
Dingchang Lin
206 posts

@DingchangLin
Asst. Prof @JohnsHopkins; Broadly interested in neurotechnology, genetically encoded tools, and materials science.
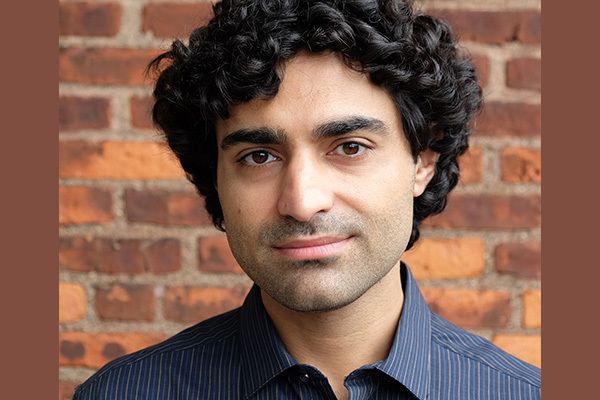

















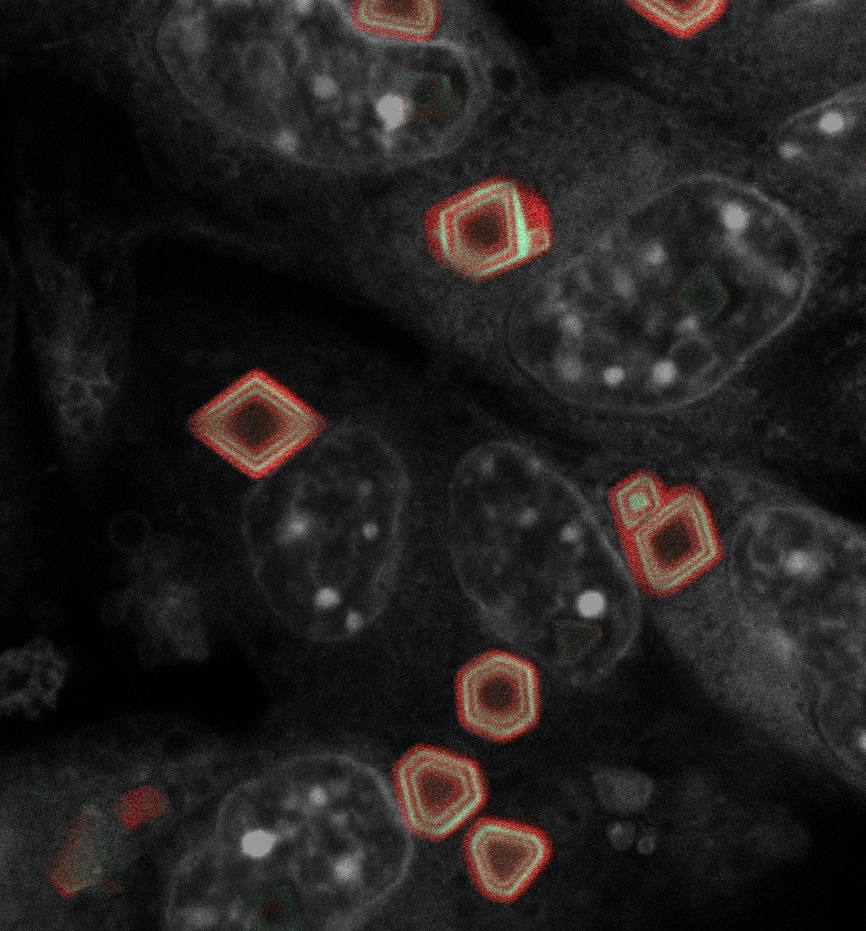
🚨 Today in @Nature, we report GEMINI—a genetically encoded intracellular memory device that writes cellular dynamics into tree-ring-like fluorescent patterns within cytoplasmic protein assemblies.[1/n] nature.com/articles/s4158…





🚨 Today in @Nature, we report GEMINI—a genetically encoded intracellular memory device that writes cellular dynamics into tree-ring-like fluorescent patterns within cytoplasmic protein assemblies.[1/n] nature.com/articles/s4158…





🚨 Today in @Nature, we report GEMINI—a genetically encoded intracellular memory device that writes cellular dynamics into tree-ring-like fluorescent patterns within cytoplasmic protein assemblies.[1/n] nature.com/articles/s4158…