Biology+AI Daily@BiologyAIDaily
Design of pseudosymmetric protein hetero-oligomers @NatureComms
🚀 New paper from David Baker!🚀
1. This study introduces a divide-and-conquer strategy for designing pseudosymmetric protein hetero-oligomers with multiple unique subunits, which are essential for creating complex biological systems and materials.
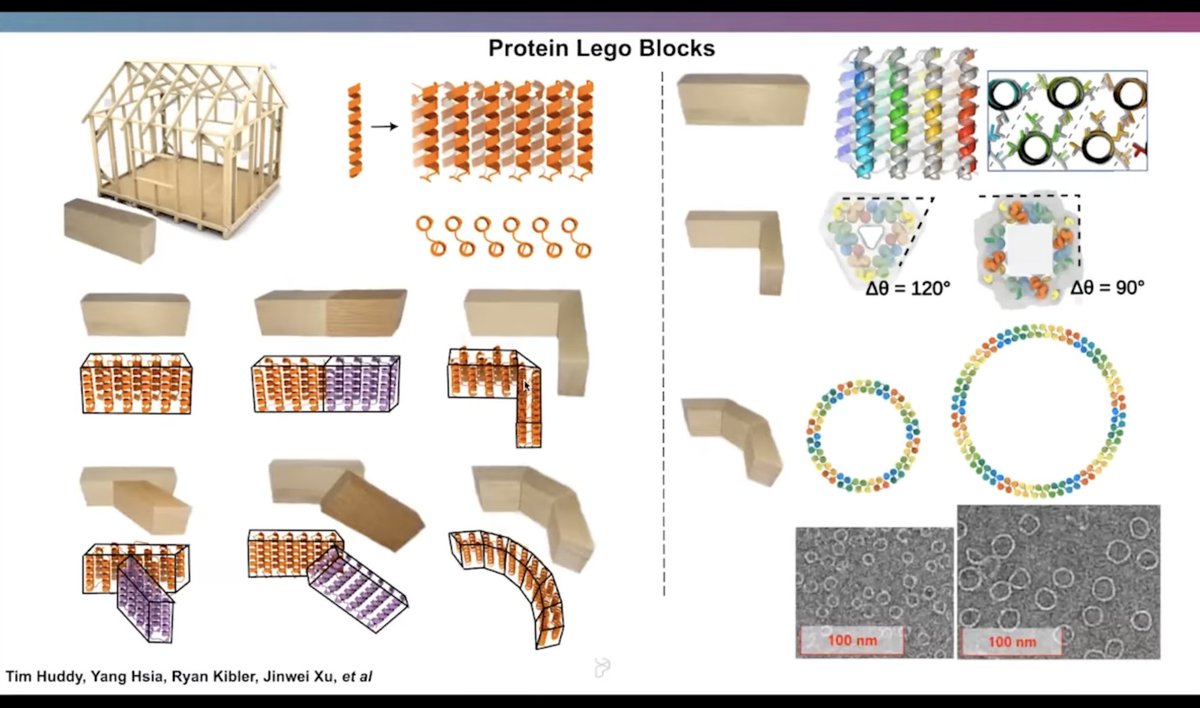
2. The method starts with de novo designed symmetric homo-oligomers, redesigning individual subunit interfaces using hydrogen bond networks (HBNets) to ensure specificity, then recombines these to form hetero-oligomers.
3. This approach was successfully applied to create ABC heterotrimers, A2B2 heterotetramers, and more complex structures like A2B2C2 heterohexamers, showcasing high specificity and structural integrity.
4. The pseudosymmetric assemblies created here provide a modular platform for constructing larger protein architectures, including functional nanocages, with applications in drug delivery, signaling, and materials science.
5. Evolutionary insights are mirrored in the design, using principles observed in natural pseudosymmetric complexes, like Sm and LSm protein assemblies, to enhance structural and functional diversity.
6. Experimental validation through structural and biophysical methods, including crystallography, SAXS, and mass spectrometry, confirms the accuracy and stability of the designed hetero-oligomers.
7. The platform demonstrates scalability and versatility, allowing for the systematic exploration of diverse structural and functional protein assemblies.
@KostelicMarius @BasileWicky @kribler @UWproteindesign
💻Code: github.com/rdkibler/pseud…
📜Paper: nature.com/articles/s4146…
#ProteinDesign #StructuralBiology #Bioengineering #SyntheticBiology