Patrick Sulem
478 posts

Patrick Sulem
@patsule
MD, Co-managing director deCODE genetics - Amgen


















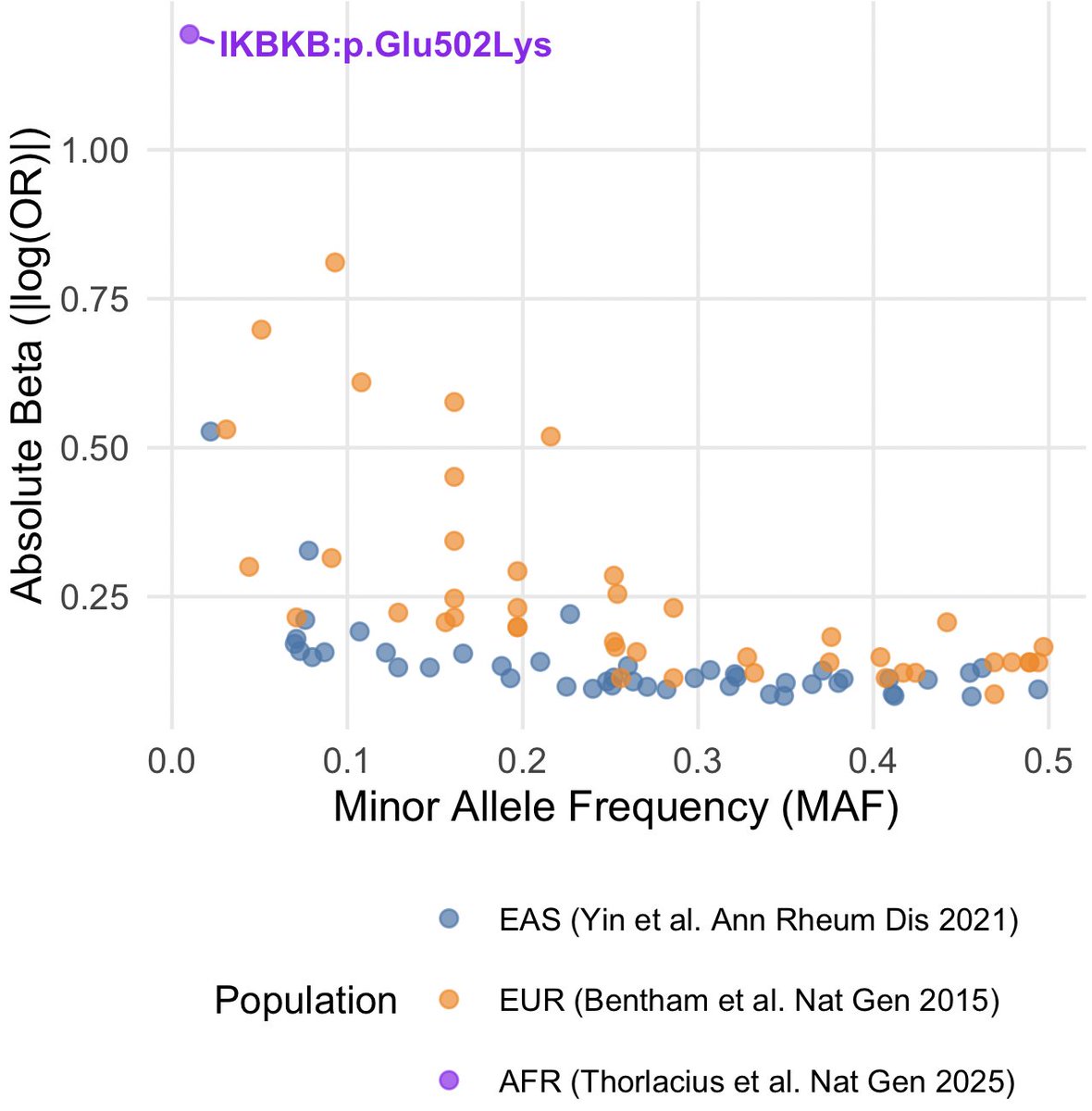
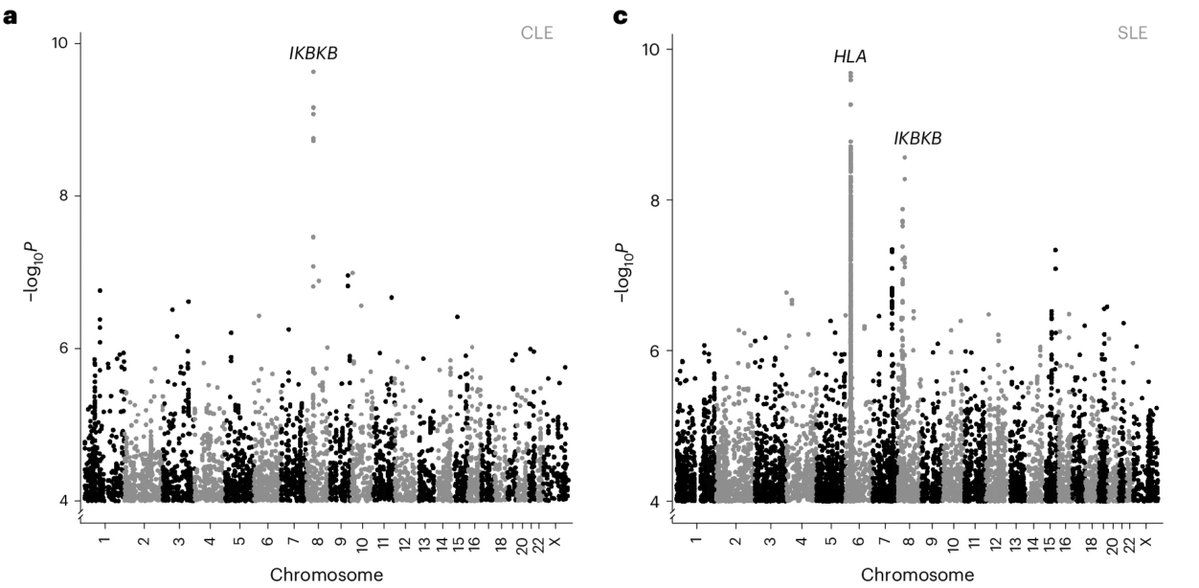

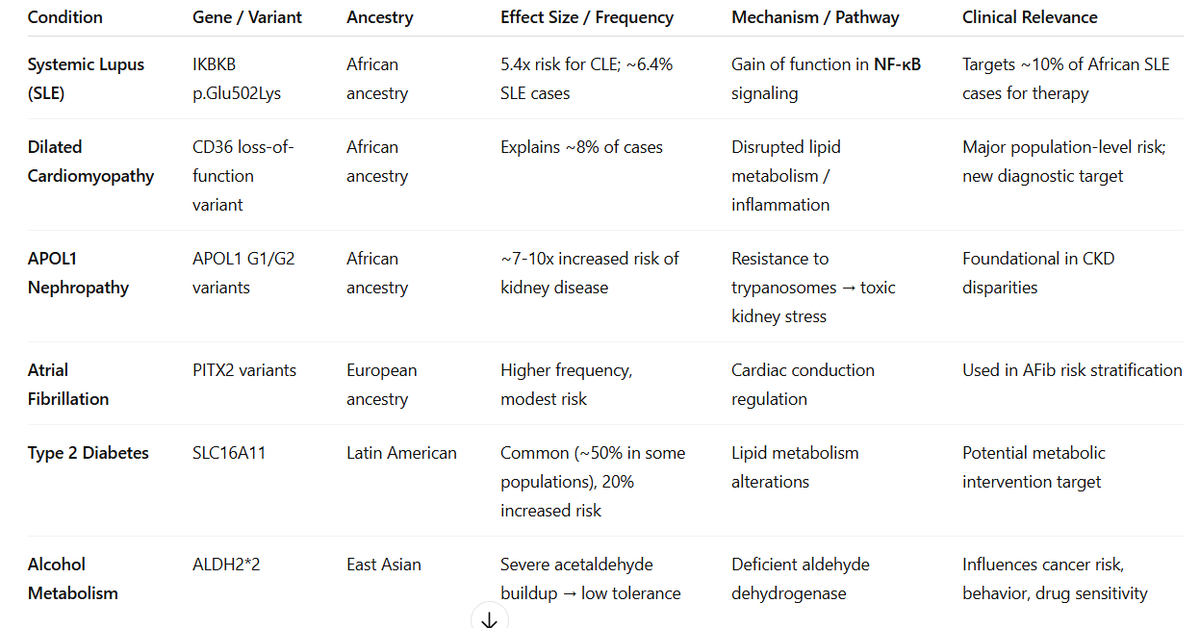
Congratulations to my colleagues Gudny Ella and Erna. Studying diverse ancestries as part of a large collaboration enables the discovery of association, with lupus, of an IKBKB missense quite specific to individuals with African ancestry. @NatureGenet nature.com/articles/s4158…








Today in @NatureGenetics, we report the discovery of a common CD36 variant specific to individuals of African ancestry that increases risk of DCM, and may help explain the ~2x higher rate of DCM/heart failure in this population nature.com/articles/s4158…










Today, @NatMetabolism published new preclinical research in mice around blocking GIPR and activating GLP-1R. 🧬🔬 This study was led by @DanielJDrucker, Endocrinologist and Professor at @UofT, and Murielle Véniant-Ellison, Scientific VP at Amgen leading #obesity research. 🔗 Learn more: amgen.ly/44e09Yz