Nick Youngblut
40 posts

Nick Youngblut
@nick_youngblut
Working on computational biology at the Arc Institute
Katılım Aralık 2013
100 Takip Edilen235 Takipçiler


@bycloudai Normalize by API price and the chart will look very different
English
Nick Youngblut retweetledi

Revealed: the millions of dollars in time wasted making papers fit journal guidelines nature.com/articles/d4158…
English

@mruehlemann @pam_ferretti @PLOSCompBiol Thanks! The proportion of misassemblies is highest for the longest and the shortest contigs. See Fig D in the supplemental
English

Excited to share our latest article published in @ploscompbiol: journals.plos.org/ploscompbiol/a… #ploscompbiolauthor
ResMiCo accurately identifies misassemblies, which seem to be quite prevalent in MAG datasets (~7% of contigs misassembled).
English

If you assemble metagenomes or are interested in deep learning methods applied to metagenome data, check this paper out
Metagenomics Papers@metagenomic_lit
ResMiCo: Increasing the quality of metagenome-assembled genomes with deep learning ift.tt/wdgJXKp
English

ResMiCo is easy to install and use.
Check out the code at github.com/leylabmpi/ResM…
English
Nick Youngblut retweetledi

Now @emily_fogarty11's pre-print is officially out (🎉), I'd like to tell you more about one of the most curious things we (or at least me, my ignorant self) stumbled upon in the human gut microbiome.
This is about a cryptic plasmid, called pBI143.
biorxiv.org/content/10.110…
English
Nick Youngblut retweetledi

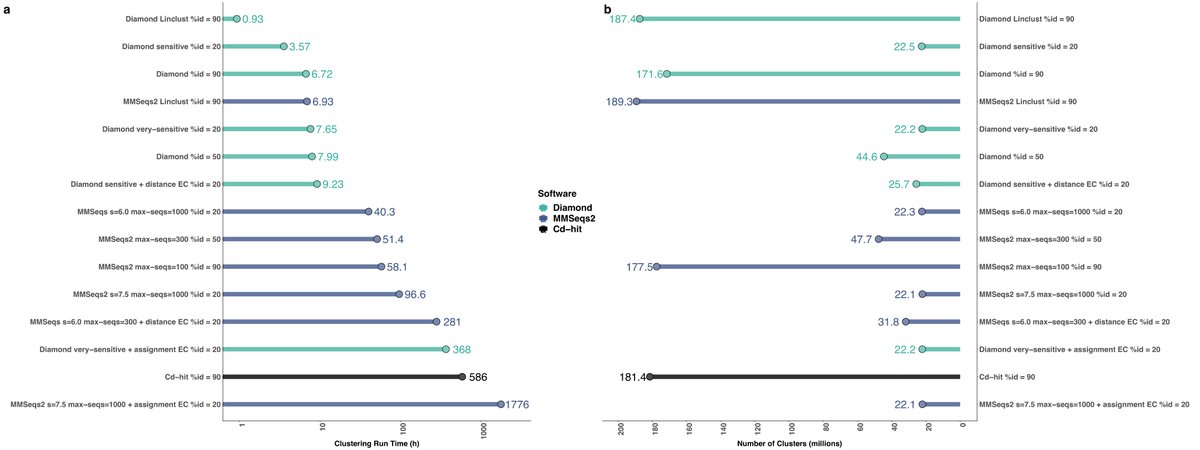
An extraordinary effort with @bbuchfink, @Haim_Ashkenazy, Klaus Reuter & John Kennedy 👉 with fantastic support by @PlantEvolution, @MPI_for_DB & @maxplanckpress.
With DIAMOND DeepClust protein clustering is up to 13x faster than MMSeqs Linclust or 82x faster than CD-hit.
English

Need to interpret ML models for microbiome or other (biological) data?
Interested in methanogens (archaea) in the human gut?
For either: check out the endoR manuscript, now published at PLOS Comp Bio
dx.plos.org/10.1371/journa…
English
Nick Youngblut retweetledi
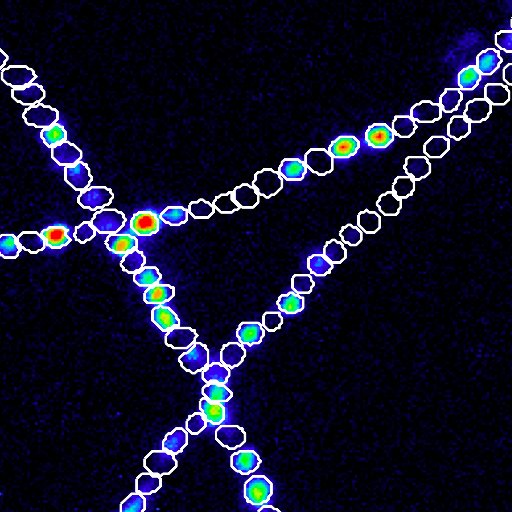
🦠+🧠=⚠️ New #preprint that we are super excited about: #archaea in #human gut #microbiome linked to #Parkinson’s disease pathogenesis. @uni_lu @lcsb_uni_lu @ResearchLux Funded by @ERC_Research @FnrLux @MichaelJFoxOrg @ParkinsonDotOrg @Rotary #Luxembourg
researchsquare.com/article/rs-182…
English

We developed a new deep learning model for predicting misassemblies in metagenome assemblies, with much better performance than existing methods
ResMiCo: increasing the quality of metagenome-assembled genomes with deep learning biorxiv.org/content/10.110…
English
Nick Youngblut retweetledi

Our paper published today @nature describes an ancient epidemic that occurred in central Eurasia in the years 1338-1339. We find that the epidemic was caused by plague (Y. pestis) and that it was associated with the beginnings of the Black Death. nature.com/articles/s4158…
English
Nick Youngblut retweetledi
The human microbiome: there is much left to do @Nature
Perspective piece by @RuthLeyMicro
on the 10th anniversary of the Human Microbiome Project on where we go next.
nature.com/articles/d4158…
English
Nick Youngblut retweetledi
Nick Youngblut retweetledi

Our story of horizontal gene transfer in nematodes has made the cover story in Molecular Evolution and Biology. Thanks to Daphne Welter for her fantastic illustration. Link to the article: academic.oup.com/mbe/article/39…

English
Nick Youngblut retweetledi

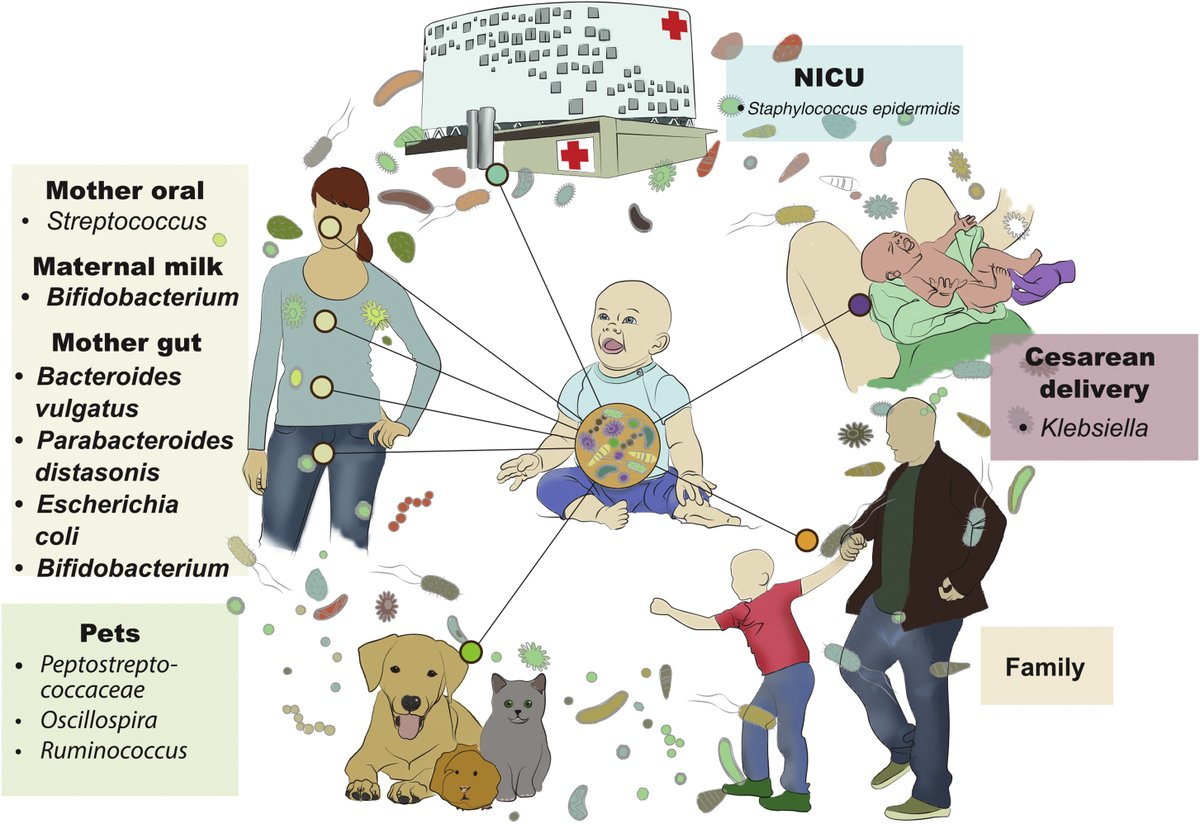
The developing infant gut #microbiome: A strain-level view. @HagayEnav @BackhedLab @RuthLeyMicro review recent studies on #infant #gutmicrobiome, moving from broad taxonomic studies to strain level analyses & shedding new light on colonization & maturation cell.com/cell-host-micr…
English