Mihailo Jovanovic
46 posts


Mihailo Jovanovic
@mihailoxyz
building biology answer engine @AppliedSciAI; ai/ml/data engineer with bits of bioinformatics

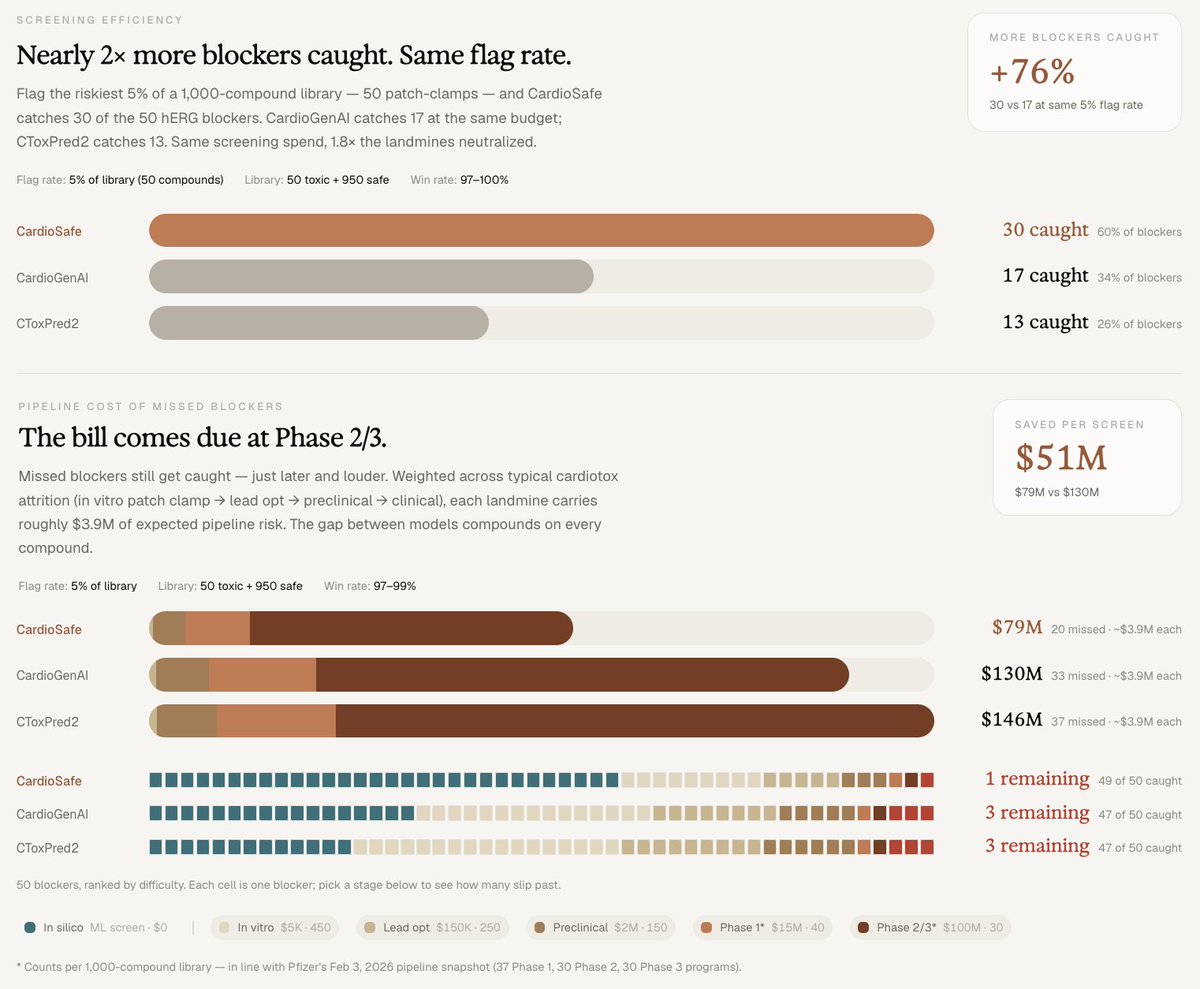

When most people hear a drug was withdrawn from the market, they picture something dramatic: a scandal, a coverup, a clinical disaster. The reality is usually quieter and stranger. A medicine that helped millions gets pulled because of an arrhythmia that hits a few patients out of thousands. The drug worked. It was safe for nearly everybody. But for an unlucky minority, it nudged the heart's electrical timing just enough to cause a fatal rhythm. A single ion channel - hERG - is now the leading cause of safety-related drug attrition in pharma. ~60% of new molecular entities show hERG-blocking liability in early screening. The cardiac safety filter is one of the dominant cost-of-capital decisions in early drug development. And yet the predictive tools for it have been surprisingly limited. Introducing CardioSafe CardioSafe is a multi-task neural network that predicts blocking activity across hERG, Nav1.5, Cav1.2, and IKs simultaneously - from a single chemical structure, in microseconds. CardioSafe aims to catch cardiac safety failures earlier, when they cost thousands of dollars instead of hundreds of millions, and efficiently rescue safe compounds buried in pharma archives for a fraction of the cost. The Results vs. the best published baselines on leak-free benchmarks: • AUC 0.919 vs 0.849 (CardioGenAI) and 0.819 (CToxPred2) • $51M avoided pipeline liability per 1,000-compound screen • 76% more blockers caught at equal patch-clamp budget • 39% lower cost per confirmed-safe lead in drug rescue For a mid-sized biotech running 3-5 focused libraries annually, the cumulative effect is meaningful: roughly $150 to $250M in avoided pipeline risk per year. More details in our new preprint below. Why it Works Two reasons: more data, and sharper resolution between near-identical compounds. First, CardioSafe was trained on substantially more data. Cardiac ion channel datasets are scattered across public databases in formats - censored values, inhibition-percentage votes - that prior models discarded. We kept them, with a curation policy that respects what each measurement actually means experimentally. That single choice contributes more to performance than any architectural decision we made. Second, CardioSafe was also trained to resolve activity cliffs - pairs of compounds that look nearly identical but have opposite cardiac profiles. Terfenadine (withdrawn for arrhythmias) and fexofenadine (safe, multi-billion-dollar antihistamine): same scaffold, opposite hearts. We curated 30 such pairs from the cardiac literature and fine-tuned the model to explicitly rank the blocker above its safer twin. With both molecules held out of training, CardioSafe resolves the cliff correctly. Other models flag the whole class as dangerous. The Bigger Picture CardioSafe is a proof point for how biology-native AI can run: • Multi-task prediction across structurally related targets • A closed loop with multimodal experimental assays - model proposes, MEA measures, model updates • Ruthless curation of heterogeneous public data On that last point: a @demishassabis quote recently re-surfaced on the heels of the Isomorphic raise saying the bottleneck in AI x bio isn't data - it's algorithm sophistication: "You do have enough data - if you were innovative enough on the algorithm side." Our preprint suggests a third answer. We tested multiple architectures. Cross-attention fusion, ChemBERTa embeddings, predicted transcriptomics across 978 landmark genes. They moved the headline number, but not significantly. What did: Keeping the measurements everyone else threw away and understanding what they actually mean pharmacologically. That single curation decision contributed more to performance than the architectural choices. The bottleneck isn't just more data. It isn't just better algorithms. It's also domain understanding applied to the data that already exists. These principles can extend to DILI, nephrotoxicity, neurotoxicity, and beyond. CardioSafe is the first module. The same architecture that learns to predict a drug's effect on the heart might be able to do the same in the liver, the kidney, the brain, even in plants. The platform is what we're building at ASI. Preprint & early access links below ↓

When most people hear a drug was withdrawn from the market, they picture something dramatic: a scandal, a coverup, a clinical disaster. The reality is usually quieter and stranger. A medicine that helped millions gets pulled because of an arrhythmia that hits a few patients out of thousands. The drug worked. It was safe for nearly everybody. But for an unlucky minority, it nudged the heart's electrical timing just enough to cause a fatal rhythm. A single ion channel - hERG - is now the leading cause of safety-related drug attrition in pharma. ~60% of new molecular entities show hERG-blocking liability in early screening. The cardiac safety filter is one of the dominant cost-of-capital decisions in early drug development. And yet the predictive tools for it have been surprisingly limited. Introducing CardioSafe CardioSafe is a multi-task neural network that predicts blocking activity across hERG, Nav1.5, Cav1.2, and IKs simultaneously - from a single chemical structure, in microseconds. CardioSafe aims to catch cardiac safety failures earlier, when they cost thousands of dollars instead of hundreds of millions, and efficiently rescue safe compounds buried in pharma archives for a fraction of the cost. The Results vs. the best published baselines on leak-free benchmarks: • AUC 0.919 vs 0.849 (CardioGenAI) and 0.819 (CToxPred2) • $51M avoided pipeline liability per 1,000-compound screen • 76% more blockers caught at equal patch-clamp budget • 39% lower cost per confirmed-safe lead in drug rescue For a mid-sized biotech running 3-5 focused libraries annually, the cumulative effect is meaningful: roughly $150 to $250M in avoided pipeline risk per year. More details in our new preprint below. Why it Works Two reasons: more data, and sharper resolution between near-identical compounds. First, CardioSafe was trained on substantially more data. Cardiac ion channel datasets are scattered across public databases in formats - censored values, inhibition-percentage votes - that prior models discarded. We kept them, with a curation policy that respects what each measurement actually means experimentally. That single choice contributes more to performance than any architectural decision we made. Second, CardioSafe was also trained to resolve activity cliffs - pairs of compounds that look nearly identical but have opposite cardiac profiles. Terfenadine (withdrawn for arrhythmias) and fexofenadine (safe, multi-billion-dollar antihistamine): same scaffold, opposite hearts. We curated 30 such pairs from the cardiac literature and fine-tuned the model to explicitly rank the blocker above its safer twin. With both molecules held out of training, CardioSafe resolves the cliff correctly. Other models flag the whole class as dangerous. The Bigger Picture CardioSafe is a proof point for how biology-native AI can run: • Multi-task prediction across structurally related targets • A closed loop with multimodal experimental assays - model proposes, MEA measures, model updates • Ruthless curation of heterogeneous public data On that last point: a @demishassabis quote recently re-surfaced on the heels of the Isomorphic raise saying the bottleneck in AI x bio isn't data - it's algorithm sophistication: "You do have enough data - if you were innovative enough on the algorithm side." Our preprint suggests a third answer. We tested multiple architectures. Cross-attention fusion, ChemBERTa embeddings, predicted transcriptomics across 978 landmark genes. They moved the headline number, but not significantly. What did: Keeping the measurements everyone else threw away and understanding what they actually mean pharmacologically. That single curation decision contributed more to performance than the architectural choices. The bottleneck isn't just more data. It isn't just better algorithms. It's also domain understanding applied to the data that already exists. These principles can extend to DILI, nephrotoxicity, neurotoxicity, and beyond. CardioSafe is the first module. The same architecture that learns to predict a drug's effect on the heart might be able to do the same in the liver, the kidney, the brain, even in plants. The platform is what we're building at ASI. Preprint & early access links below ↓










something of significant occurrence has arrived





