
MLiNS Lab
41 posts

@mlins_lab
The MLiNS Lab studies how AI/ML can be used to improve the diagnosis and treatment of patients with neurosurgical diseases.





















⚡️📣Delighted to announce MMP, a prototype-based multimodal framework combining histology and transcriptomics for cancer outcome prediction, to appear in #ICML 2024 @icmlconf. Congratulations to our superstar postdoc @GreatAndrew90 and rest of the team who helped the study. Paper: arxiv.org/pdf/2407.00224 Code: github.com/mahmoodlab/MMP This represents the latest iteration of the multimodal fusion frameworks our lab has investigated since Pathomic Fusion by @richardjchen in 2019. Few interesting facts to know about MMP - Multimodal extension of PANTHER (CVPR 2024), combining morphological prototypes and transcriptomic prototypes (pathways) - Outperforms other multimodal baselines with ~10x less computation - Intuitive prototype-oriented cross-modal interpretability analyses #ComputationalPathology #DigitalPathology #ICML2024 #MultimodalFusion
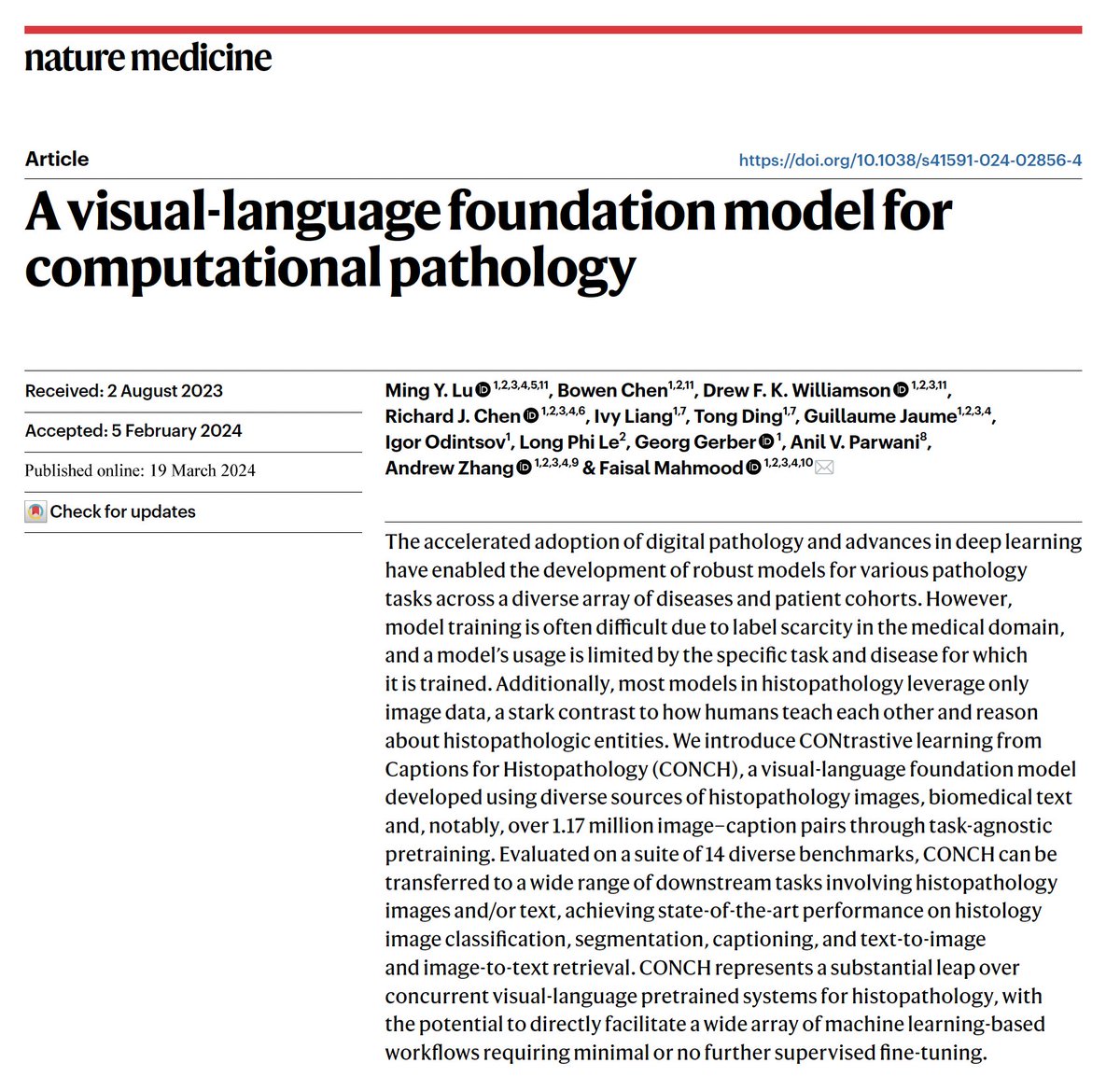











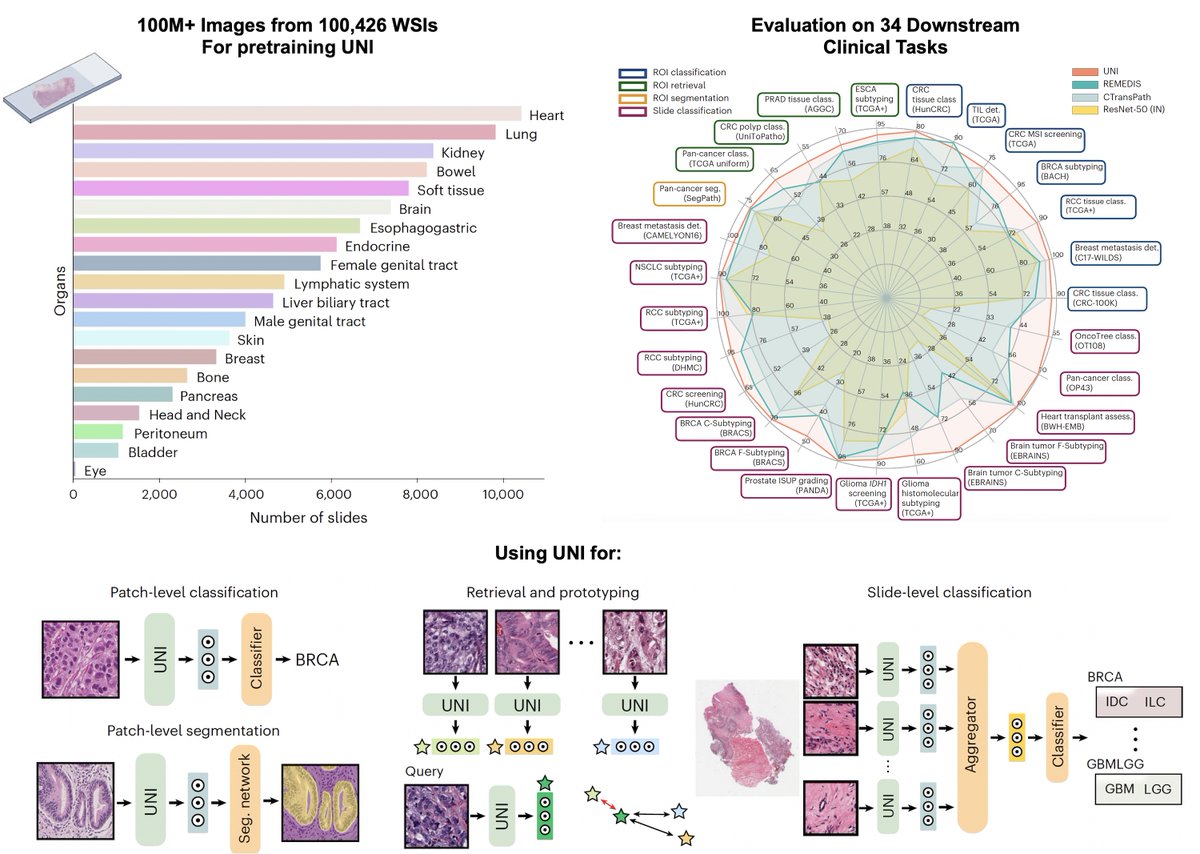
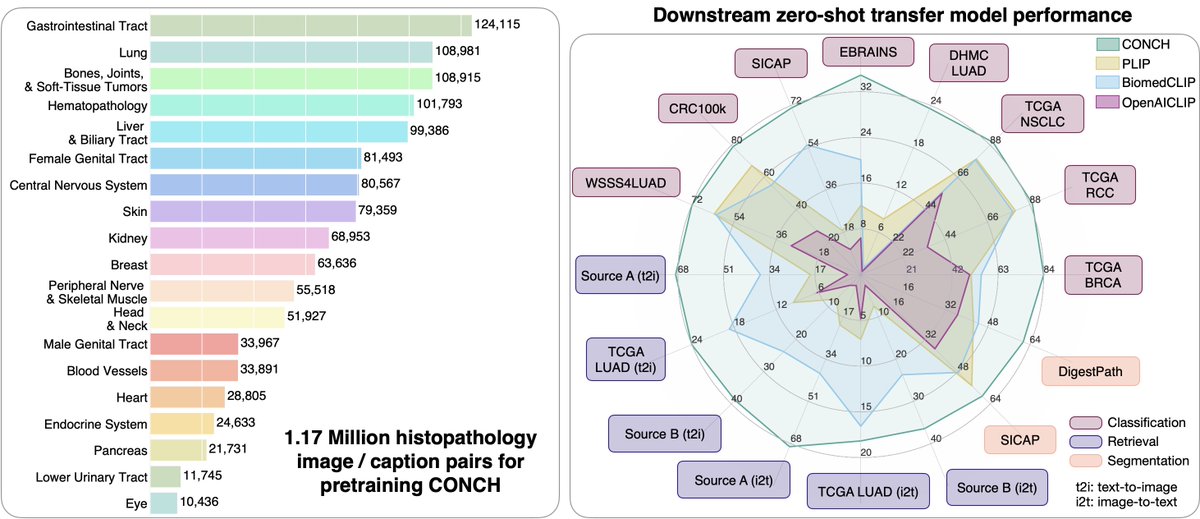

Pretrained using over 100,000 diagnostic histopathological slides across 20 major tissue types, a self-supervised model is shown to outperform existing baselines across various clinically relevant tasks @AI4Pathology @harvardmed nature.com/articles/s4159…




