Julien Roy retweetledi

I am particularly bullish on using mechanistic interpretability (especially SAEs) to better understand and discover new knowledge about biology and medicine.
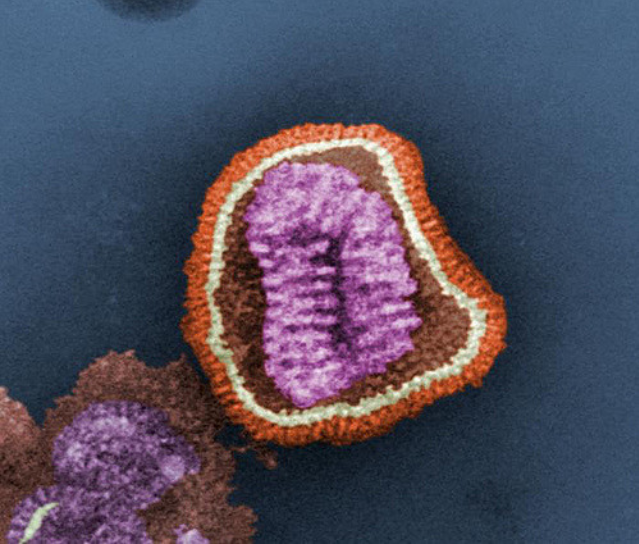
"Towards scientific discovery with dictionary learning: Extracting biological concepts from microscopy foundation models" is a recent paper by @valence_ai that demonstrated the use of mech interp to discover features that correspond to biologically relevant structures in a microscopy image.
A self-supervised foundation model (MAE) was trained on a large dataset of microscopy images of cell cultures that have undergone either knockout of specific genes or perturbations with small molecules.
A novel mech interp technique that is similar to SAEs called Iterative Codebook Feature Learning was applied to the MAE.
By examining which image patches exhibit the highest cosine similarities with specific feature directions, the authors demonstrated these features pick up specific biological concepts.
For example, the researchers identified a feature associated with the disruption of adherens junction proteins ("glue proteins" that allow cells to "stick" to each other). This feature is activated in parts of the microscopy image where there are small, bright and isolated cells which appear unable to establish proper connections with the neighboring cells.
This paper is just a very early demonstration and proof-of-concept, but I think there is significant promise of this approach:
1. collect lots of observations of cells/tissues/patients/etc. under different conditions to create a huge dataset
2. train large-scale self-supervised foundation models on such a dataset
3. Train mech interp models like SAEs on the foundation models and identify biologically/clinically-relevant features
4. Use such features to derive novel scientific and clinical insights!

English