Benjamin J. Buchfink
178 posts

Benjamin J. Buchfink
@bbuchfink
Independent computer scientist. Developer of the DIAMOND protein aligner.
Tübingen, Deutschland Katılım Ağustos 2014
1.3K Takip Edilen577 Takipçiler
Benjamin J. Buchfink retweetledi

🦠🧬🖥️ Bakta v1.12.0 is out
with tons of tiny improvements and bug fixes, too many to list all:
- partial genes on linear seqs
- improved errror handlings & runtimes
- support Python 3.12 & 3.13
- ...
Thank you to all bug reporters and contributors!
github.com/oschwengers/ba…
English

DIAMOND v2.1.17 has new output fields sRANK to print taxonomy nodes of the given rank associated with the subject sequence, where RANK can be any rank in the NCBI taxonomy, e.g. sdomain, skingdom, sphylum, sorder, sgenus, sspecies, etc. github.com/bbuchfink/diam…
English
Benjamin J. Buchfink retweetledi

🚨New preprint out!
We present a foundational genomic resource of human gut microbiome viruses. It delivers high-quality, deeply curated data spanning taxonomy, predicted hosts, structures, and functions, providing a reference for gut virome research (1/8) biorxiv.org/content/10.110…

English
Benjamin J. Buchfink retweetledi

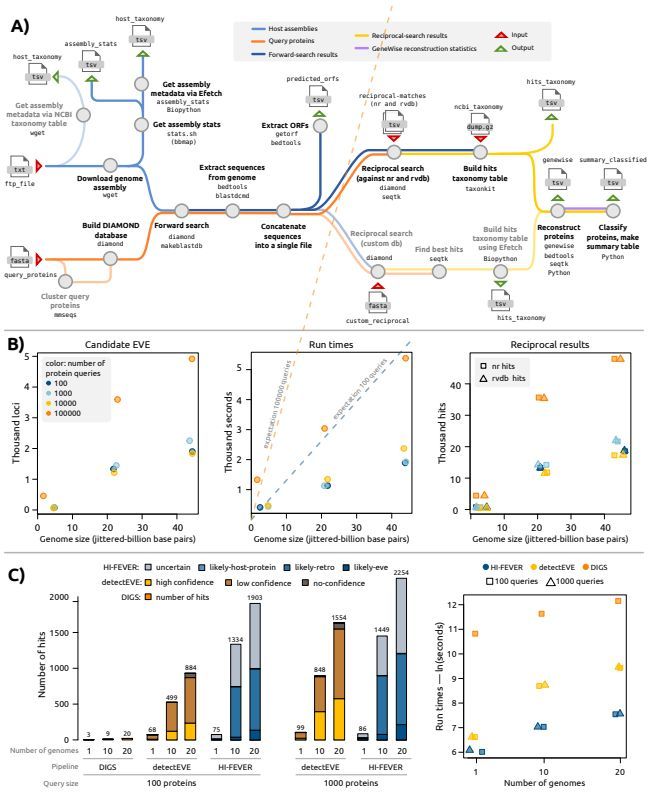
HI-FEVER: a Nextflow pipeline for the high-throughput discovery and annotation of endogenous viral elements. #EndogenousViralElements #EukaryoticHostGenomes #Genomics #Bioinformatics
academic.oup.com/bioinformatics…
English
Benjamin J. Buchfink retweetledi

Sup' #python #bioinformatics ! Didn't announce it so far, but now that it's working, I'm pleased to say you can now pip install BLAST+ (and more)! Technicalities below🧶 1/12

English

DIAMOND v2.1.15 now supports all taxonomy features for BLAST databases, and support for using BLAST databases has also been added to the Bioconda version github.com/bbuchfink/diam…
English
Benjamin J. Buchfink retweetledi
Benjamin J. Buchfink retweetledi

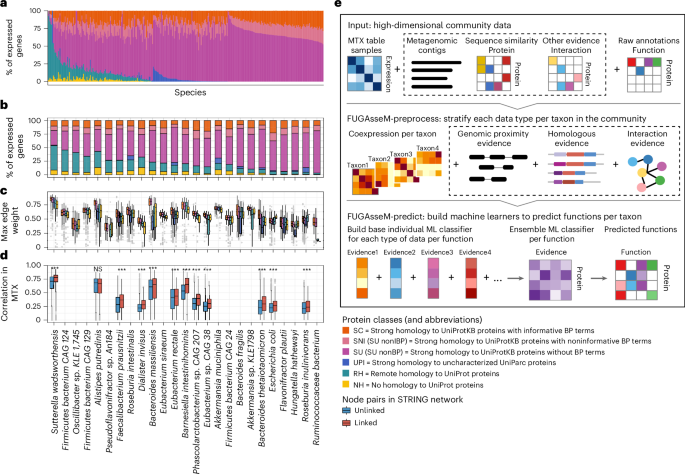
Predicting functions of uncharacterized gene products from microbial communities go.nature.com/435C4lp
English
Benjamin J. Buchfink retweetledi

Ragnarok: a flexible and RApid GeNe Annotation (ROcKs) pipeline deployed through Nextflow biorxiv.org/content/10.110… #biorxiv_bioinfo
English
Benjamin J. Buchfink retweetledi

CluSeek: Bioinformatics Tool to Identify and Analyze Gene Clusters biorxiv.org/content/10.110…

English
Benjamin J. Buchfink retweetledi

New preprint shows that DALI, TM-align and Foldseek overestimate protein structure alignment significance by 5-6 orders of magnitude
biorxiv.org/content/10.110…](biorxiv.org/content/10.110…
English
Benjamin J. Buchfink retweetledi

TaxTriage: An Open-Source Metagenomic Sequencing Data Analysis Pipeline Enabling Putative Pathogen Detection. #Metagenomics #Sequencing #PathogenDetection #Bioinformatics @biorxiv_bioinfo 🧬 🖥️
biorxiv.org/content/10.110…

English
Benjamin J. Buchfink retweetledi

It has been a while… but an updated, faster & more accurate version of OrthoFinder is now out!
Scales to thousands of species on conventional compute resources with higher accuracy than ever before and the same data rich fully phylogenetic outputs
👉biorxiv.org/content/10.110…
English
Benjamin J. Buchfink retweetledi

Out in @NatureBiotech: Metagenome taxonomy profilers usually ignore unknown species. SingleM is an accurate profiler which doesn't, even detecting phyla with no MAGs. Profiles of 700,000 metagenomes at sandpiper.qut.edu.au. doi.org/10.1038/s41587… A 🧵

English
Benjamin J. Buchfink retweetledi

We have a new version of OrthoFinder:
biorxiv.org/content/10.110…
Faster scaling to larger species sets, phylogenetically resolved orthogroups, reduced RAM usage.
Thanks to @dr_belcher, Yi Liu & Jonathan Holmes who have joined the team, and as ever OrthoFinder's PI @Steve__Kelly!
English

DIAMOND v2.1.12 provides support for the new NCBI taxonomic ranks and various other fixes. github.com/bbuchfink/diam…
English
Benjamin J. Buchfink retweetledi
Benjamin J. Buchfink retweetledi

Our proposal supported lab automation software infrastructure, with backing from major robotics companies. It was due in a week. Pure idiocy.
grants.nih.gov/grants/guide/r…
English
Benjamin J. Buchfink retweetledi

The saddest thing about this is academic science doesn’t reward software development, and many fields have serious tech debt that’s a huge drag on scientific productivity.
Emma J Chory@chorye
NIH just early-expired its 1st & only software NOFO: “Building Sustainable Software Tools for Open Science.” I guess “sustainable” is too risky now. So much for funding the STEM workforce.
English