
Alex Federation
679 posts


@afederation
Co-founder, CEO of Talus Bio illuminating the regulome | controlling the genome | drugging the 'undruggable' #proteomics, #teammassspec, #chembio



🐠 Everything we know about biology has been built on an incomplete picture. DNA tells us what a cell might do. Proteins tell us what it’s actually doing. Pumpkinseed announced their $20M Series A today (led by Future Ventures and NfX) to build the platform that reads proteins directly—for the first time. Proteomics has always faced a fundamental constraint: you can only measure what you already know to look for. The current workhorse, mass spectrometry, requires matching protein fragments against reference databases. If a protein isn't in the database, or doesn't ionize reliably, it's invisible. Other approaches rely on fluorescent labels or antibody-based affinity methods, which introduce their own biases and blind spots. The result is a field that has spent decades generating an increasingly detailed map of a small, well-lit corner of the proteome, while biology’s most important data layer remains hidden. This isn't a sensitivity problem. It's a category problem. Existing tools were never designed to read proteins directly de novo. They were designed to find what researchers already suspected was there. Pumpkinseed is built to find everything else. And proteomics is harder than most people outside the field appreciate. When we account for post-translational modifications, non-canonical amino acids, and glycan decorations, there are roughly a thousand distinct chemical monomers in the proteomic alphabet, compared to the four bases of DNA. deSIPHR (de novo Sequencing and Identification of Proteins with High-throughput Raman spectroscopy) is Pumpkinseed's proprietary nanophotonic chip platform, fabricated with semiconducting manufacturing. With over 100 million sensors per square centimeter, it reads proteins, known or unknown, letter by letter — amino acid by amino acid — without a reference catalog of proteins, and at high-throughput. The result is direct, high-resolution proteomic data, including post-translational modifications, non-canonical amino acids, and single-cell detail, that mass spectrometry-based approaches cannot match. What is Raman spectroscopy? Rather than tagging or fragmenting proteins, Raman spectroscopy reads the molecular vibrations of individual molecules. Each amino acid vibrates at a characteristic frequency, producing a unique physical signature that deSIPHR detects directly. This is physics reading biology in the most literal sense. With conventional Raman spectroscopy, only about one in ten million photons interacts with a molecule usefully, far too weak for single-molecule work. Pumpkinseed's answer is a silicon photonic chip patterned with a billion sensors per wafer. Those sensors concentrate light into volumes smaller than a single protein, amplifying Raman scattering efficiency by over 10 million-fold. And their future ventures? “The longer-term ambition is the virtual cell, a computational model that simulates not just how proteins fold but how they interact, respond to drugs, and behave under perturbation inside a living system. AlphaFold demonstrated what structural AI can do once a sequence is known. The gap that cannot be closed is determining the sequence itself from biological samples, particularly for proteins carrying modifications absent from existing databases. Pumpkinseed is designed to supply that input layer. "If the Human Genome Project was the data infrastructure that enabled genomic medicine, we believe the high-resolution proteomic dataset Pumpkinseed is building could be the analogous foundation for AI-driven biological discovery," co-founder Dr. Jen Dionne says. "In our vision, the molecular signatures driving disease, aging, and ecosystem health become fully legible. Medicine shifts from reactive to proactive. Optimal healthspan moves from aspiration to achievable reality." —synbiobeta.com/read/pumpkinse… • The biology mining company: Pumpkinseed.Bio • Today’s News: pumpkinseed.bio/news/pumpkinse…

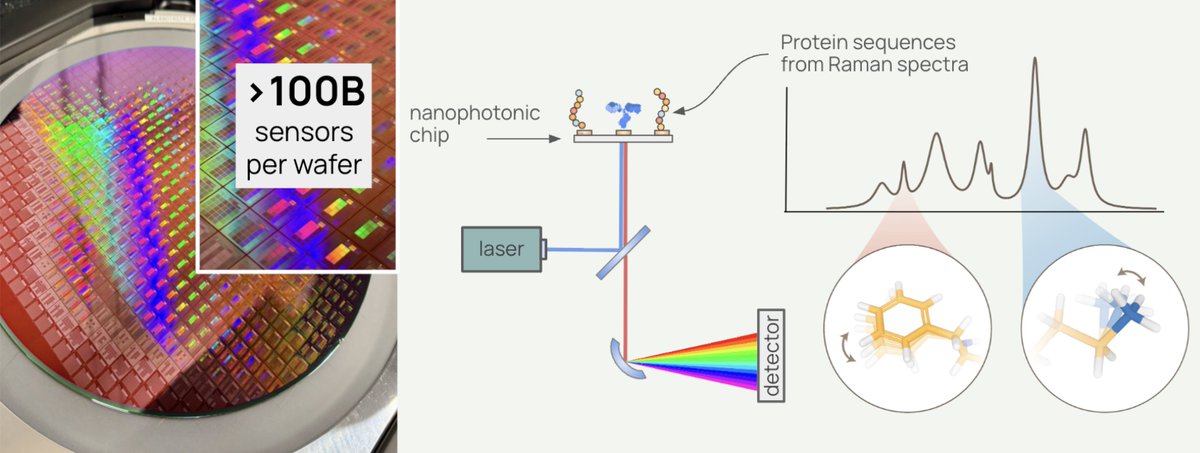

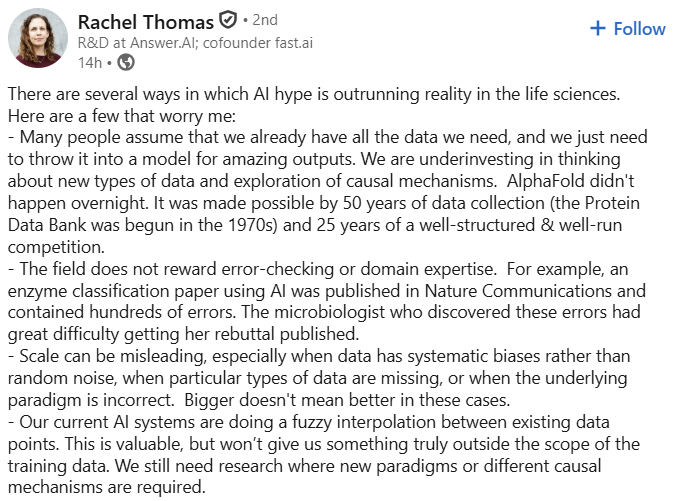

AI x Bio teams like Origin have coding agents, scaling laws, and a wave of big biotech deals all at their backs. This is barely touched territory. Crazy what this small team can do now.

In a phase 1 study of the oral p53 reactivator rezatapopt in heavily pretreated patients with TP53 Y220C–mutated solid tumors, the most common adverse events were nausea and vomiting, and the overall response was 20%. Full PYNNACLE study results: nej.md/3OIQC5P Science behind the Study: Restoring Function to a Variant of p53 in Solid Tumors nej.md/3N0pQW8


The Humanity Project People always ask me: if we want to cure all diseases with AI, shouldn't we be building a huge robotic warehouse to run closed loop experiments on mice, human cells, and so on? From a naive outside perspective, this may seem like the obvious thing to do, analogous to what Periodic Labs is doing for materials and what the robotics companies like Physical Intelligence are doing for laundry and so on. The problem is, for most diseases, experiments in cultured cells, mice, and other models are poorly predictive of success in actual humans. If you synthesize a material in the lab and find it has a certain conductivity, then that’s the conductivity; by contrast, if something works in a mouse, you still have to test it in a human, and mouse biology is so poorly predictive of human biology that for some diseases success in mice actually anticorrelates with success in humans. Closed loop reinforcement learning in mice might be a great way to learn how to cure mouse diseases, but the only way to cure human diseases is to run more experiments in humans. To cure all diseases, in other words, what we actually need is a massive, centralized effort to scale up clinical trials testing new AI-generated therapeutic hypotheses. Such an effort would serve several goals: it would identify novel mechanisms for treating diseases; it would train AI systems to understand human biology; and it would build the infrastructure needed to streamline and accelerate clinical trials going forward. Centralization would allow us to minimize duplication, maximize data availability for training, and ensure the resulting infrastructure is shared. An effort of this scale -- which would dwarf the genome project and consume resources similar to the entire NIH budget -- is not just our best chance to cure all diseases. It is also our best chance to remain ahead in biotechnology, where the US is rapidly losing ground to China. I call it the Humanity Project, and I describe it in a new essay, linked below. The first emphasis in the Humanity Project is on novelty. Medicine moves forward when we identify new therapeutic mechanisms -- think GLP-1s, CAR-T cells, and checkpoint inhibitors. Within a few years, AI agents will be as good as or better than humans at proposing novel therapeutic mechanisms. However, testing novel therapeutic mechanisms is risky and rarely pays off, so most biopharmas largely avoid it. Instead, 96% of clinical trials today are aimed at obtaining approval for new drugs that use existing therapeutic mechanisms, rather than testing new ones. If we want to cure all diseases, we need to massively expand the number of clinical trials we run that examine novel hypotheses, both as a way of identifying novel therapeutic mechanisms and as a way of training AI models to better understand human biology. This brings us to the second emphasis of the Humanity Project, which is on scale. The Humanity Project would be a centralized project to run 2000 drug programs testing novel, AI-generated therapeutic hypotheses through clinical proof of concept over 5 years. The Project would cost between $50B and $200B -- equivalent to the amount allocated for the California High Speed Rail, or the 5-year budget for the NIH --, would increase the number of first-in-class drug trials by 400% to 800% over that period, and would benefit from various advantages of AI in proposing and prosecuting clinical trials. In the process, the Project would lay down the infrastructure needed to accelerate and scale future trials run by the private sector, accelerating US biotechnology as a whole. The logistical challenges associated with such a problem are intense. We need better infrastructure to recruit and identify patients; we need to streamline regulation to make it easier to get drugs into patients faster while ensuring safety; we need to scale up and accelerate manufacturing; and we need to plug all of this infrastructure into AI systems that can efficiently integrate information, generate hypotheses and plan experiments, and learn from outcomes. The Humanity Project would work on developing these, and would make the resulting infrastructure available subsequently to accelerate US biotech as a whole. And, importantly, if we are serious about curing all diseases in the coming decades and maintaining US leadership in biotechnology and medicine, there is no alternative. With all the wealth and value set to be created by AI, finding the resources to accelerate medicine in this way should be straightforward. We should begin now. Read the full proposal below.



Underrated Ideas in Biotech (#7) AlphaFold predicts a protein's structure solely by looking at its sequence. But structure does not always suggest function. The next frontier is to train a model that can predict any protein's FUNCTION solely from its sequence. This is incredibly important because the most useful tools in biotechnology come from NATURE. The thermostable polymerase used in PCR came from microbes in a Yellowstone geyser. The "repeats" in CRISPR were first identified in extremophilic microbes in Spain. GFP was discovered in a jellyfish. The list goes on. If we had a good model for sequence-to-function prediction, we could send biologists into nature to hunt for proteins with new, useful functions. They could sequence samples from the Arctic and jungles and oceans, upload those data into the model, and then discover lots of useful proteins that could be adapted into tools! This vision is not so far from reality, either. A nonprofit, called The Align Foundation, is already collecting the datasets needed to train this predictive model. The training dataset needs three variables: 1. The amino acid sequences of proteins; 2. Some kind of "quantitative functional score" indicating how well each protein performs in an experiment; 3. A function-definition, aka a detailed description of the experiment used to benchmark what the protein does. (There are many "classes" of protein functions, like antibodies, proteases, transcription factors, and so on.) Align is collecting millions of datapoints. They have scaled up a lot, and are now routinely collecting hundreds of thousands of data points in a single experiment, at a cost of ~$0.05 per protein. Say Align wanted to collect data on antibiotic resistance proteins. They would take antibiotic resistance genes, mutate them to make thousands of variants, and insert those sequences (with unique 'barcodes') into living cells. Next, they'd expose the cells to an antibiotic. Cells with functional proteins survive; many others die. Some variants will do 'OK,' but those cells grow slowly. By sequencing the barcodes, they can quantify the population of each variant, and thus figure out which proteins are REALLY GOOD, JUST OKAY, or fail entirely. Given these data, the next step is to train a model that predicts whether a given protein variant will be good or bad at defending a cell against antibiotics. The model will be good at doing this for antibiotic resistance proteins, but not much else! Align's goal, then, is to do these assays for proteins with many different functions and then "merge" them to build a model that can generalize across proteins. "As the dataset grows and more islands are sampled [they write], the models will become more generalized and capable of predicting the function of protein sequences that are increasingly distant from those that have been...measured." I hope this works. It would legitimately be one of the greatest unlocks for biotech progress more broadly.


The recent breakthroughs from @nablabio & @chaidiscovery emphasize a split in early biotech strategy. For the specific range of problems that antibodies address, making the binder, is becoming trivial. This forces a choice between 'fast but competitive' and 'AI intractable'. 🧵
