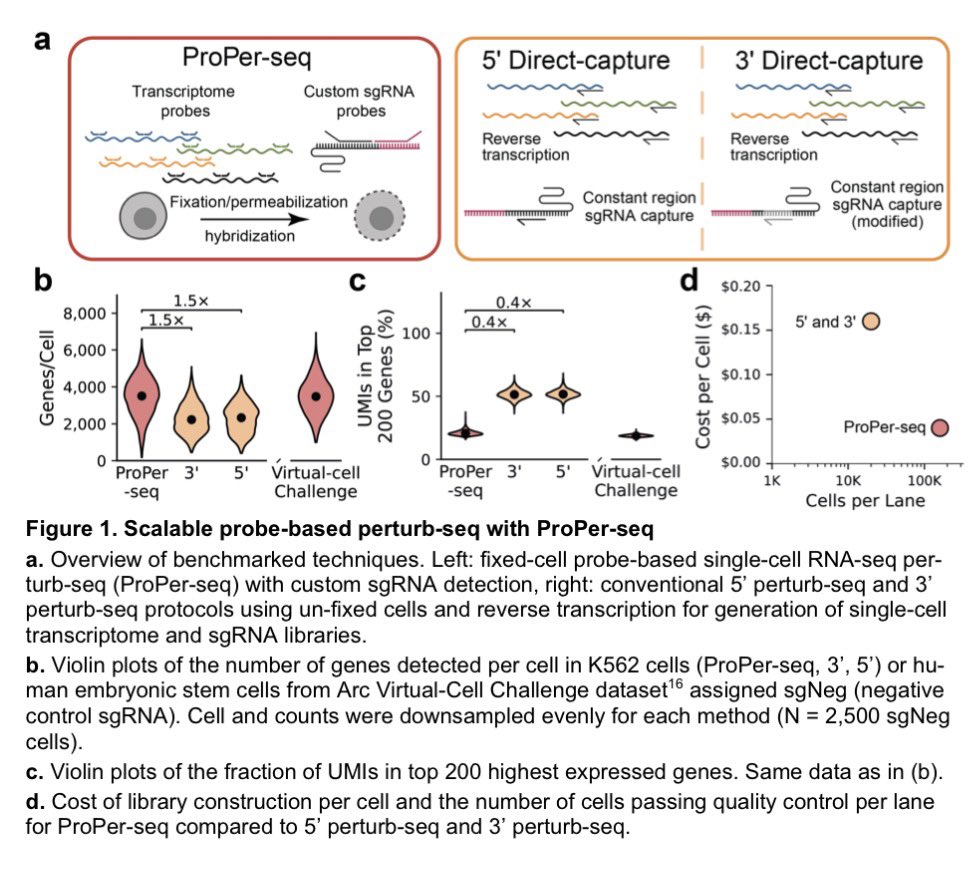
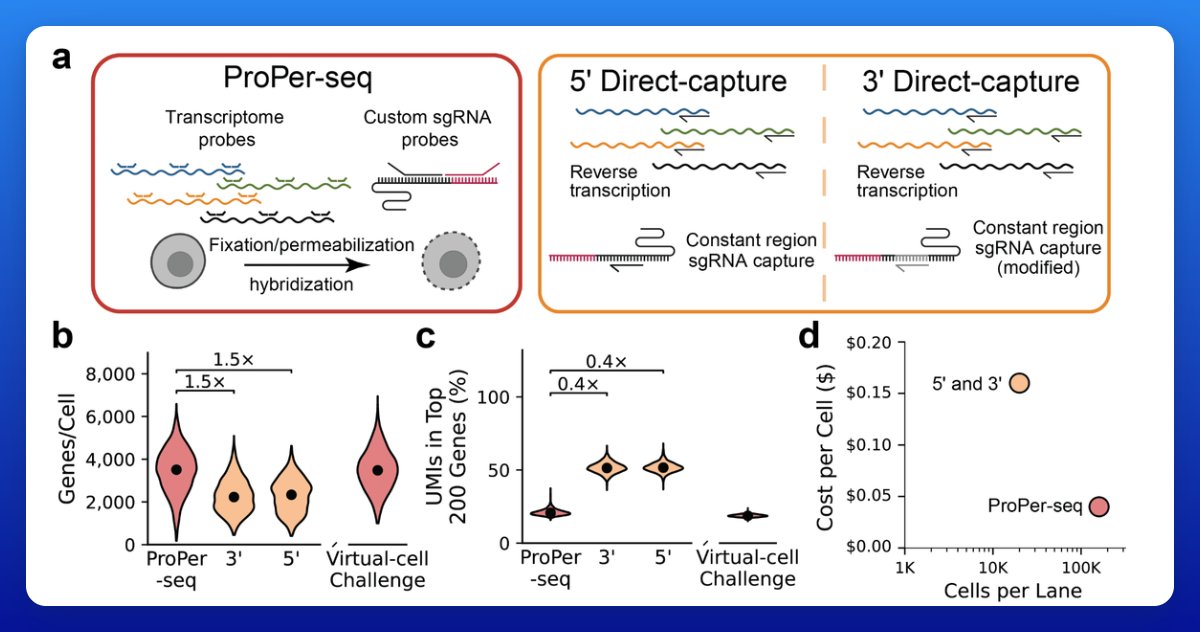

More exceptional technology from the Arc Institute: "ProPer-seq faithfully reproduces gold-standard perturb-seq phenotypes while achieving 4-fold cost reduction and 50% increased gene detection per cell." Seems like a big improvement (in cost and resolution) for one of the most central methods in modern functional genomics. The required inputs to scale "virtual cell" models continue to plummet in cost...












