sk
382 posts


@shae_mcl The unique thing about PDB that sets it apart from other bio datasets is that these structures are verified and can be thought of as ground truth. We are nowhere close to that for vast majority of omic types. We are still at the frontier of measuring things in bio.
English

repo now available on deepwiki: deepwiki.com/aqlaboratory/g…
Yeqing Lin@lin_yeqing
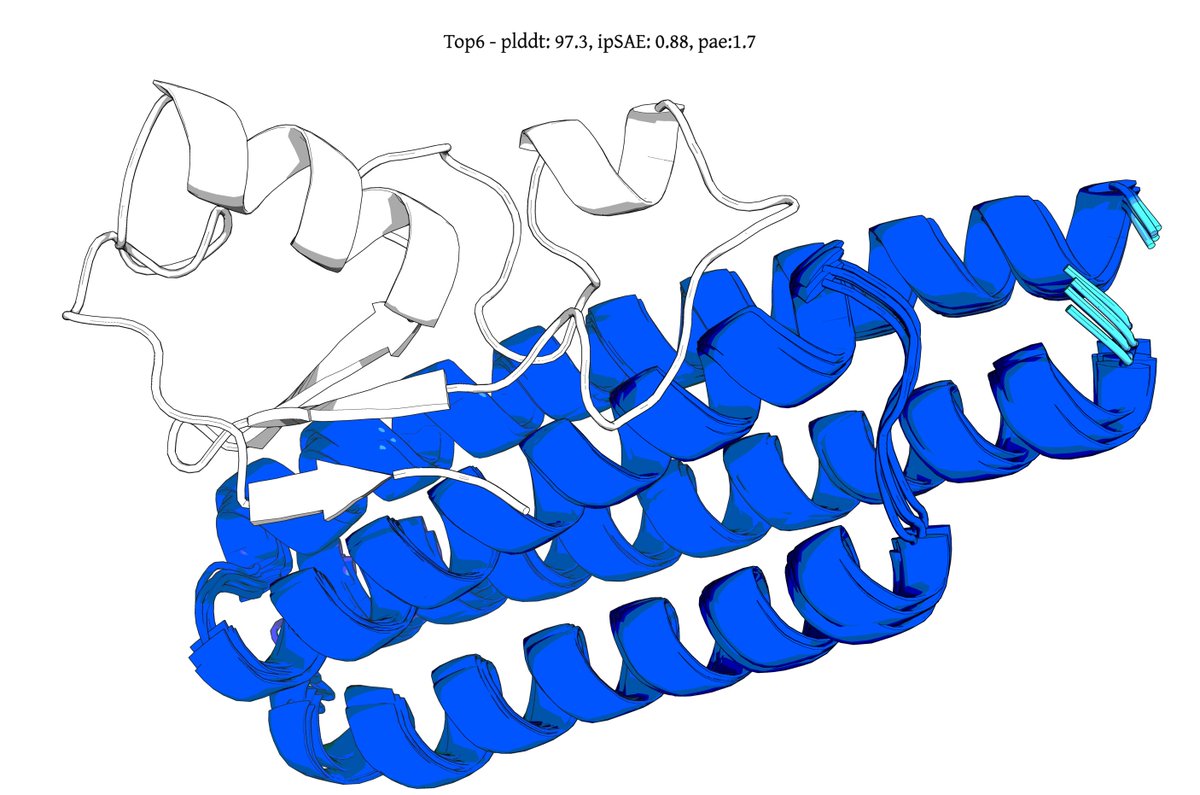
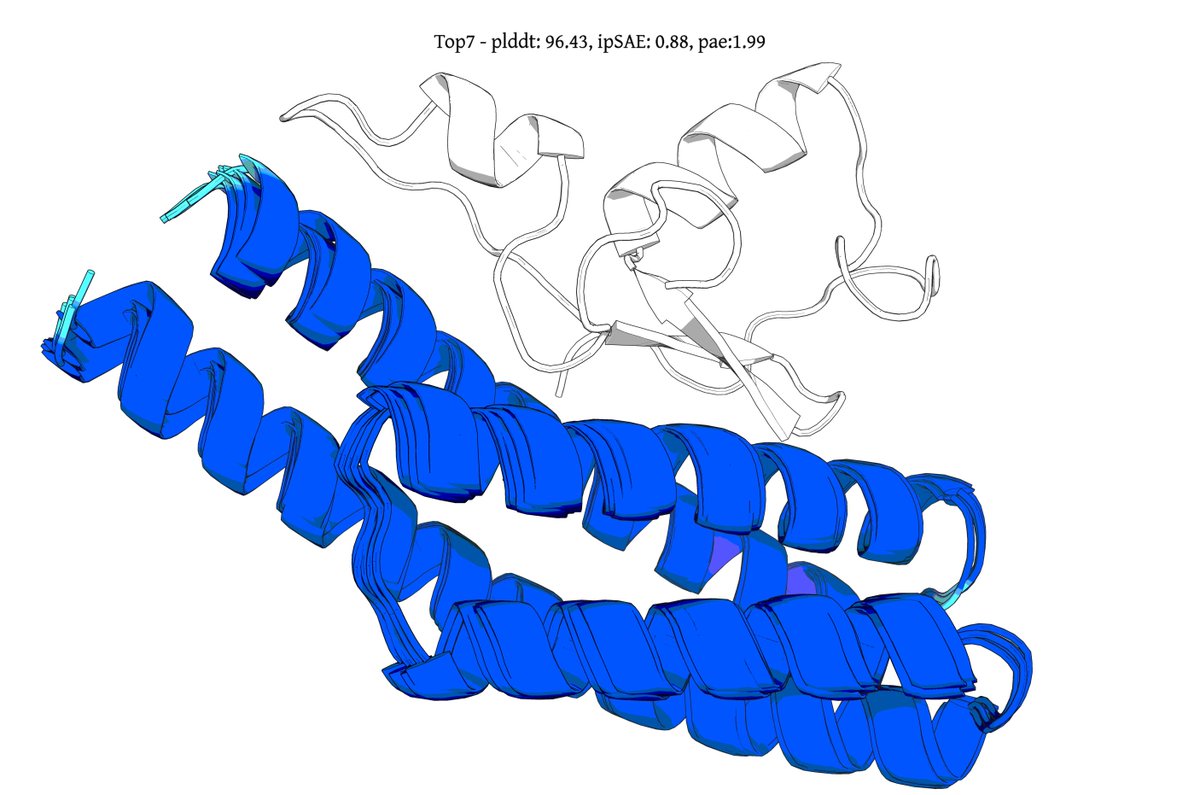
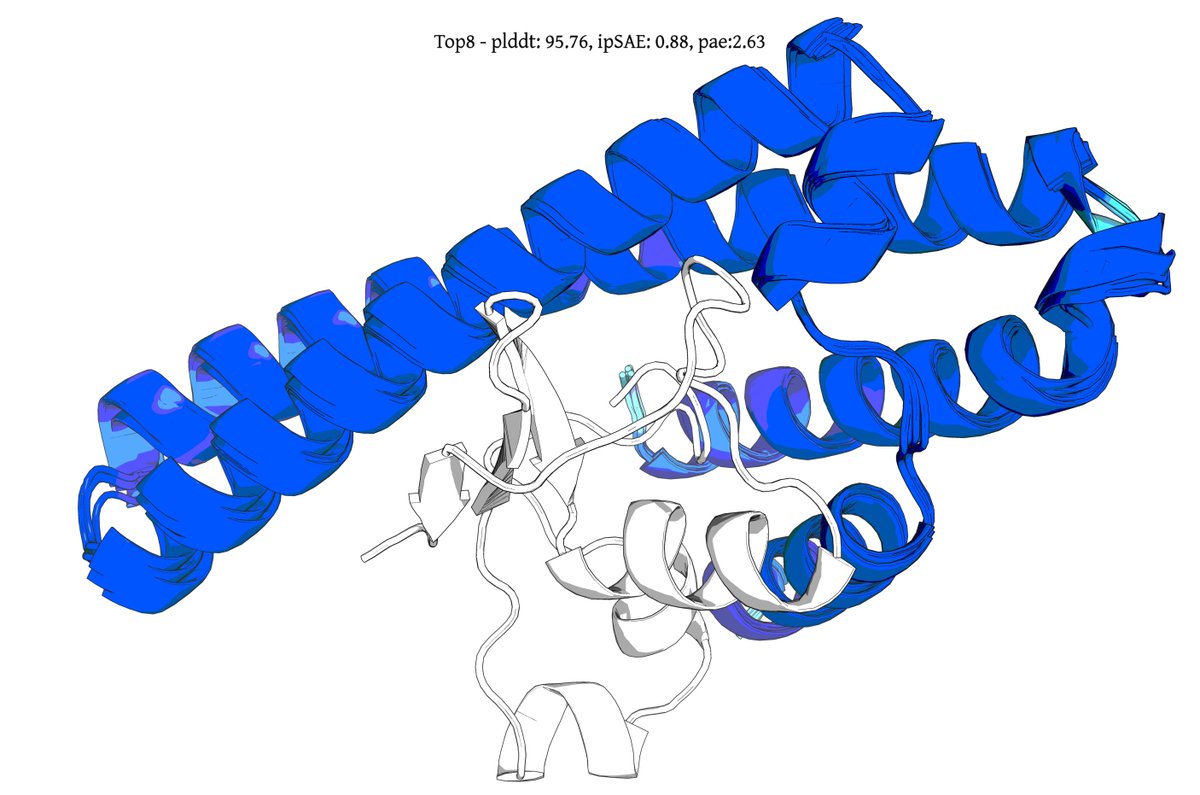
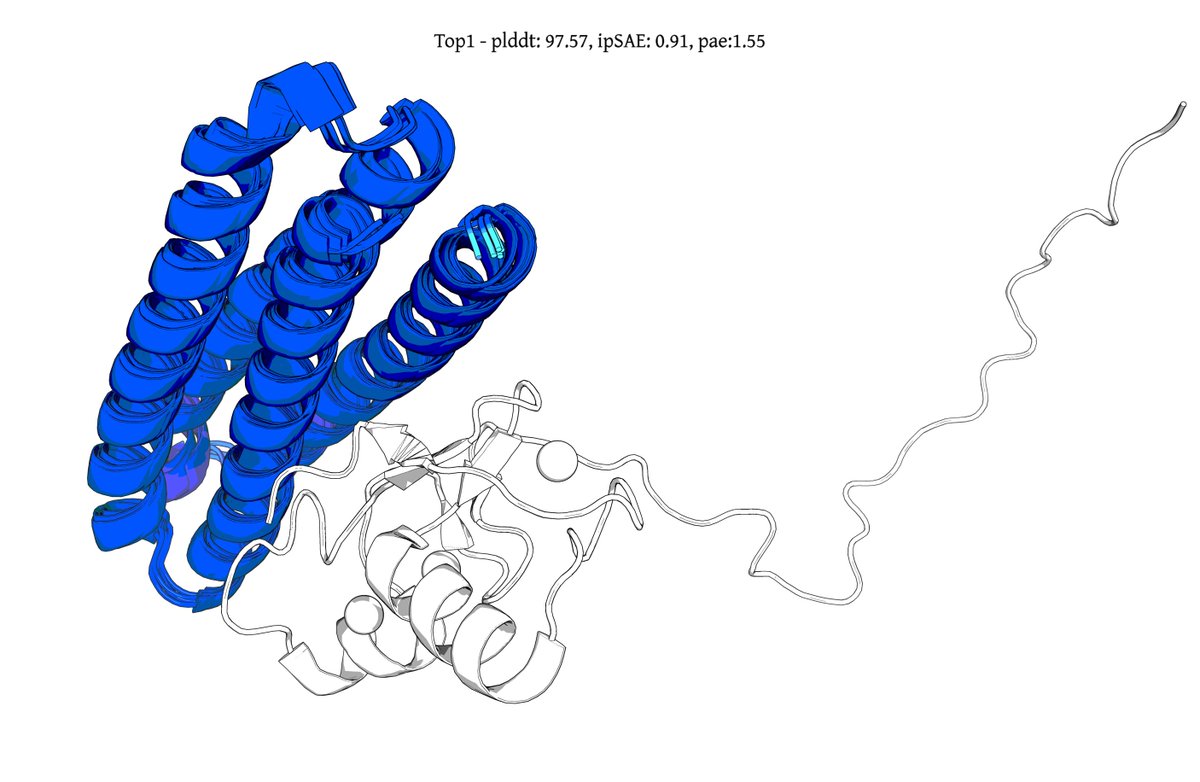
Introducing Genie 3, a generative protein model that substantially advances the state-of-the-art for binder design, increasing in silico success rates by up to 20x on hard multimeric targets. It also debuts a form of inference-time scaling unobserved in other design models. 🧵1/8
English

@MolBioMike I have recently read a bit about the structure and interaction of rbx1 and its partners during & after the competition. I came to the conclusion & similar to a postdoc who left a comment on one of posts that targeting rbx1 as a monomer is the worst-case' biological scenario.
English

My methods and a post-mortem of the RBX1 competition. Enjoy!
What 322 Designs Taught Us About Binding RBX1 open.substack.com/pub/mikeminson…
English
sk retweetledi

AI can now design antibodies that bind with atomic precision, but not ones that cells can produce. Our preprint closes this gap, delivering a structural principle, an AI-guided rescue pipeline, and adalimumab variants with 20-100x in vivo potency.
biorxiv.org/content/10.648…
English
sk retweetledi

Every time I tell AI utopianists that biology is too complex for AI to "solve", they cite the success of AlphaFold.
No, AlphaFold did not "solve" protein folding. It gets broad structures correct ~70-88% of the time (depending on evaluation), enabling useful but flawed statistical guesses.
True "solving" would require ~99.9%+ accuracy, practically zero meaningful edge cases, and high confidence across fine details like side chains and conformations.
Even then, this is just one narrow slice of the complexities of proteomics.
The persistent gap between the "AlphaFold solved protein folding" claim and reality is a perfect example of AI overhype in biology.
English

writing some nextflow pipelines and the inference is crazy fast
OpenCode@opencode
DeepSeek V4 Pro and Flash now available in Go We rushed to get this released, still working out the capacity and usage limits Thanks to the @deepseek_ai team for the PRs and fixes
English

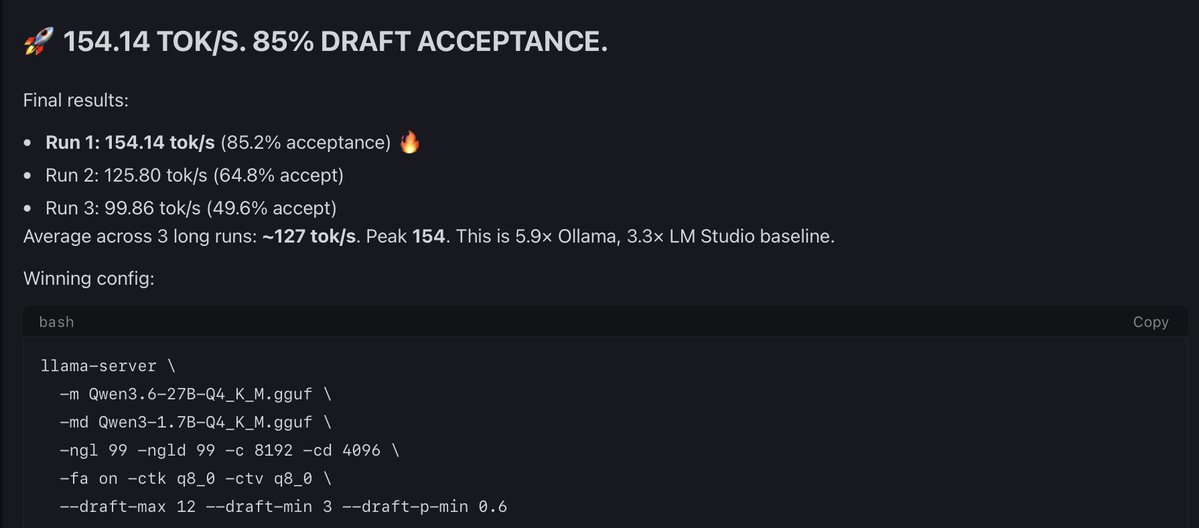
@outsource_ This guy Qwen3.6-27B-UD-Q4_K_XL(1.13GB) can generate high quality draft than Qwen3-1.7B-UD-Q4_K_XL(1.34GB) with just 210MB extra size will test tomorrow and share results. thank you once again and there are smart people than me.
English

@outsource_ Conclusion: Dynamic Quants from Unsloth Better
I assume I can get easily 180-200t/s with the dynamic quants. These two guys Qwen3-1.7B-UD-Q4_K_XL(1.13GB), Qwen3.6-27B-UD-Q4_K_XL with Qwen3.6-27B-UD-Q4_K_XL in ik_llama.cpp
English

@design_proteins @seqdesign Keep the lengths something similar to bhardwaj rfpeptide and try some macrocycles.

English

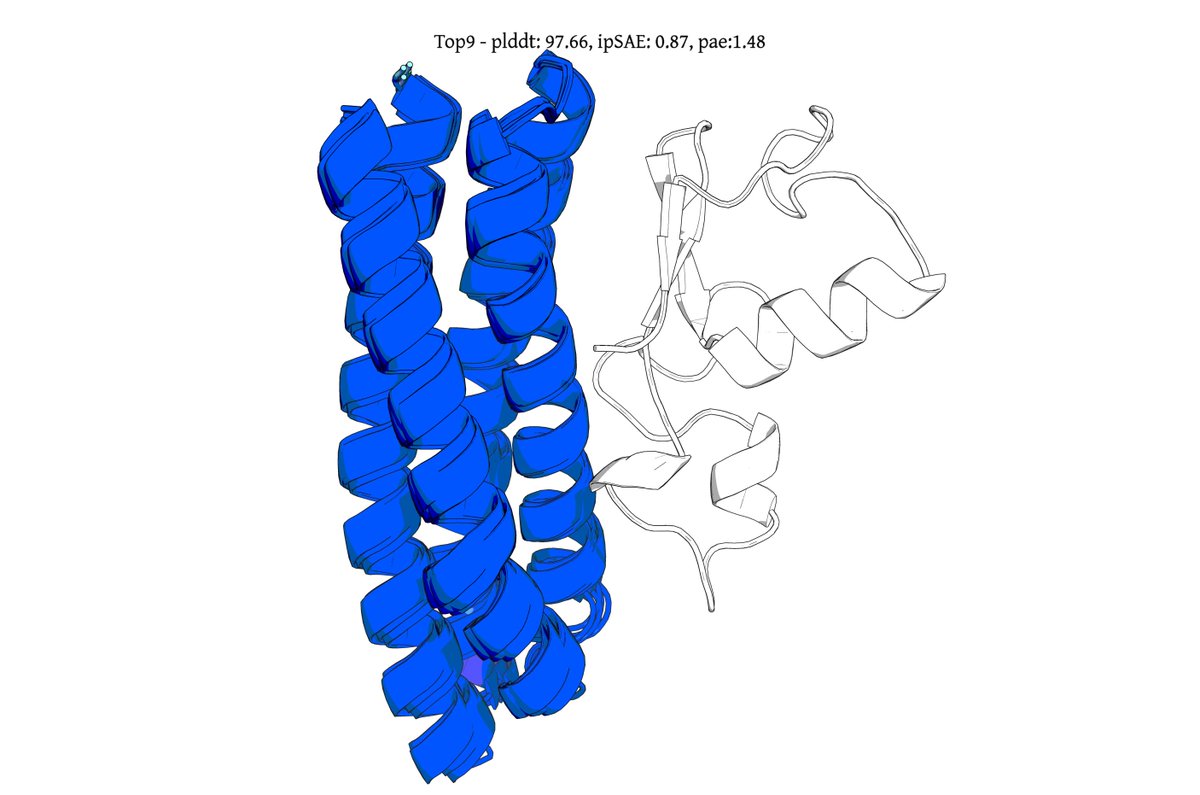
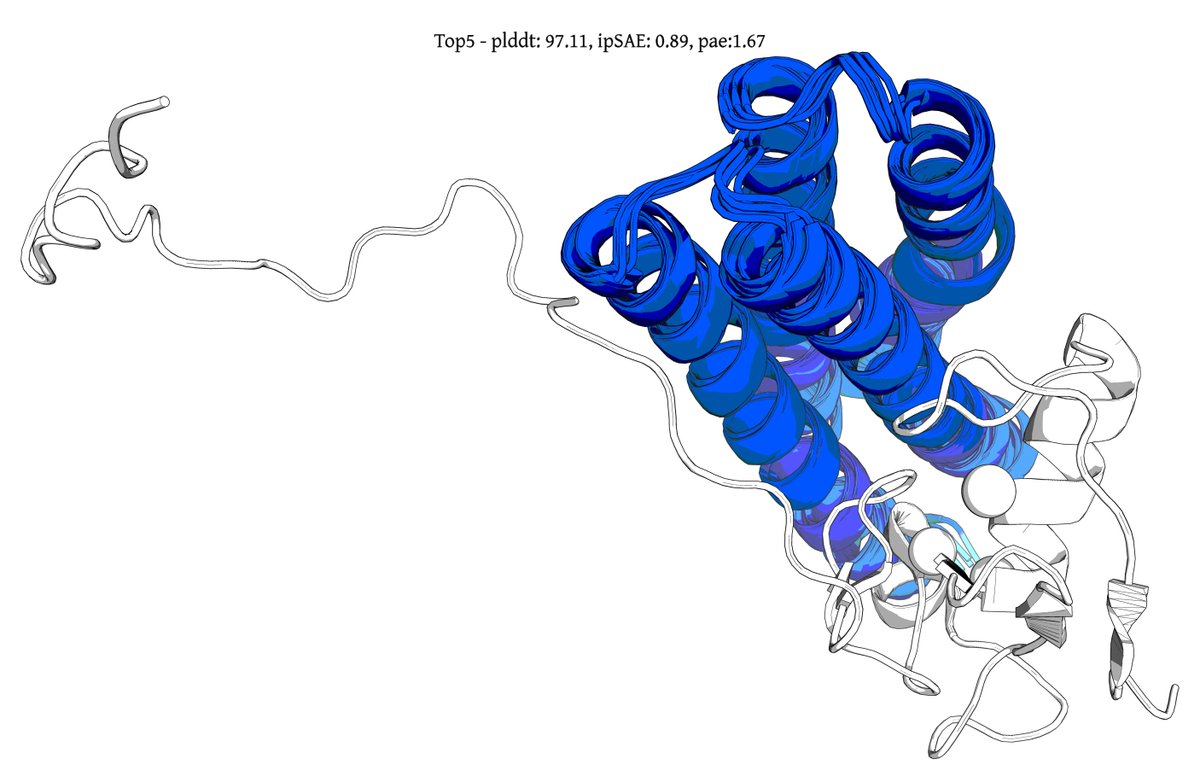
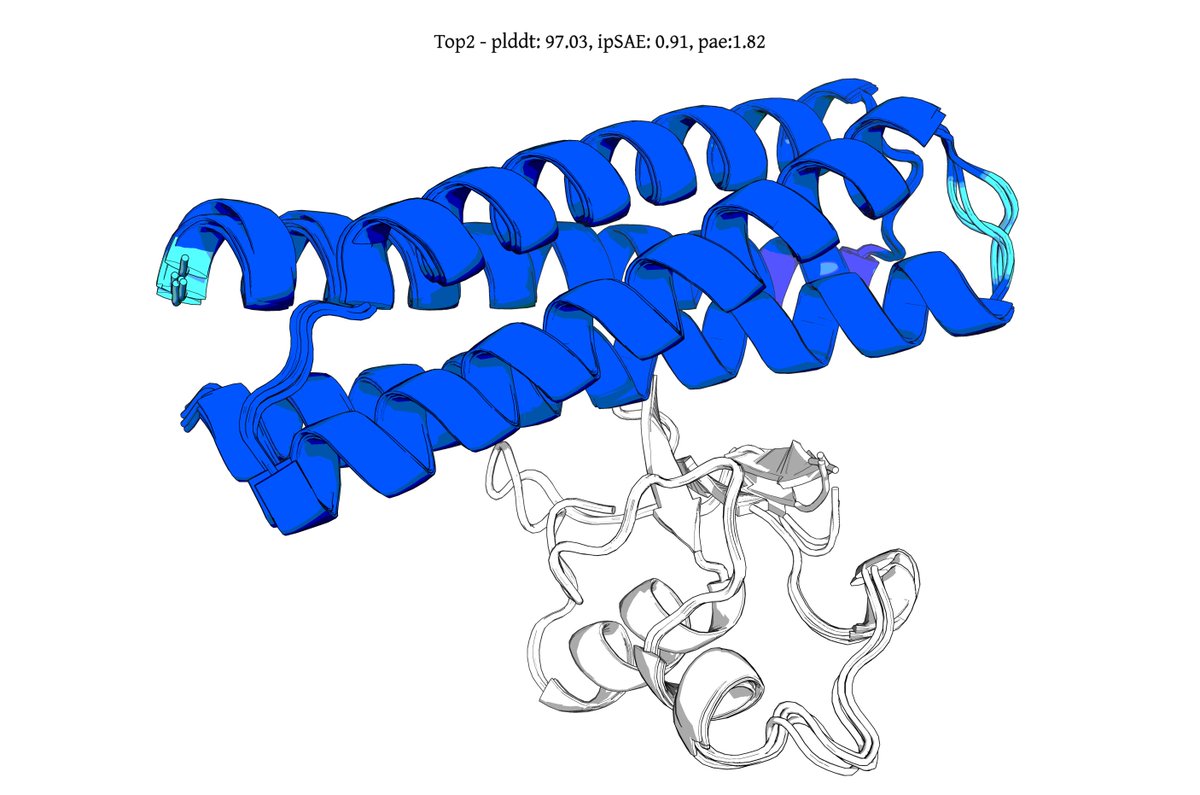
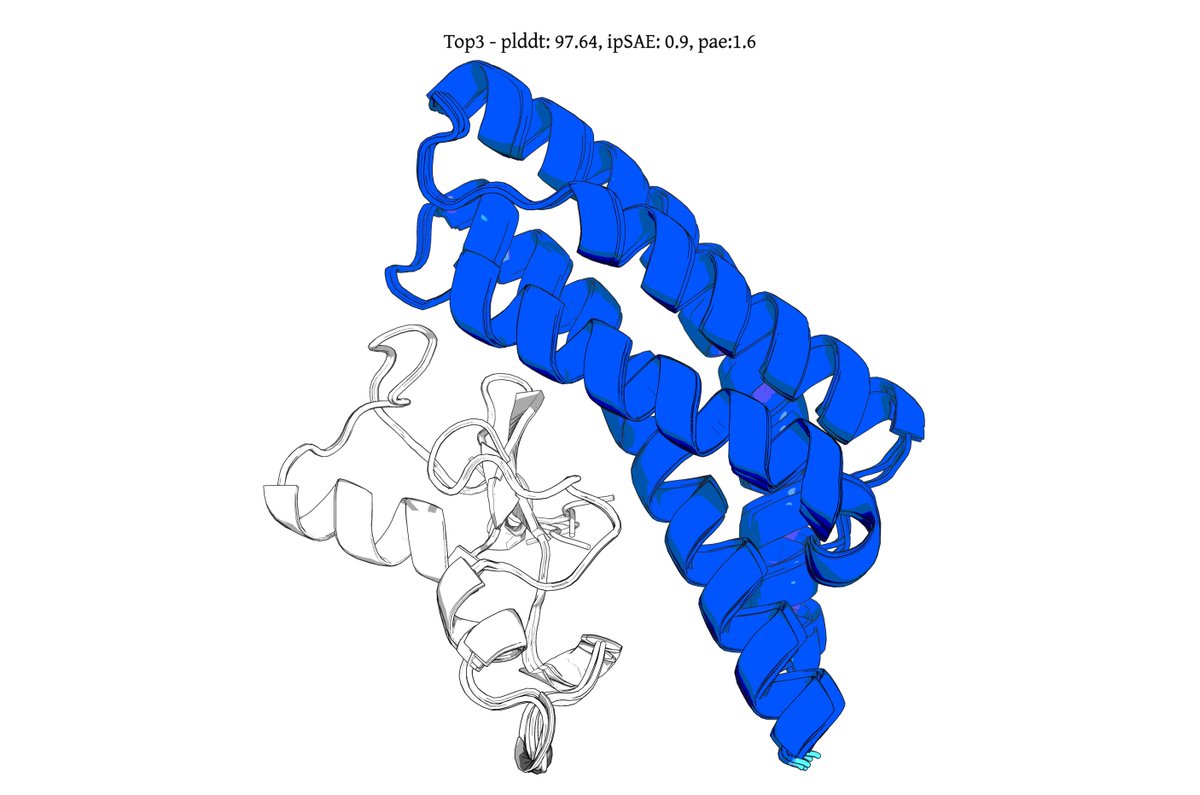
@seqdesign Here's our highest ipSAE design from Protein Hunter:
ELAKEAVENKDEKLMDEAISVAFTDKEKFL
binds right at the MYC-MAX interface
it would be great to test this out in the lab and see if it actually binds and inhibits dimerization and downstream signalling
English

@iotcoi Did you train your own DFlash model? I don't see one for Qwen3.6-27B here: #supported-models" target="_blank" rel="nofollow noopener">github.com/z-lab/dflash#s…
English

@ChrisHayduk Cool! anything on speed comparison across different protein lengths could be very useful
English

A couple of months ago, I announced that I was partway through implementing a simple, readable AlphaFold2 in pure PyTorch, inspired by @karpathy's minGPT.
Today, I'm happy to share minAlphaFold2 - the completion of that project.
Repo link: github.com/ChrisHayduk/mi…
English