Sabitlenmiş Tweet
Caleb Ellington
596 posts


Caleb Ellington
@probablybots
Scientist @genbioai | PhD @CMUCompBio | Creator/maintainer https://t.co/E3h3NJS6S7 | multi-task learning, graphical models, and personalized medicine
San Francisco, CA Katılım Aralık 2017
364 Takip Edilen805 Takipçiler
Caleb Ellington retweetledi

Last month, we shared research on why LLMs fail at data analysis: even the best models hallucinate answers when reasoning over structured data.
Today we're launching what we've built to fix it.
Summand is now live at summand.com.
What most teams want is simple: plug AI into their data and get answers they trust. Most "chat with your data" tools try to deliver that by translating your question into SQL and hoping for the best. Summand does something harder: it builds up a real understanding of your data. What your columns actually mean, how your tables relate, where the edge cases live. You can contribute to that understanding too, and so can the agent.
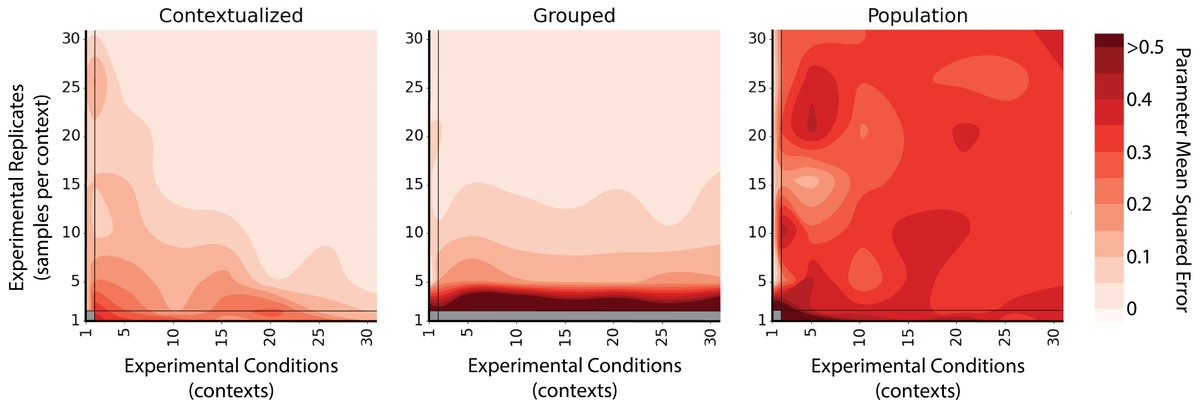
Under the hood, that understanding is grounded in interpretable ML and a semantic layer purpose-built for structured data. That's what makes the answers trustworthy.
Why the name “Summand”? Just like how a summand is a term in a summation, Summand decomposes your data into interpretable reasoning components. By breaking complicated outcomes into simple patterns, Summand makes downstream AI systems reliable and transparent.
*What this means to you:* Connect your data to Summand.com, start asking questions immediately, and power your downstream AI applications through Summand’s MCP access.
Try it today → summand.com
English

Huge thanks to the excellent co-authors behind this work: Sohan Addagudi, @JiaqiWang_, @ben_lengerich, and @ericxing.
This is the final chapter of my phd, but you'll continue seeing this kind of work reflected at @genbioai in our work on general-purpose biological simulators.
English

We curated DDR-Bench and DTR-Bench to validate virtual cells on practical drug discovery tasks and enable hill-climbing on useful hills. We make one contribution to cell-level screening with CellVS-Net, but the ceiling is still quite far away!
Pre-print: biorxiv.org/content/10.648…
English
Caleb Ellington retweetledi

Computer systems have long been developed to consider adaptive memory problems (e.g. cache invalidation, hierarchical storage, speculative reuse, routing, and scheduling). Some people say "Hashing and caching is 50% of computer science."
As generative models scale, many of these same problems are starting to reappear inside the generative models.
Two new papers from our group, both led by Dong Liu, explore this idea through adaptive caching and memory management:
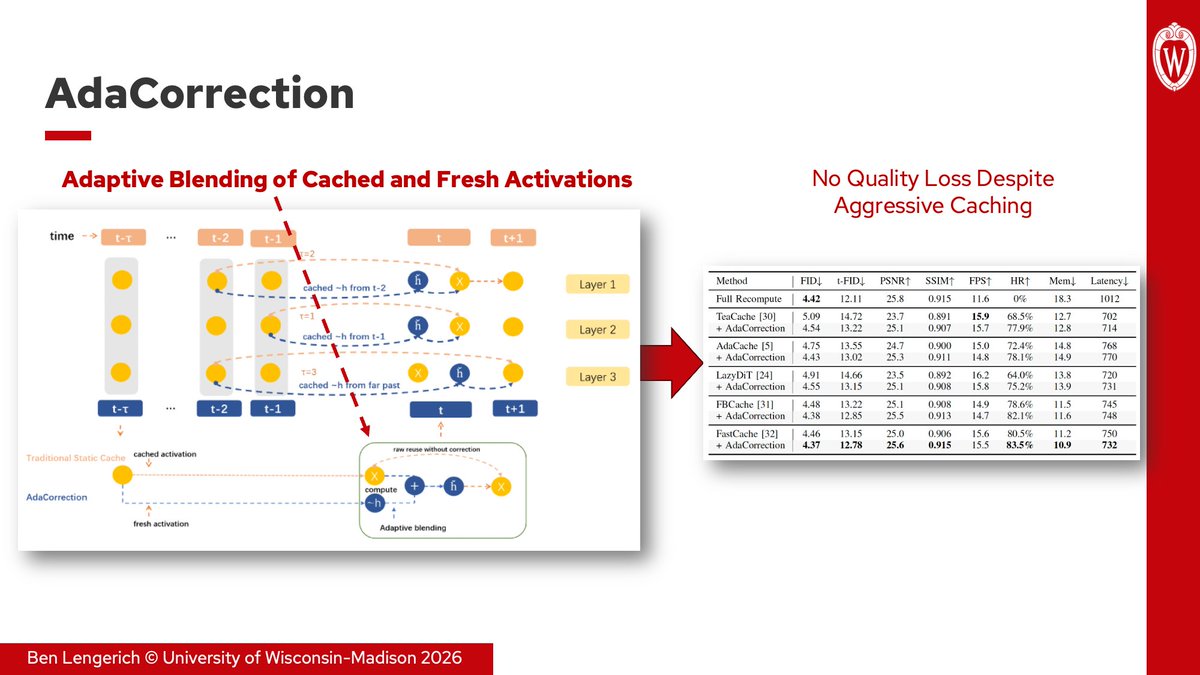
1. AdaCorrection (ICIP 2026, arxiv.org/abs/2602.13357) measures activation drift during diffusion inference and adaptively blends cached and fresh features instead of relying on static reuse schedules. Combined with FastCache, AdaCorrection reduces memory by 40% and latency by 30% without any loss to FID.
2. Memory-Keyed Attention (Computing Frontiers 2026, arxiv.org/abs/2603.20586) organizes KV memory into local, session, and long-term tiers, then learns per-token routing across them. It achieves up to 5× faster training and 1.8× lower decode latency vs. MLA with comparable perplexity.
These build on our earlier PiKV work (ICML ES-FoMO 2025, arxiv.org/abs/2508.06526), which explored adaptive KV cache management for MoE serving.
Common thread across all three: reuse policies cannot remain static as models scale. Efficient generative systems increasingly require adaptive decisions about what to reuse, when to refresh, and what memory should be attended to for a given token or timestep.

English
Caleb Ellington retweetledi

Excited to share Orthrus is now published in Nature Methods! This was a work from our PhDs in which we showed 3 things:
- There's lots of room for new biologically grounded self-supervised objectives
- The "y - intercept" in scaling is important! We show that representations from 10 million parameter Orthrus outperform a 7 billion parameter model, 700 its size.
- Orthrus works in the low-data regime where data acquisition is especially expensive: low throughput experimental data and clinical trials
Ian and I are now building BlankBio to apply these ideas at a bigger scale. I'm going to be at AACR get in touch if you want to chat!

English
Caleb Ellington retweetledi

I wrote a little reflection after my PhD since it would be a shame not to commemorate 5 years of life
I. A PhD committee meeting is sort of like a roast session with the best intentions
II. I was able to witness two paradigm shifts (1) FMs (2) LLMs
III. Mismatch between the training objectives and the biology
IV. Our lunchtime conversations about devaluation of intellectual labor took on a more anxious tone
V. During periods of upheaval good scientists need to do two things
English
Caleb Ellington retweetledi

🎉 New on Arxiv:
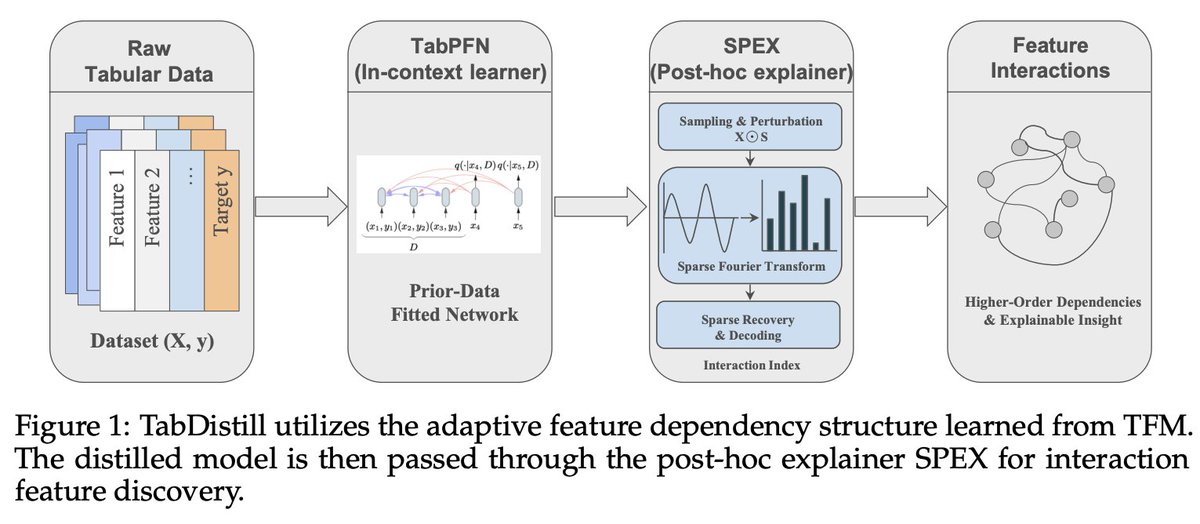
A fundamental challenge of statistical learning is discovering which features interact with one another to influence outcomes. Looking for pairwise intx. requires O(n^2) calculations, three-way intx. requires O(n^3), etc.
Can we skip this statistical challenge by tapping into the rich prior learned in foundation models?
Our approach TabDistill shows that we can.
Read more:
arxiv.org/pdf/2604.13332
English
Caleb Ellington retweetledi

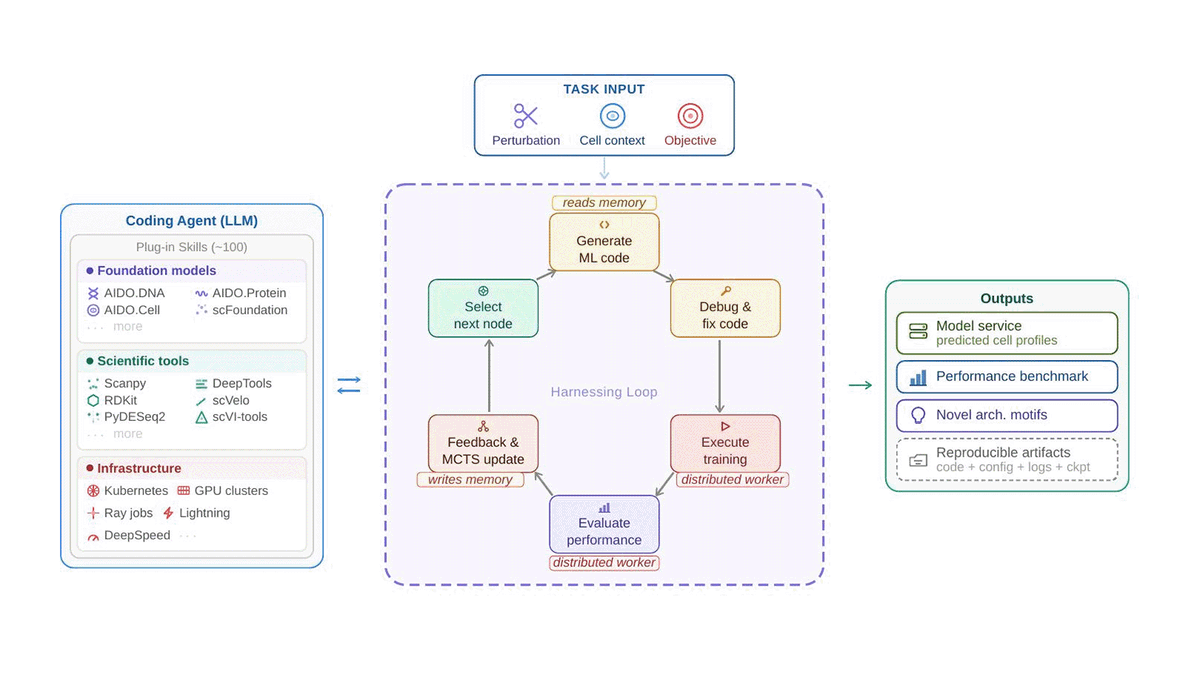
What if AI could build its own biology models — and beat human experts?
We built VCHarness: an autonomous system that proposes models, writes code, runs experiments, and learns from results in a closed loop.
Tested on CRISPR gene knockdown across 4 human cell lines — it outperforms expert-designed baselines.
From months of manual model-building → days of autonomous search.
🔬 Blog: genbio.ai/harnessing-ai-…
📄 Paper: biorxiv.org/content/10.648…
🌐 Website: genbio-ai.github.io/VCHarness
English

@maxhodak_ I've done a number of studies on this, but my favorite is this one,
pnas.org/doi/10.1073/pn…
We compare representation-based subtypes to reference molecular subtypes for 25 cancer types. The representation dramatically improves improves stratification of cohort survival.
English

@maxhodak_ One promising approach is learning a disease representation that stratifies the actual mechanism of interest, and then define cohorts in this space. Cohorts are still necessary, they're just hard to identify with individual biomarkers. We need a way to "compile" mechanisms...
English

> But this creates a combinatorial explosion! If you have 20 binary biomarkers, that’s over a million possible patient subgroups. No trial, no matter how well-funded, can enumerate that space.
I continue to believe that RCTs are often the wrong tool for modern oncology
owl@owl_posting
this is an essay about cancer, how it is one of the most 'detailed' diseases in existence, and why we must delegate the understanding of that complexity to machine intelligence owlposting.com/p/cancer-has-a… 3.4k words, 15 minute reading time
English