Arthur Ko
1.5K posts


Arthur Ko
@arthur5ko
Human geneticist at Children’s National Hospital. NHGRI DATA Scholar.








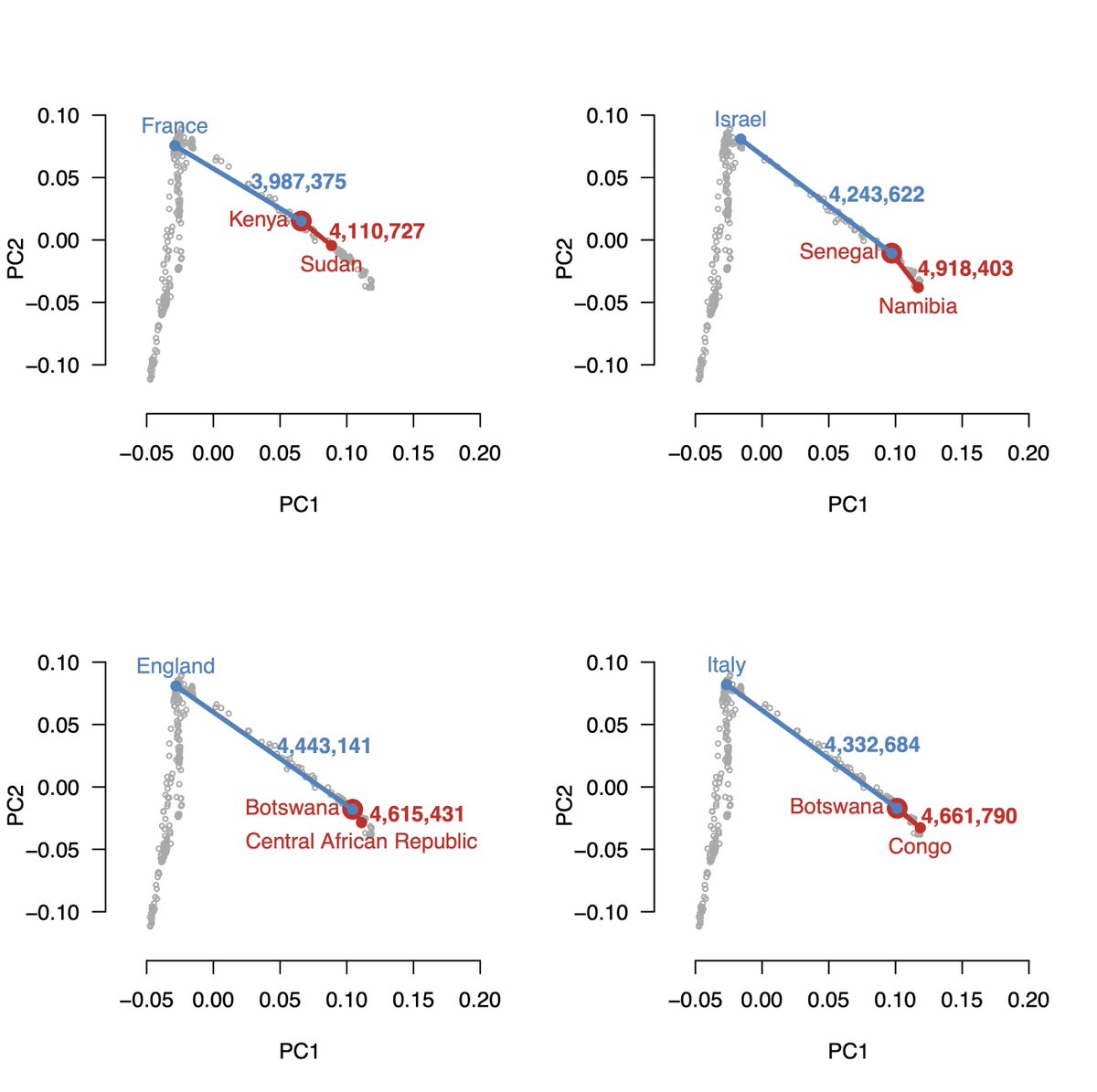
At @herasight, we wanted to compare our genetic predictors (PGS) to those from @nucleusgenomics. However, in many cases, we couldn’t reconcile plausible performance of their PGSs with customer risk reports we saw — this may have misled customers about their disease risks.

virtual cells are currently bottlenecked by compute, not novel data: drug discovery is an iterative search process (design, test, analyze) through therapeutic design space guided by a dynamics model directly trawling this therapeutic space with large hypothesis-free perturbational screens is an incredibly inefficient, expensive means of doing this search. the combinatorial space of cell states x perturbations is too large to brute-force it is even more of a fool's errand to run these screens solely for the purpose of generating training data for the dynamics model (i.e. "virtual cell") we'll use to navigate therapeutic design space rather, a general cellular dynamics model (there will be one model to rule them all) is most cost- and time-efficiently pretrained on large, diverse observational datasets of majority healthy cells, not perturbational atlases of diseased cells. the icing and cherry on top are useless if you haven't baked the cake we have plenty (petabytes) of this observational data already and are currently FLOP-constrained, not novel biological data (bioFLOP) constrained. therefore, we'd be better served spending on pretraining compute, not assaying millions of single cells we are doubly FLOP-constrained because the future is scaling up inference-time compute running in silico experiments on our mechanistically interpretable virtual cell, in order to select the most promising targeted perturbational experiments to run in the wet lab the compute demands of this inference-time experimental search will far exceed those of pretraining the virtual cell this virtual hypothesis-driven approach will direct us toward the regions of cell state x therapeutic design space where collecting perturbational data has the highest return. rather than trawl, we will precision-guided spearfish this is the only way to efficiently search therapeutic design space, using FLOPs to better allocate bioFLOPs