Erna Valdís Ívarsdóttir retweetledi

Another incredible GWAS finding this year: an African ancestry–specific missense variant confers the largest common-variant risk effect reported to date for SLE.
Just weeks ago, there was a GWAS paper reporting the discovery of a major genetic risk factor for dilated cardiomyopathy--an African ancestry-specific loss of function variant in CD36--explaining up to 8% of the cases in African populations. (x.com/doctorveera/st…).
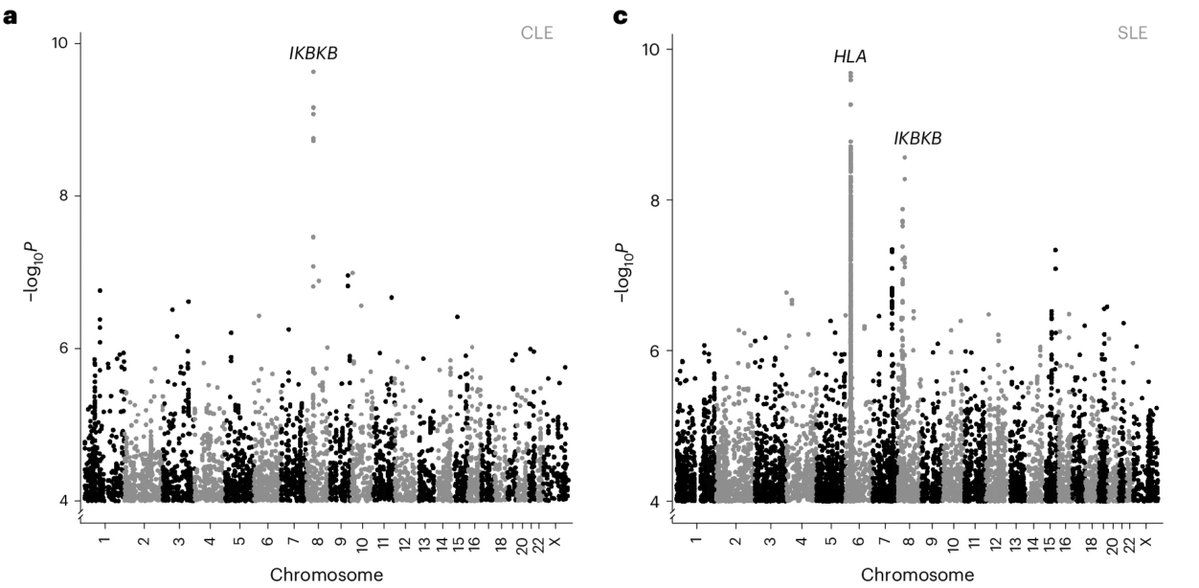
Now, deCODE Genetics has uncovered a major genetic risk factor for lupus, a missense variant p.Glu502Lys in IKBKB, explaining 10.4% of cutaneous lupus (CLE) and 6.4% of cases of systemic lupus (SLE) in African populations.
The variant increases CLE risk by 5.4 fold, uncovered by comparing just 211 cases to 25,360 controls of African ancestry from the Alliance for Genomic Discovery (AGD). It highlights how deeply African ancestries have been underrepresented in lupus GWAS efforts.
Lupus is more common (2-3x) and more severe in individuals of African ancestry compared with those of European ancestry. Certain forms, like discoid lupus, are 10 times more common in African populations. Despite that, only one of the past 33 SLE GWASs involved African-ancestry population!!
Interestingly, that one study (Langefeld et al. Nat Comm 2017) that involved African ancestry individuals actually captured this IKBKB locus (connecting the signal to a different nearby gene, PLAT), but they seemed to have never identified the right gene and overlooked the importance of the finding.
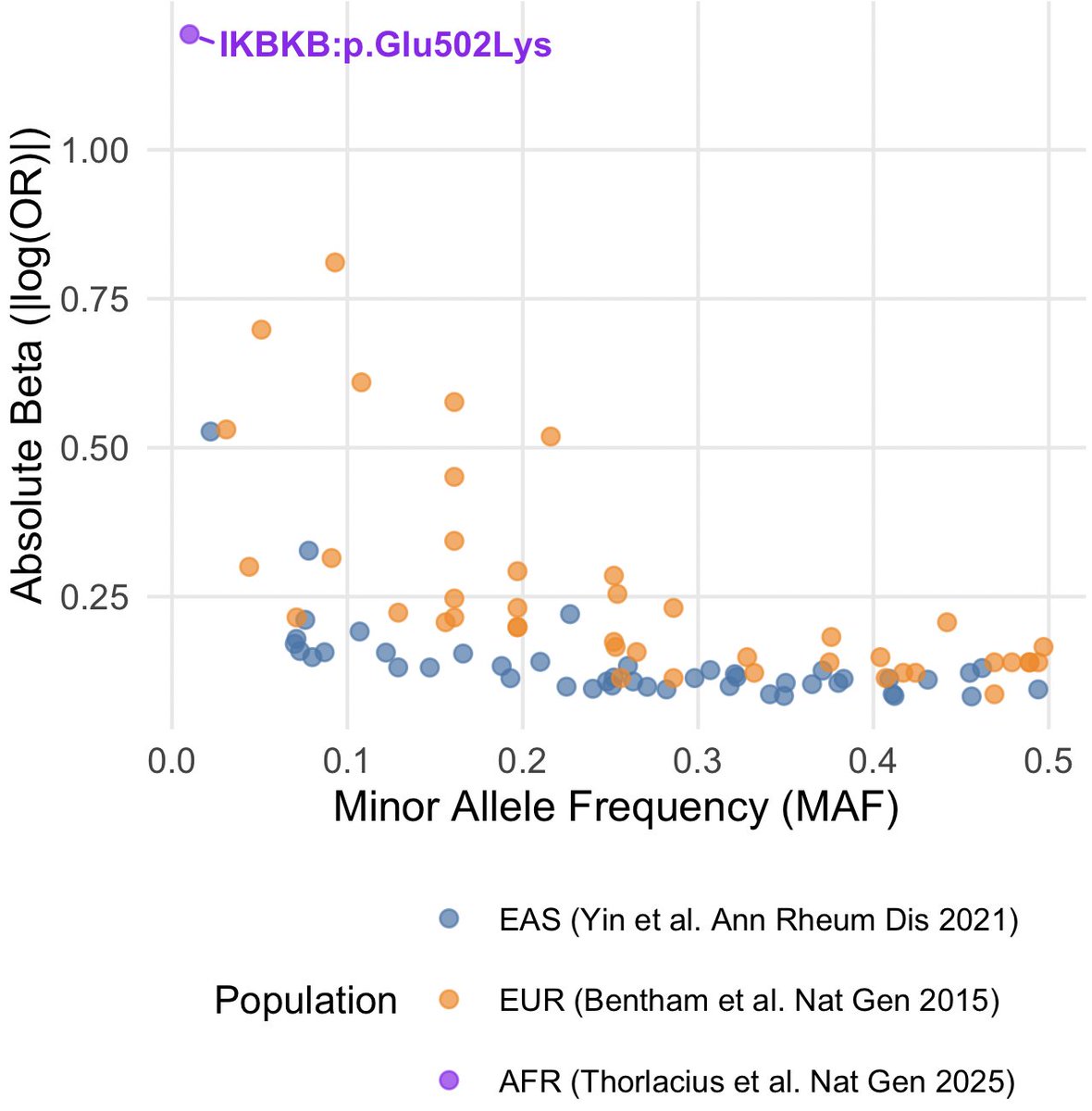
Here is a plot I roughly made to highlight the effect size of this variant. The plot compares the effect sizes vs minor allele frequency of GWAS loci reported the past large SLE studies in European (Bentham et al. Nat Gen 2015) and East Asian ancestries (Yin et al. Ann Rheum Dis 2021). I overlayed the IKBKB missense variant to show the it's effect size for SLE dwarfing even the largest effect sizes reported for HLA variants.
The allele frequency of this missense variant in African ancestries is ~1%. Around 2% of the African ancestry individuals are carriers, and among the SLE/CLE cases, more than 10% carry this variant. Amazing!
Analyzing related clinical phenotypes, the authors found that the carriers tend to have a more severe phenotype (more CLE, more glomerular involvement, proteinuria and lower blood counts).
The gene IKBKB encodes an immune protein that activates the NF-κB signaling, a key immune response pathway. Partial or complete loss of IKBKB cause Mendelian forms of immunodeficiency conditions. The missense variant likely confers a gain of function effect leading to autoimmunity.
This discovery highlights the importance of NF-κB signaling in lupus pathogenesis, particularly in individuals of African ancestries. The finding has many translational implications, for example, it suggests a large target population (~10% of African cases) for any NF-κB targeted SLE treatment.
Another great work by deCODE scientists!
Thorlacius et al. Nat Gen 2025

Patrick Sulem@patsule
Congratulations to my colleagues Gudny Ella and Erna. Studying diverse ancestries as part of a large collaboration enables the discovery of association, with lupus, of an IKBKB missense quite specific to individuals with African ancestry. @NatureGenet nature.com/articles/s4158…
English