Sabitlenmiş Tweet

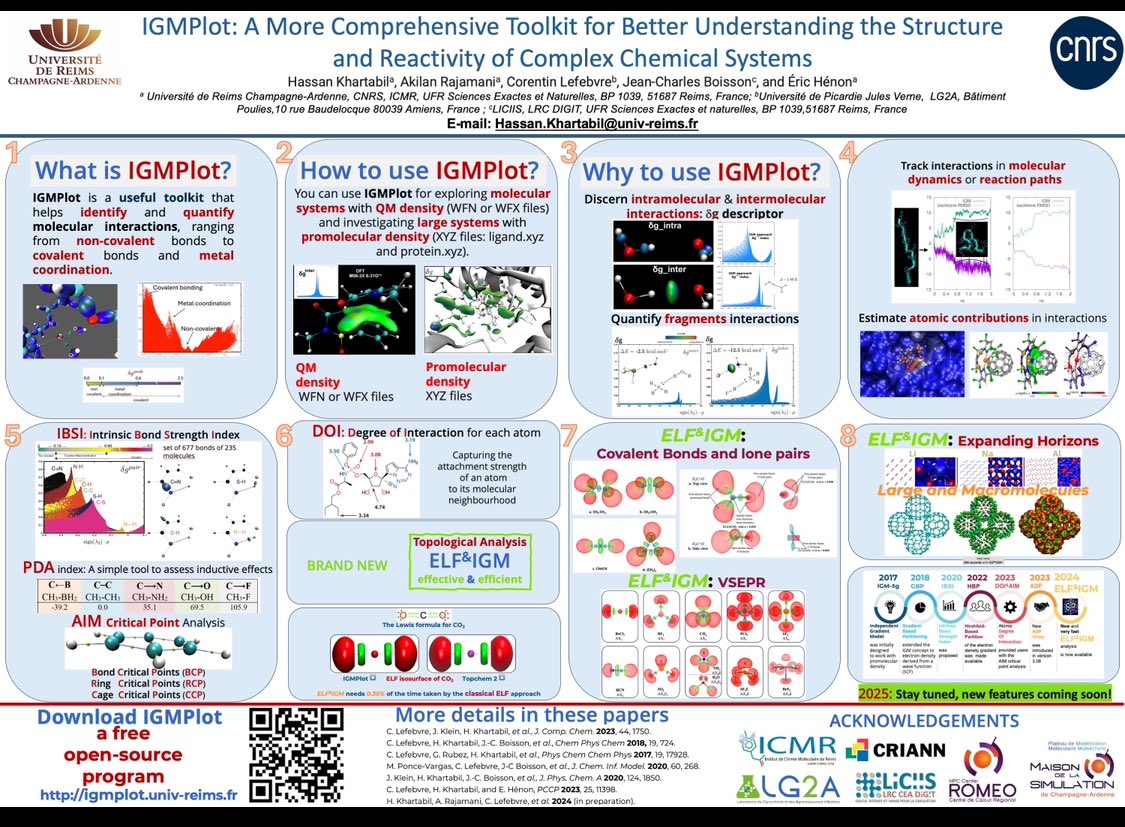
👋Hi @LatinXChem, presenting our work🖥️‘IGMPlot: A More Comprehensive Toolkit for Better Understanding the Structure and Reactivity of Complex Chemical Systems’🧬🧪⚗️⚛️at #LatinXChem24 #LatinXChemComp #Comp022 #CompChem #Chemistry #ElectronDensity #Descriptor #IGMPlot #ELF #DFT
English