Arjun Krishnan
1.1K posts

Arjun Krishnan
@compbiologist
ML & data-driven discovery; Complex traits & diseases; Data reuse & open science. Associate Professor | Group leader @KrishnanLab | @compbiologist everywhere




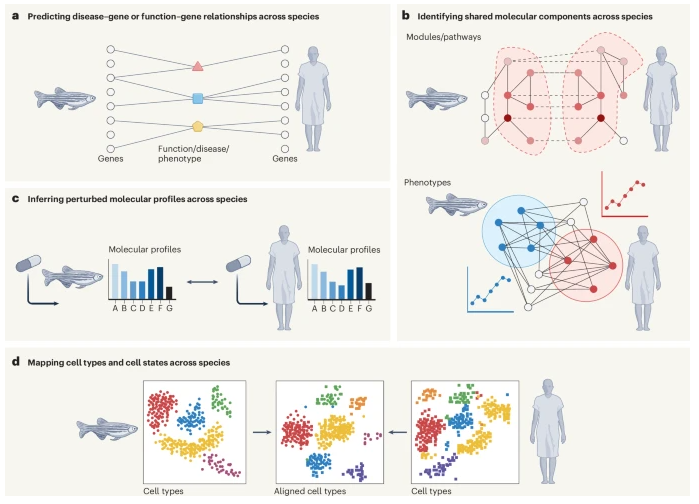



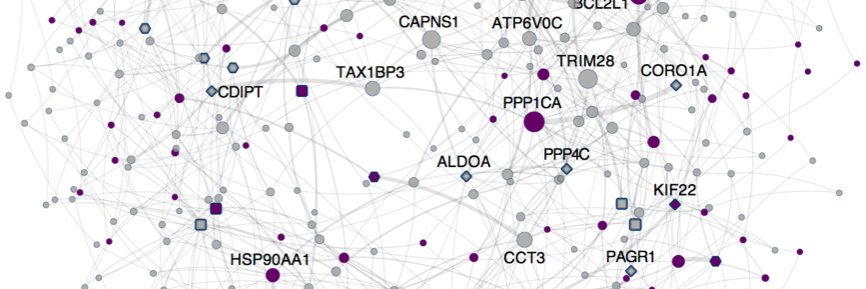
Preprint 🚨 A review state-of-the-art computational strategies for cross-species knowledge transfer in biomedicine 💻👩🦰🐭🐟🪰🪱🧬🫁⚕️ Led by an excellent team at @KrishnanLab: @yhbioinfo, @ChrisAMancuso, & @kaylainbio in collab w/ @FishEvoDevoGeno 🧵 arxiv.org/abs/2408.08503

Remember, mixing up Type I and Type II errors is called a Type III error

Understanding Type I and Type II errors is the secret to unlocking the full potential of your statistical analysis. These errors are pivotal in hypothesis testing, where Type I errors represent false positives (incorrectly rejecting a true null hypothesis) and Type II errors represent false negatives (failing to reject a false null hypothesis). Handling these errors effectively can greatly improve the accuracy and credibility of your analyses. By meticulously managing these errors, you can ensure your statistical conclusions are both reliable and valid, ultimately leading to more trustworthy and impactful research findings. Cons of Mismanaging Type I and Type II Errors: ❌ Misleading Results: High rates of Type I errors can result in false claims of significance, leading to incorrect conclusions. ❌ Missed Discoveries: Excessive Type II errors can cause important findings to be overlooked, as genuine effects are dismissed as insignificant. ❌ Reduced Trust: Frequent errors undermine the credibility of your analysis, leading to mistrust in your results and decisions. Pros of Effectively Managing Type I and Type II Errors: ✔️ Minimized False Positives: By carefully setting thresholds, you can reduce the number of false positives, ensuring that positive results are genuinely significant. ✔️ Accurate Conclusions: Proper management of Type I and Type II errors helps draw more accurate conclusions from data, enhancing the overall validity of your study. ✔️ Improved Decision-Making: With fewer errors, the decisions based on your data will be more reliable and informed. To manage Type I and Type II errors effectively in practice: 🔹 R: Use the p.adjust function from the stats package to control for multiple comparisons and reduce Type I error rates. 🔹 Python: Utilize the statsmodels library, specifically the multipletests method, to adjust p-values and maintain control over error rates. The visualization originates from a wikipedia image (link: en.wikipedia.org/wiki/Type_I_an…) and shows the results of negative samples (left curve) overlapping with positive samples (right curve). Adjusting the cutoff value (vertical bar) helps balance false positives (FP) and false negatives (FN), impacting the rates of true positives (TP) and true negatives (TN). To explain this topic in further detail, I collaborated with Micha Gengenbach to create a comprehensive tutorial: statisticsglobe.com/type-i-and-typ… Eager to advance your skills in statistics and R programming? My online course, "Statistical Methods in R," might be ideal for you. More details are available at this link: statisticsglobe.com/online-course-… #statisticians #DataScience #DataAnalytics #RStudio #RStats #database


Preprint 🚨 A review state-of-the-art computational strategies for cross-species knowledge transfer in biomedicine 💻👩🦰🐭🐟🪰🪱🧬🫁⚕️ Led by an excellent team at @KrishnanLab: @yhbioinfo, @ChrisAMancuso, & @kaylainbio in collab w/ @FishEvoDevoGeno 🧵 arxiv.org/abs/2408.08503

Annotating Publicly-Available Samples and Studies Using Interpretable Modeling of Unstructured Metadata 1. This study introduces txt2onto 2.0, an improved NLP and ML-based tool that automates the annotation of unstructured biomedical metadata, linking samples and studies to controlled disease and tissue vocabularies without manual intervention . 2. By using a TF-IDF-based feature extraction approach instead of averaging word embeddings, txt2onto 2.0 offers more interpretable results, allowing it to accurately identify key predictive terms within sample and study metadata . 3. The model outperforms its predecessor in both tissue and disease annotation tasks, excelling particularly in scenarios with limited training data, thus making it ideal for infrequent or rare biomedical terms . 4. A notable strength of txt2onto 2.0 is its ability to work across different biomedical text sources (e.g., GEO, PRIDE, ClinicalTrials), providing consistent annotations by capturing meaningful semantic relationships even with unseen terms . 5. The interpretability of txt2onto 2.0 is highlighted through word clouds of predictive terms, where it captures domain-specific keywords without requiring explicit mentions of target terms, showcasing its robustness and potential to adapt to new datasets . 6. This tool’s transparent prediction process and scalability support its application across various data repositories, advancing the FAIR data principles (Findable, Accessible, Interoperable, Reusable) in biomedical research . @compbiologist 💻Code: github.com/krishnanlab/tx… 📜Paper: doi.org/10.1101/2024.0… #BiomedicalNLP #DataAnnotation #MachineLearning #FAIRdata #ComputationalBiology





Interested in learning more about the Human Medical Genetics and Genomics Program at CU Anschutz? Join us at 1PM Mountain on Monday, October 21, for the 2024 BIOMED PHD EXPO!






SfN is proud to announce the recipients of the 2024 Promotion of Women in Neuroscience awards, who will be honored at #SfN24. Through mentorship, professional development and outstanding careers, these seven researchers have made significant contributions to the advancement and inclusion of women along the length of the research pipeline paving the way for a more inclusive and impactful field of neuroscience. Learn more about the recipients and their remarkable contributions to #neuroscience. 🔗 bit.ly/3TA00s2 The Bernice Grafstein Award for Outstanding Accomplishments in Mentoring is supported by Bernice Grafstein, PhD. #NeuroTwitter #AcademicTwitter #SciTwitter








