erva retweetledi

🚀 Exciting news! Our project, CROssBARv2, is now live as an online tool and its article (preprint)!
We've built a unified AI-powered platform that brings together the fragmented world of biomedical data — and lets you talk to it in plain English.
Here's the story 👇
The problem, in plain terms: Imagine being a detective, but your clues are scattered across 34 different filing cabinets, written in different languages, with no index. That's what biomedical researchers face every day — genes in one database, drugs in another, diseases somewhere else. Connecting the dots is slow & painful without serious programming skills.
CROssBARv2 is our answer to this. 🧬💊🔬
What did we build?
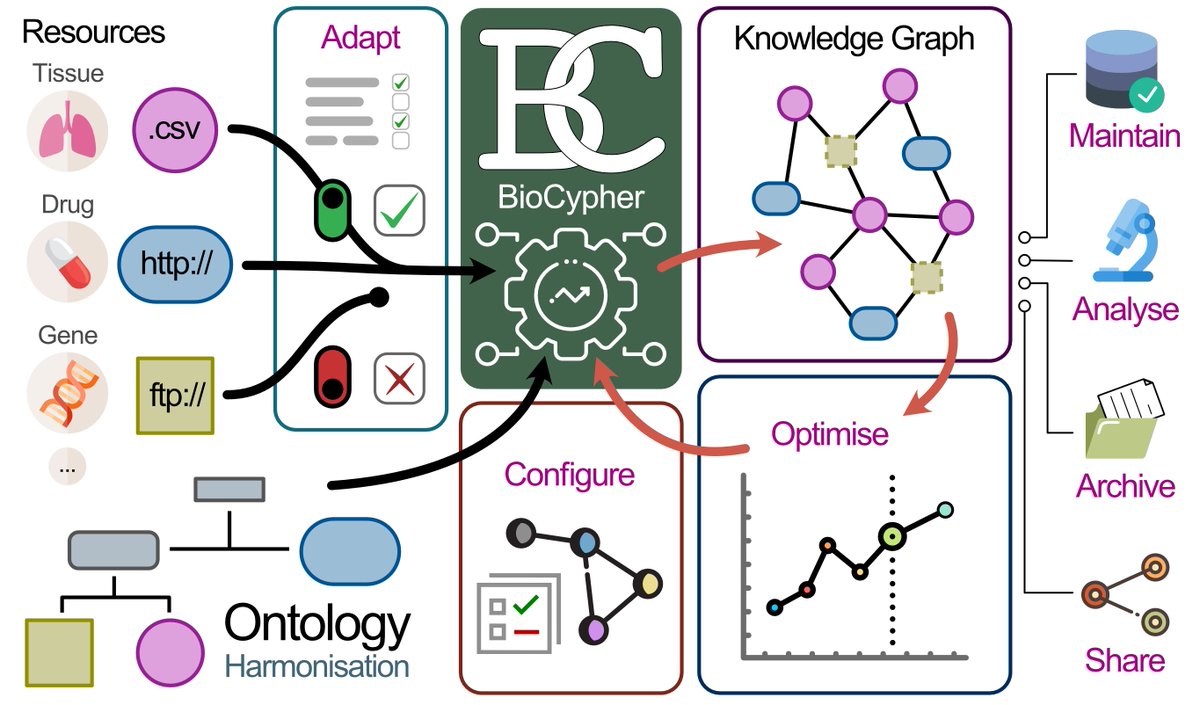
CROssBARv2 is a biomedical Knowledge Graph (KG) — think of it as a giant, intelligent map of biology. We integrated data from 34 well-established databases into a single, structured, and searchable system:
🔵 ~2.7 million nodes (proteins, genes, drugs, diseases, pathways, side effects, & more)
🔗 ~12.6 million edges (biomedical relationships)
🧠 14 node & 51 edge types
🏷️ Rich metadata
Now the exciting part 🤖 — meet CROssBAR-LLM
Large Language Models (LLMs) like ChatGPT or Gemini are brilliant at conversation — but they hallucinate. They confidently tell you things that sound right but aren't. In biomedicine, that's dangerous.
Our solution: ground the LLM in the knowledge graph.
CROssBAR-LLM lets you ask complex biomedical questions in plain English, like:
"What proteins are encoded by genes regulated by NFKB1, participate in the Endocytosis pathway, and are targeted by drugs used to treat diseases comorbid with osteoporosis?"
CROssBAR-LLM tutorial: youtu.be/FjuoKAmzjNM
What happens under the hood:
🗣️ Your natural language question is translated into a formal DB query
⚡ The query runs on the CROssBARv2 KG in real time
📊 Structured, verified results are retrieved
💬 The LLM turns results into a clear, readable answer
No hallucinations. Every answer is traceable back to real data
How does it compare to just directly asking LLMs on a biomedical Q&A benchmark?
GPT/Claude/Gemini: ~50-65%
CROssBAR-LLM (w/ Gemini 1.5 Pro): 98% accuracy 🎯
CROssBAR-LLM (w/ GPT-4o): 97% accuracy 🎯
Everything is open and accessible 🔓
🌐 Web platform: crossbarv2.hubiodatalab.com
📡 GraphQL API
🖥️ Neo4j Browser for visualisation
📂 Full data on Hugging Face & Google Drive
💻 All code on GitHub
No paywall. No programming required. Just ask your question!
Huge thanks to the team!
Bünyamin Şen, Erva Ulusoy @ervaulusy , Melih Darcan, Mert Ergun, Sebastian Lobentanzer, Ahmet S. Rifaioğlu @ahmet_rifaioglu , Dénes Türei, Julio Saez Rodriguez @JulioSaezRod , and me, in collaboration with Saez Lab @saezlab
spanning Hacettepe University @Hacettepe1967 , Heidelberg University @HeidelbergU , Helmholtz Munich @HelmholtzMunich , and European Bioinformatics Institute | EMBL-EBI @emblebi
📄 Read the preprint: biorxiv.org/content/10.648…
⚙️Try the tool: crossbarv2.hubiodatalab.com/llm
***Please repost!***
#AI #LLM #ML #Bioinformatics #KnowledgeGraph #DrugDiscovery #Biomedicine #OpenScience

YouTube
English