Yi Zhou
126 posts


Yi Zhou
@_y1zhou
Protein design @BiomapI. Views are my own.








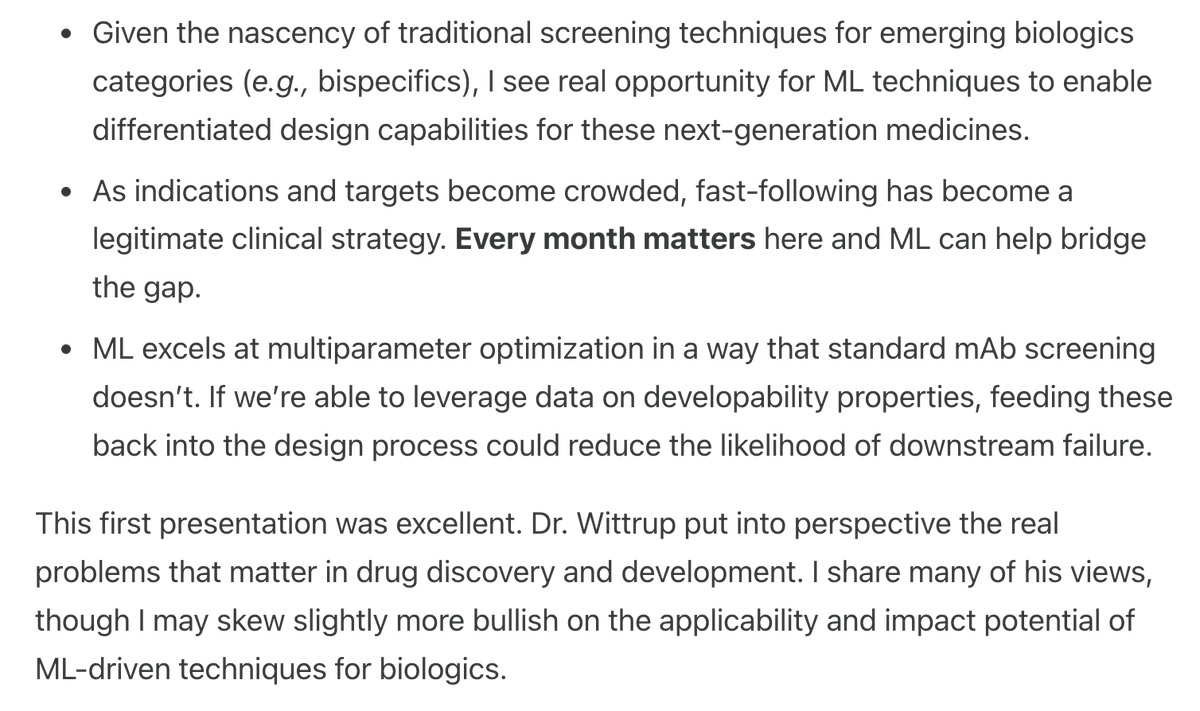
turns out everything nabla’s model claims it can do, chai’s can too! so i guess the suspicion that developability being a naturally emergent property of a well-trained model is true the GPCR result also seems emergent (surprising!), given that chai-2 could do it from the start but just was never tested on it in the original release insane speed from chai, i wonder if this result was just sitting on ice or they literally contracted a CRO the second they saw Nabla’s release. the post at 11:48pm PST makes me feel like it was the latter, which is a fun story it does beg the question a little of whats next to hill climb on in this subfield if the traits i assumed are next to optimize for (solubility, etc) are simply going to naturally pop out of any good model, regardless of who are the ones developing it. in-vivo properties i guess?



Today, we’re releasing new data showing that Chai-2 can design antibodies against challenging targets with atomic precision. >86% of our designs possess industry-standard drug-quality properties without any optimization. Thread👇




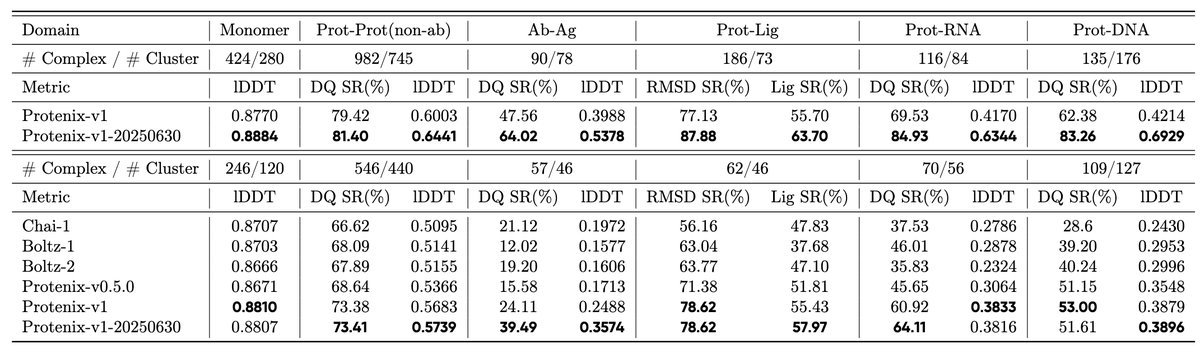
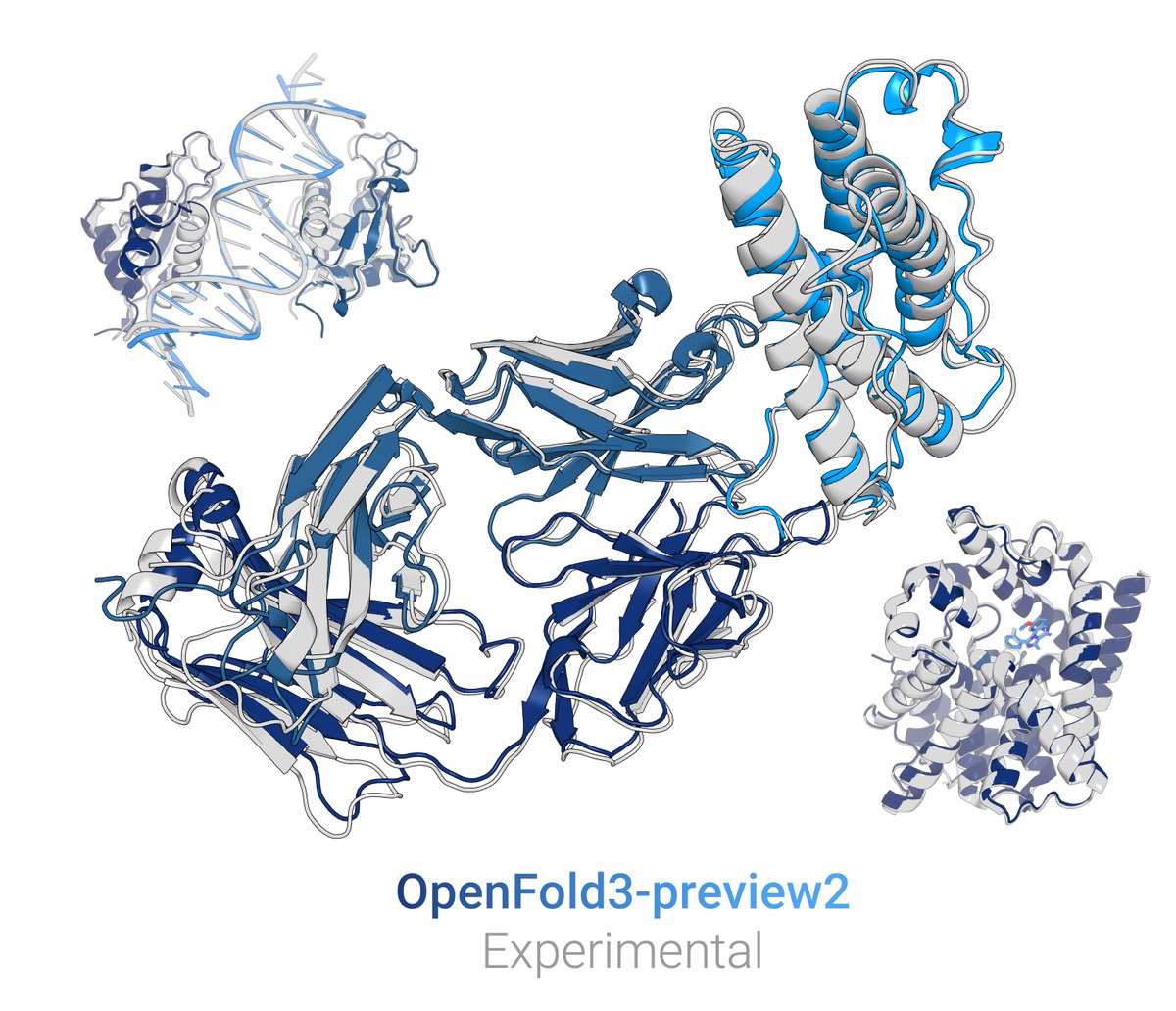
OF3p is already quite good. For any modality, it is comparable (or better) to the best existing OSS model for that modality. On RNA, where we spent considerable effort, it is at AF3-parity. RNA is a challenging modality where all models (incl AF3) leave much room for improvement.





Accurate inverse folding models are crucial for antibody CDR design, especially with the rise of LLMs in protein engineering. This benchmark evaluates these models, aiming to improve antibody therapeutics. biorxiv.org/content/10.110…










We’re excited to release another update to the Boltz repo: v0.3.0. This release includes several important features, including our confidence model and low memory mode. Give it a try! github.com/jwohlwend/boltz