
Daniel Chang
105 posts

@danielchang2002
Genetics PhD student @Stanford w/ @BrianHie | Prev @UMNComputerSci





Out now! In collaboration with @LeifuChangLab, we uncover the molecular and structural underpinnings of CRISPR-Cas12f-like RNA-guided transcription systems! Links to the articles in the following tweet:





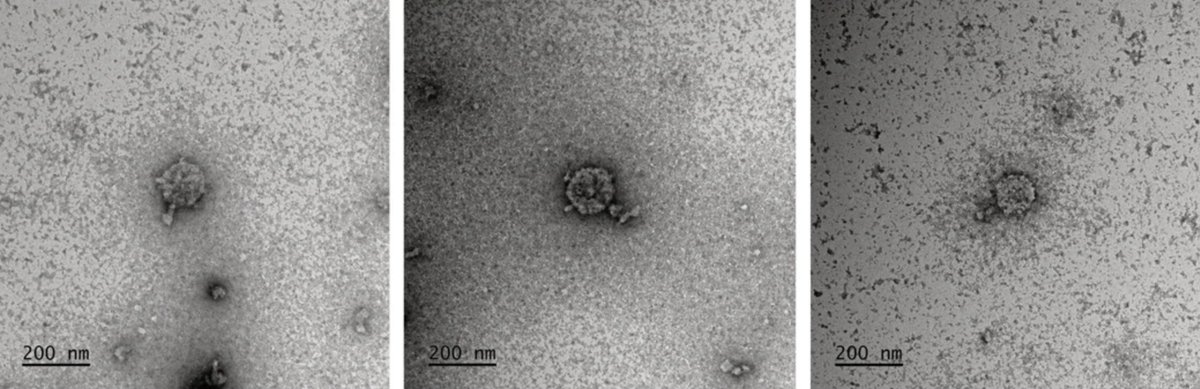
New work out from Bhatt lab led by Jakob Wirbel and @Angela_Hickey98 on uncovering gut prophage biology via long-read metagenomics! biorxiv.org/content/10.110… A small 🧵 highlighting the tale of IScream🍦phage (Fig. 4) which I was lucky to work on while rotating in the Bhatt lab!

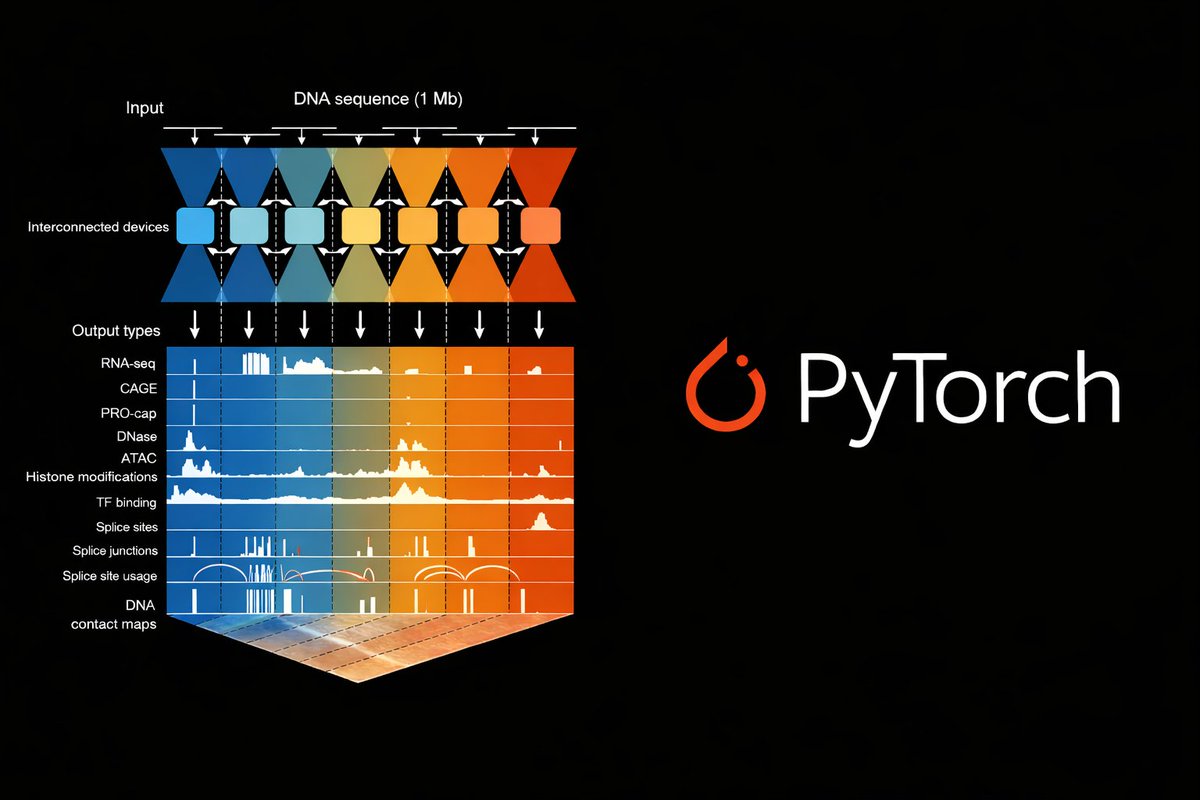
What if we could autocomplete DNA based on function? Today in @Nature, we share semantic design—a strategy for function-guided design with genomic language models that leverages genomic context to create de novo genes with desired functions.🧵 nature.com/articles/s4158…






Sliding window attention (SWA) is powering frontier hybrid models for efficiency. Is there something better? Introducing Phalanx, a faster and better quality drop-in replacement for sliding window attention (SWA). Phalanx is a new family of hardware and numerics-aware windowed layers designed with a focus on data locality and jagged, block-aligned windows that map directly to GPUs. In training, Phalanx delivers 10–40% higher end-to-end throughput at 4K–32K context lengths over optimized SWA-hybrids and Transformers by reducing costly inter-warp communication. Today, we are releasing both the technical report, a blog, and Phalanx kernels in spear, our research kernel library. We are hiring.

How do protein language models (PLM) think about proteins?🧬 We answer this w/ #InterPLM, just published in @naturemethods! Using sparse autoencoders + LLM agent, we identify 1000s of interpretable concepts learned by PLMs, pointing to new biology 🧵

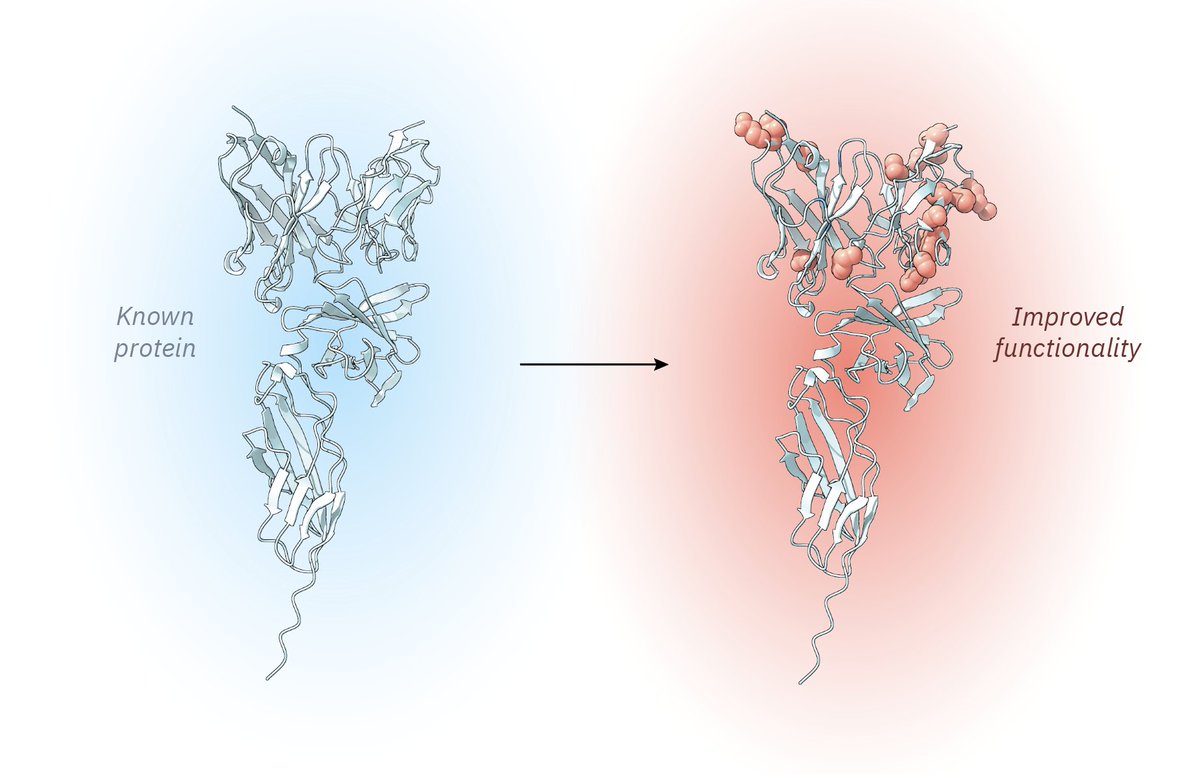
The ability to design antibodies against any protein of interest has major implications for medicine, biotech, and basic science. Today, we introduce Germinal, a pipeline for epitope-targeted de novo antibody design achieving 4–22% success rates with efficient experimental validation.

The ability to design antibodies against any protein of interest has major implications for medicine, biotech, and basic science. Today, we introduce Germinal, a pipeline for epitope-targeted de novo antibody design achieving 4–22% success rates with efficient experimental validation.