Honggui Wu
492 posts


@Wu_Honggui
Postdoc in Feng Zhang lab @BroadInstitute. Graduated from Sunney Xie Lab at Peking University.





Very excited to share our work on high-resolution single-cell Micro-C, which is on bioRxiv now. Here we reveal that cohesin-mediated loop extrusion participates in transcription with kilobase-resolution single-cell 3D structures. biorxiv.org/content/10.110…









I am excited to share our latest #preprint: ChromoGen: Diffusion model predicts single-cell chromatin conformations researchsquare.com/article/rs-463… As the title suggests, we achieved de novo prediction of 3D chromatin structures using DNA sequence and ATAC-seq using AI.


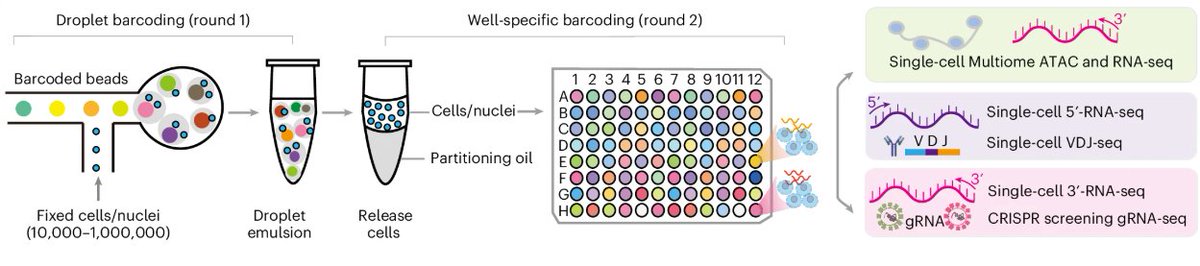
Thrilled to introduce a new single-cell 3D genome analysis tool to our toolkit! Our high-throughput droplet Hi-C (dscHi-C) and dscHi-C-multiome methods use droplet microfluidics to generate tens of thousands of single-cell Hi-C profiles with high sensitivity in a single batch.