Kenneth Loi
419 posts

Kenneth Loi
@kenjmloi
Decoding life's longest war Doudna Lab @UCBerkeley


Bridge recombinase enables versatile rewriting of bacterial genomes biorxiv.org/content/10.648… #biorxiv_synbio

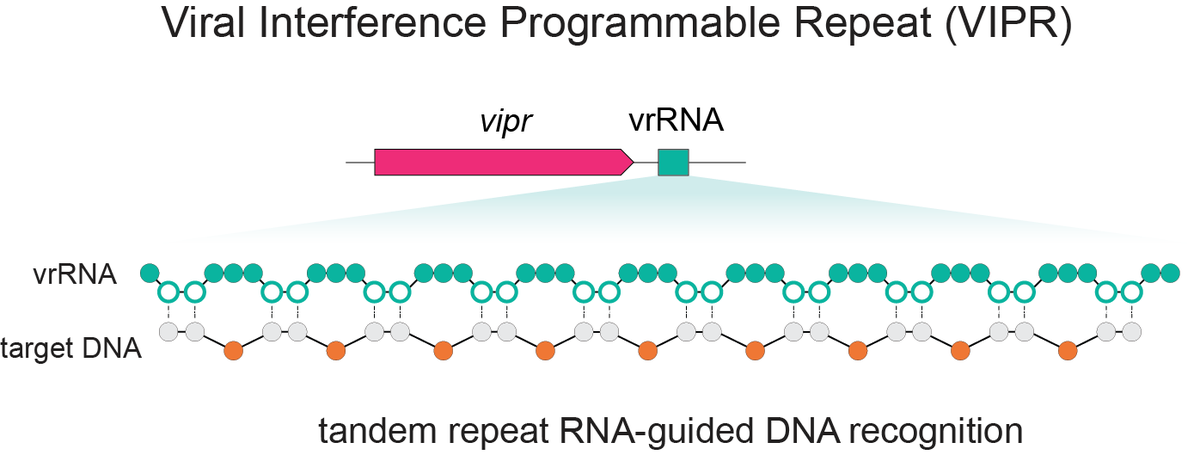
Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.


Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.


Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.

Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.

Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.

Excited to share our discovery of a new programmable RNA-guided DNA-targeting system hiding inside bacteriophages that predates CRISPR. We call it VIPR (Viral Interference Programmable Repeat), and it uses an entirely new logic to find its targets. Thread + link below.

A beautiful story on the origins of CRISPR -- congrats to @PeterHYoon @kenjmloi and team! Cool to see the Evo 2 likelihoods being useful for identifying a tricky repeat region. biorxiv.org/content/10.648…