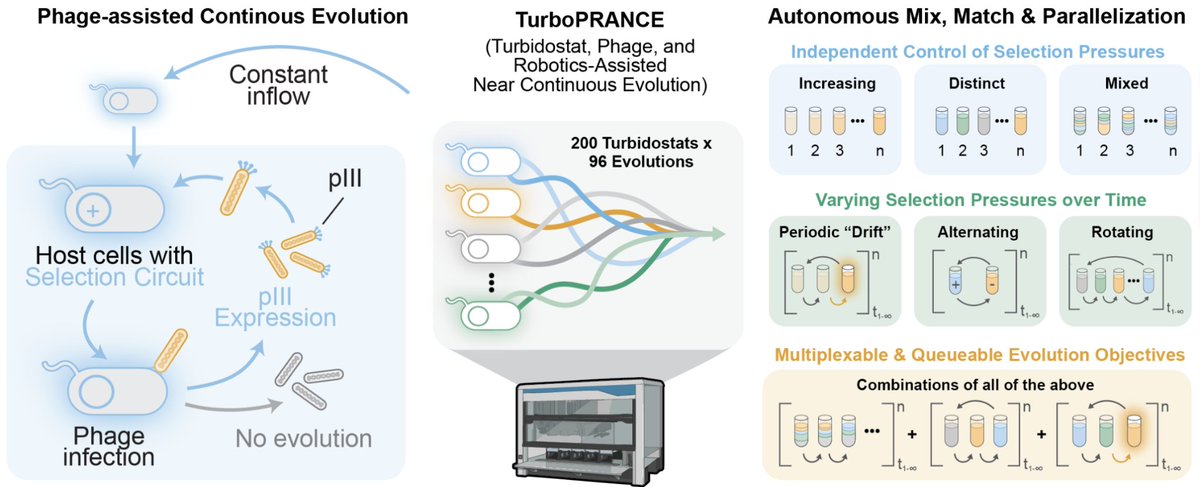
Aaaand it’s online ahhhhh!!! 🥳🥳 So excited!! The first glimpse of my postdoc work with @chorye @dukecagt. Here, @stefanmgolas and I developed TurboPRANCE, an open-source robotics platform for rapid and scaled phage-assisted continuous evolutions. 🧪Tweetorial party!👇1/n