Sabitlenmiş Tweet

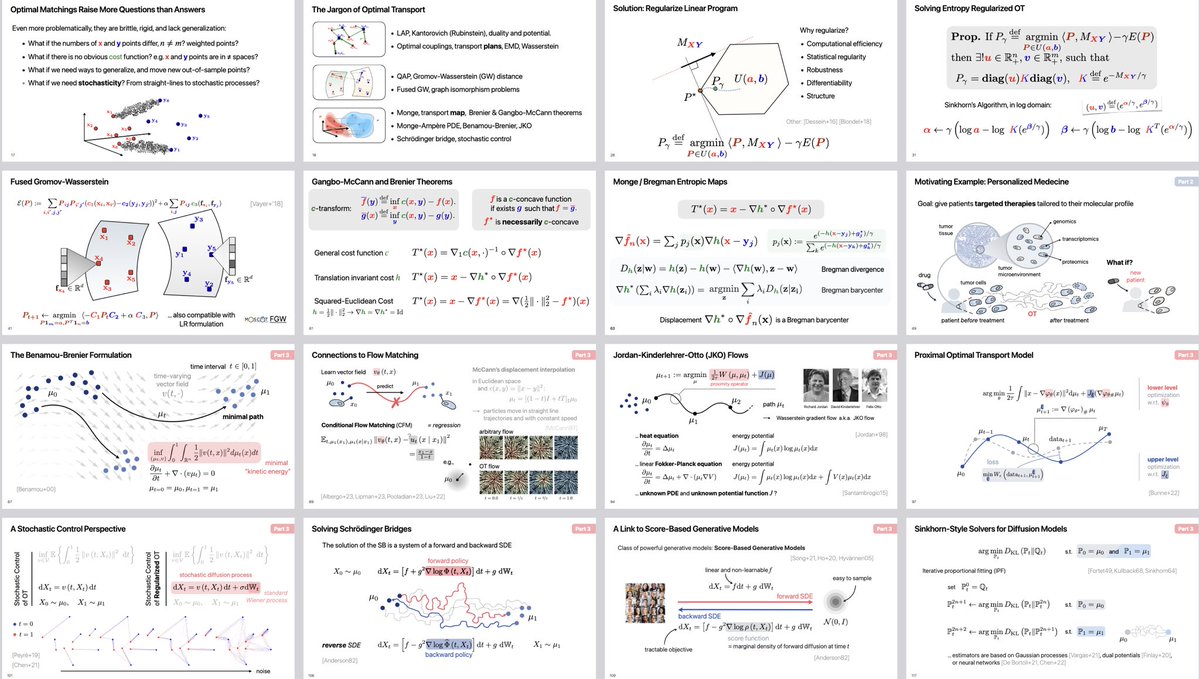
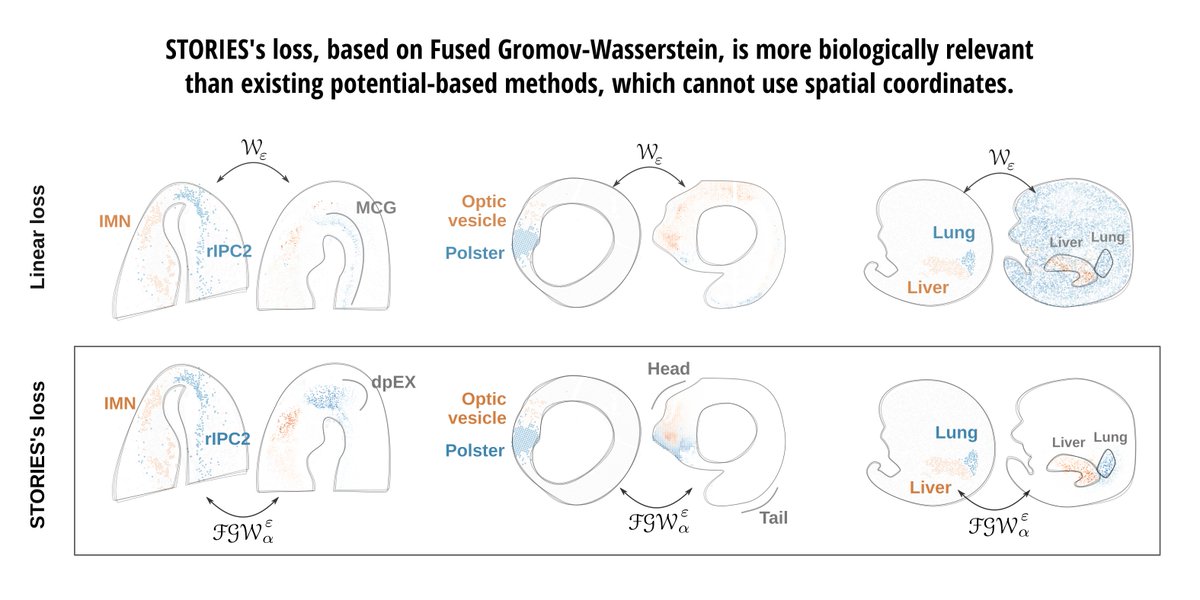
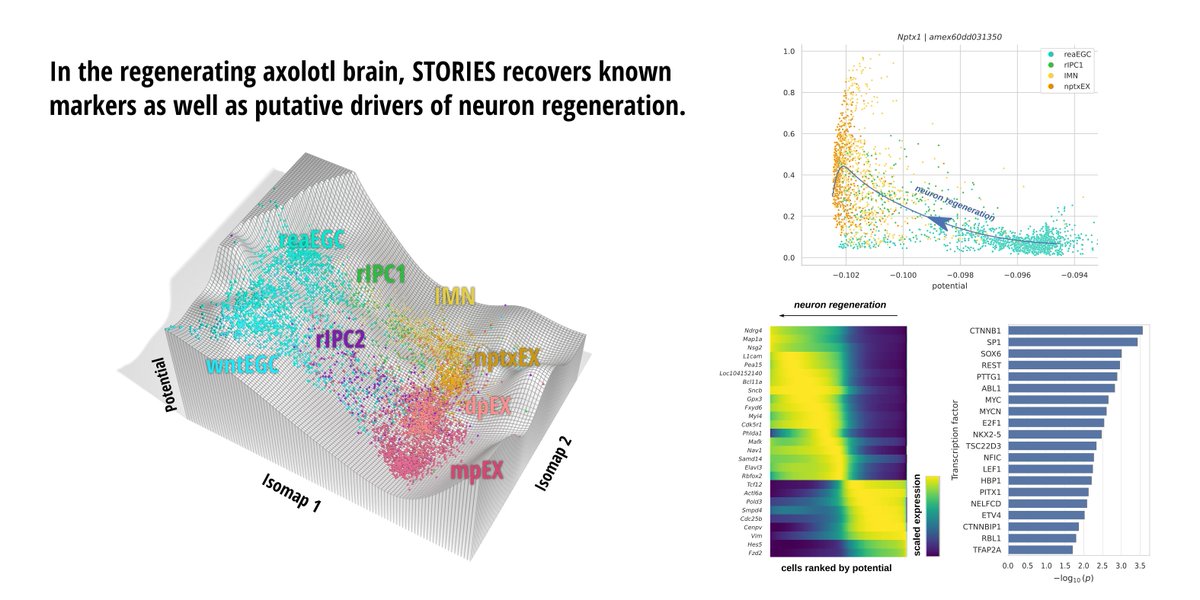
🎉 New preprint! biorxiv.org/content/10.110… STORIES learns a differentiation potential from spatial transcriptomics profiled at several time points using Fused Gromov-Wasserstein, an extension of Optimal Transport. @gabrielpeyre @LauCan88


English