Patrick Lim retweetledi
Patrick Lim
342 posts


Patrick Lim
@psl_lim
Postdoc @WhiteheadInst @MIT
Cambridge, MA Katılım Temmuz 2018
815 Takip Edilen174 Takipçiler
Patrick Lim retweetledi

New Essay: CULTURE SHIFT
The human immune system is, in one sense, a detection mechanism. It has evolved, over millions of years, to scan the body for molecular signals that tell it whether to attack or stand down. Most of these signals come from pathogens, damaged cells, or the body’s own hormones. But in 2019, a lab in Germany published a finding that pointed to a much stranger source: one of the signals sensed by the immune system is found in sauerkraut.
When people eat sauerkraut, a molecule called phenyllactic acid (D-PLA) — found in fermented foods — enters their bloodstream and activates a receptor, known as HCA3, on immune cells, triggering an anti-inflammatory response. In addition to lactic acid, phenyllactic acid is one of many compounds produced by lactic acid bacteria during the fermentation of sauerkraut and related fermented foods. Prior to this study, other molecules had been found to bind HCA3, but D-PLA was a hundredfold more potent than any of them.
This discovery advances our understanding of how fermented foods can reduce inflammation, but more striking is what it suggests about hominid physiology. Although HCA3 is part of a larger family of receptors broadly conserved across eukaryotes, HCA3 is only present in humans and other great apes like chimpanzees and gorillas — and not even in other mammals. It is a recent addition to the genome, appearing only a few million years ago. Its existence seems to suggest that our immune system evolved to recognize the microbial metabolites from fermented foods.
We tend to think of fermented foods as something humans invented and then chose to eat. But, increasingly, scientific evidence suggests the causality runs the other way: Fermented foods appear to have helped shape human biology itself, and our bodies may have been built, in part, to expect them.
The case for this runs from changes in hominid gut anatomy millions of years ago to the HCA3 receptor, to a growing body of research linking fermented food consumption to immune function and gut health. And it raises an uncomfortable question about what happened when the Western food system, in the name of safety and efficiency, quietly removed these foods from our diets in the nineteenth and twentieth centuries.
Read the full essay: press.asimov.com/articles/cultu…

English
Patrick Lim retweetledi

Have a photo from your research you'd like to see in print?
We’re collecting images for Making the Modern Lab, an upcoming book from Asimov Press about the tools, spaces, and experiments that define modern biology.
If your image is selected, we’ll pay $30 per photo and include a photo credit in the book.
Microscopy images, lab benches, instruments, experimental setups, and lab environments are all welcome. Phone photos are fine —we’re most interested in capturing the real look of modern research — but we are seeking well captured images with good composition. People are allowed in the photos, if the image clearly depicts the topic.
While this call is open, we have a list of images we are specifically searching for:
CHO cell microscope image
Liquid Handling Platforms (full view of instruments, action shots of liquid transfers)
A multiwell plate being fed into a plate reader.
Zebrafish (adult fish in tanks, microscope embryo images)
Biochemical buffers (labeled bottles lined up)
Bioreactors (any size)
C. elegans (full body or fluorescent microscopy. Ideal if neurobio related images)
Immunohistochemistry microscopy images
Electroporator setup
Liquid Chromatography setups (HPLC, HPLC-MS, or FPLC)
Mammalian tissue culture media
S. cerevisiae microscope image (ideal if you can get an image depicting cells in different parts of the life cycle.)
BONUS: Images of historical lab equipment in your lab.
Submit here: forms.gle/JBYb5YmfduaDaW…


English
Patrick Lim retweetledi
Patrick Lim retweetledi
Patrick Lim retweetledi

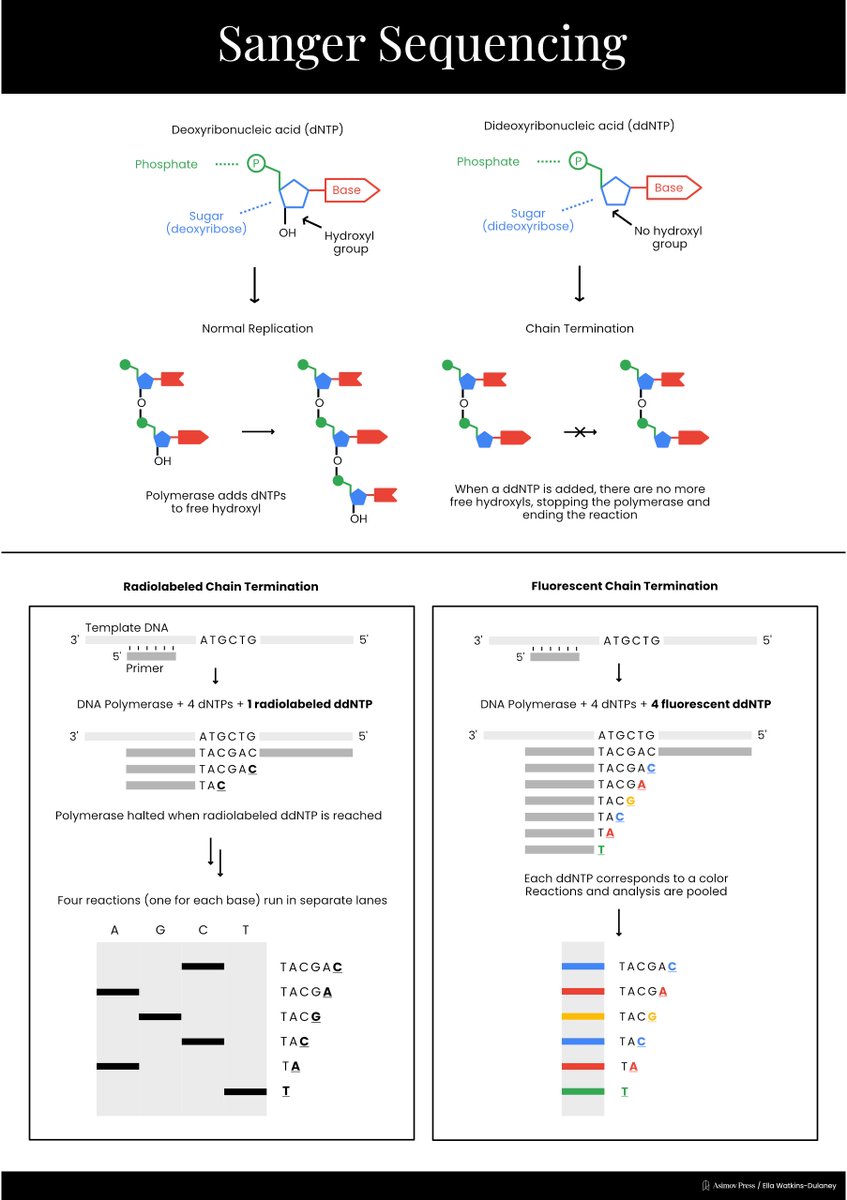
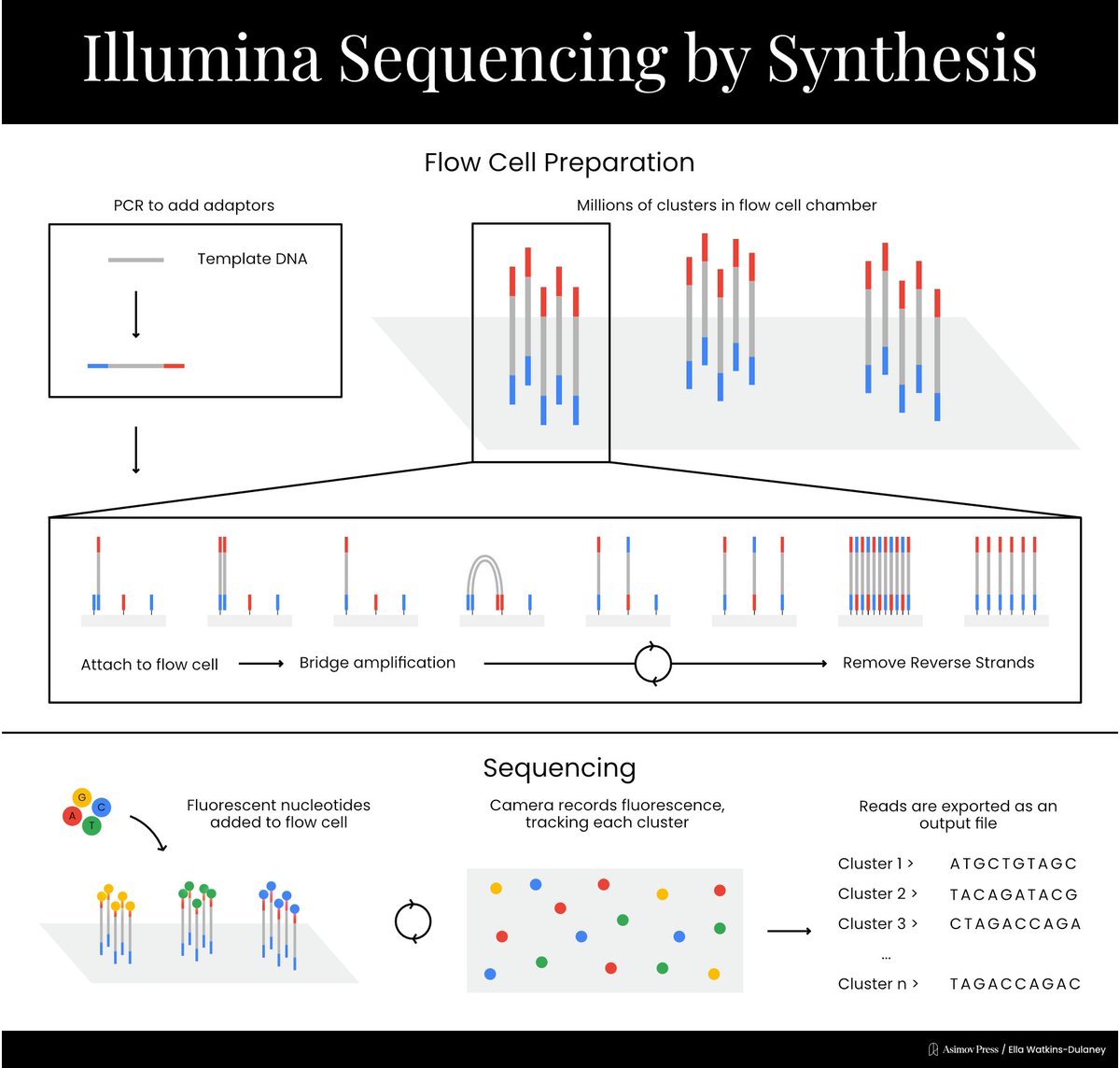
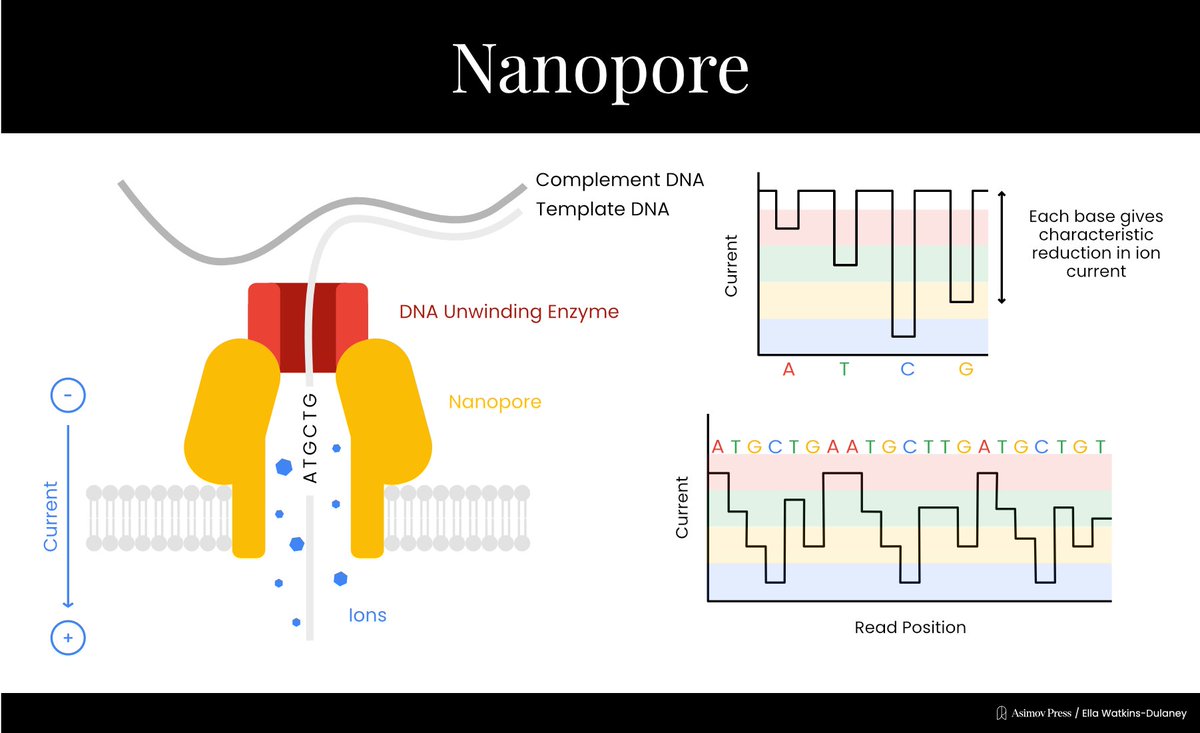
A Visual Guide to DNA Sequencing.
Learn how different DNA sequencing technologies work, from Sanger sequencing to Illumina to nanopores. (Complete with illustrations!)
Written by @evandeturk. Illustrated by @EllaWD_PhD.
GIF
English

Happy to announce that part of my PhD work on SET in the early embryo has finally been published :) !
pubmed.ncbi.nlm.nih.gov/41665009/
Eran Meshorer@meshorer1
Finally out! Patrick developed a system to study SET isoform specificity in ES cells, revealing a role for SETa/b switch in endoderm specification. Check it out🔥 @psl_lim academic.oup.com/nar/article/54…
English
Patrick Lim retweetledi

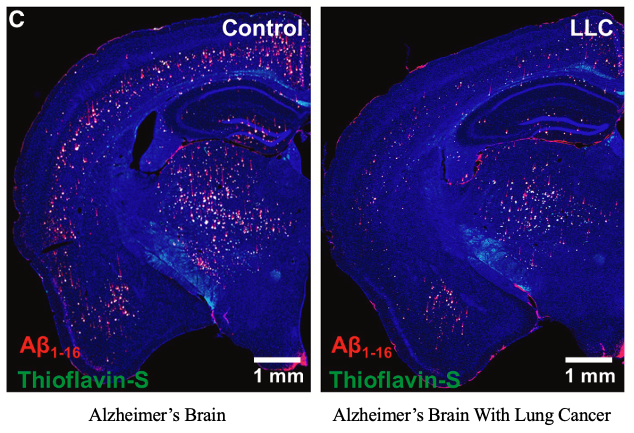
Cancer patients rarely get Alzheimer's.
And a 15-year study in @Cell just explained why. They found a protein that clears brain plaques in mice - by activating the brain's own IMMUNE CELLS 🧵
English
Patrick Lim retweetledi

Thanks to the authors for sharing all components of this mouse embryo spatial transcriptomics data from cell gene counts to per-molecule coordinates: ahajournals.org/doi/full/10.11… 🥳
So I vibe coded an app to explore the 3D subcellular transcript organization: jef.works/CellCarto-Mous…
English
Patrick Lim retweetledi

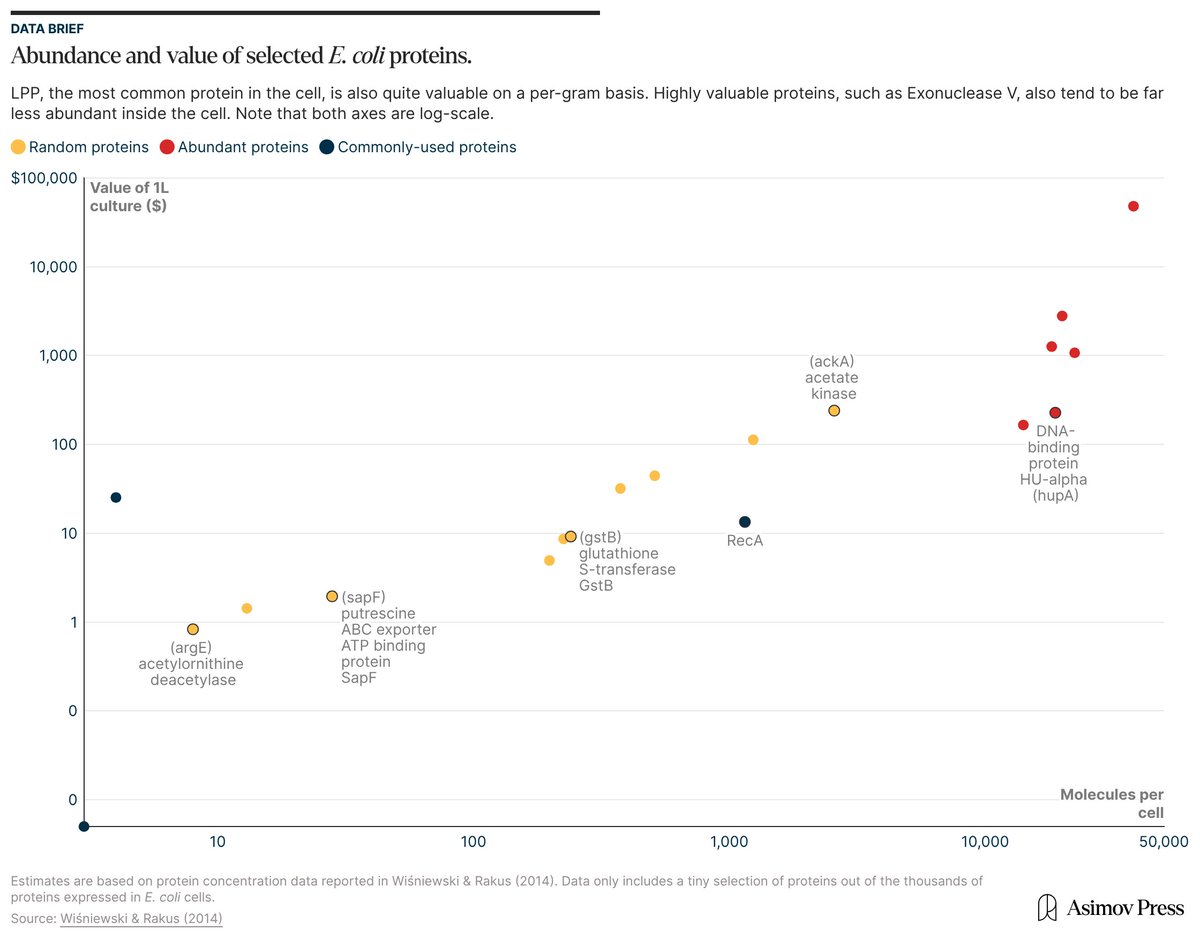
If you took a liter of E. coli cells, isolated all their molecules, and sold them, they'd be worth ~$600,000.
This is a thought experiment, of course; just doing the isolations might cost more than the molecules are worth.
But if you're keen to learn which parts of cells are worth the most money, check out our article: "The Price of E. coli."
press.asimov.com/articles/price…

English

New Veritasium video has the clearest explanation of EUV tech I've ever seen.. highly reccomended
youtube.com/watch?v=MiUHjL…

YouTube
English
Patrick Lim retweetledi

When polishing scientific figures for publication, I try to limit fonts + font sizes to reduce visual noise. But doing this by hand is tedious.
So I vibe coded a web app to standardize text in SVGs: jef.works/SVG-Text-Stand…
Try it out! Spot the differences. Hope it's helpful

English
Patrick Lim retweetledi

An in vivo imaging technique can track the way molecules move through the nuclear pore complex
go.nature.com/4pCGNnz
English
Patrick Lim retweetledi

Now that you have gathered you raw data from a cell migration assay, how do you analyze that data?
In our review, we describe methods - including AI methods - to track migrating cells and extract as much information as possible.
Read our paper here: nature.com/articles/s4159…

English
Patrick Lim retweetledi

Learning cell dynamics with neural differential equations
Cell fate is not purely deterministic. When a multipotent progenitor differentiates, it doesn't follow a single predetermined trajectory—it navigates a landscape shaped by both regulatory programs and random fluctuations. This interplay between deterministic and stochastic forces is how biology generates diverse cell types from common ancestors. Yet most computational methods for modeling single-cell dynamics treat noise as a constant background parameter, assuming stochastic behavior is the same regardless of cell state.
Michael Vinyard and coauthors take a different approach. They introduce scDiffEq, a framework that learns neural stochastic differential equations directly from single-cell RNA-seq data, parameterizing both deterministic drift and state-dependent diffusion as separate deep neural networks—allowing the model to learn how noise itself varies across the developmental landscape.
The difference matters. When they ablate cell-specific diffusion and replace it with constant noise, fate prediction accuracy drops by 11% and the model produces only single-fate trajectories, completely failing to capture multipotent outcomes. With state-dependent diffusion restored, scDiffEq achieves 58.5% accuracy on lineage-traced haematopoiesis data—a 7.8% improvement over methods like PRESCIENT—and correctly predicts cells that differentiate into multiple lineages.
The framework also enables in silico perturbation screens: simulated Gfi1 overexpression shifts progenitors toward neutrophils while depleting monocytes, matching known biology. And it generalizes beyond multi-time-point experiments to single snapshots—the majority of existing scRNA-seq data—by leveraging pseudotime inference, scaling to over 1 million cells with near-linear complexity.
The message is clear: deterministic models of development are incomplete. Cells actively modulate their susceptibility to noise during differentiation, and capturing this interplay—not just the average trajectory, but the full stochastic landscape—is essential for understanding how progenitors explore and commit to fates.
Paper: nature.com/articles/s4225…

English
Patrick Lim retweetledi

Nature research paper: Human assembloids recapitulate periportal liver tissue in vitro
go.nature.com/3Y3f3fR
English
Patrick Lim retweetledi

How does an embryo reliably "compute" its form - "cell by cell" - using only local interactions and mechanics, yet produce a precise global body plan? I’m excited to share our Nature Methods paper "MultiCell: geometric learning in multicellular development", presenting #AIxBiology research led by @HaiqianYang and the result of a great collaboration with Ming Guo, George Roy, Tomer Stern, Anh Nguyen and Dapeng Bi.
A long-standing challenge in developmental biology is to predict how thousands of cells collectively self-organize as tissues fold, divide, and rearrange. In MultiCell, we represent a developing embryo as a dual graph that unifies two complementary views of tissue mechanics with single-cell resolution: cells as moving points (granular) and cells as a connected foam (junction network). This lets the model learn dynamics from both geometry and cell–cell connectivity.
On whole-embryo 4D light-sheet movies of Drosophila gastrulation (~5,000 cells), our model predicts key cell behaviors and the timing of events, including junction loss, rearrangements, and divisions with high accuracy, at single-cell resolution. Beyond prediction, the same representation supports robust time alignment across embryos and offers interpretable activation maps that highlight the morphogenetic "drivers" of development. The broader goal is a foundation for cell-by-cell forecasting in more complex tissues, and eventually for detecting subtle dynamical signatures of disease.
Kudos to the team for this inspiring collaboration with brilliant researchers to push the boundary of AI for biology!
Citation: Yang, H., Roy, G., Nguyen, A.Q., Buehler, M.J., et al. MultiCell: geometric learning in multicellular development. Nature Methods (2025), DOI: 10.1038/s41592-025-02983-x
Code/data links are in the manuscript.
English
Patrick Lim retweetledi

There's a recent blog from @OpenAI where they used GPT-5 to optimize a common biology experiment, called Gibson Assembly. I've seen criticisms online from people who say things like, "Who cares? A human totally could have done that" or whatever. And that's true. But I still think this blog is nice for a couple reasons.
First, faster iterations / more reliable experiments is one of the best ways to accelerate biotechnology progress more broadly! Experiments take much too long, and are often much too unreliable, for scientists to move quickly. Therefore, we should invest more resources toward optimizing and improving common methods that seem "mundane".
Second, this is a simple experimental system in which to test AI; indeed, that's the whole point! Gibson Assembly has been around for nearly two decades, is widely-used, and only requires three enzymes. It is therefore a natural fit for AI companies to benchmark their models on biological questions. (The parameter space is not too large!)
To understand what OpenAI actually did, I first need to tell you about Gibson Assembly, a common method biologists use to stitch DNA molecules together. Originally developed in 2009, most scientists use Gibson because it's dead simple: Everything works at one temperature (50°C) and it requires only three enzymes. The DNA molecules to be joined together are designed such that they have 15-40 nucleotides, at either end, which overlaps with the other DNA molecule. All the DNA is then added to a tube and an enzyme, exonuclease, "chews back" several dozen nucleotides from the 5' ends of each molecule, leaving behind long single-stranded "arms." These arms float around in the liquid, collide with a matching arm in another DNA sequence, and hug each other tightly. A second enzyme, DNA polymerase, runs along these touching DNA strands and fills in parts of the arms that don't overlap or are still single-stranded. Finally, DNA ligase seals the "nick" and heals the strands, thus forming a newly assembled, double-stranded piece of DNA.
OpenAI collaborated with a new biosecurity startup, Red Queen Bio (co-founded by Hannu Rajaniemi, an excellent science fiction writer), to build the evaluation framework. The metric they settled on is called cloning efficiency, which just means this: For a fixed amount of input DNA (like one picogram) transformed into cells, how many colonies successfully grow and contain the correctly assembled DNA molecule? By the end of their blog post, the OpenAI team claims that they were able to boost this number 79x relative to a "baseline protocol" from New England Biolabs, or NEB, a common purveyor of the Gibson enzymes.
An important note is that OpenAI says no humans were involved in optimizing the reaction; all the humans did was carry out protocols generated by GPT-5, and also upload experimental results back into the model. They repeated this several times, coaxing the model to iterate each time. Their Gibson Assembly was remarkably simple, involving just two DNA molecules: a gene encoding a fluorescent protein and a plasmid to hold the gene.
(The OpenAI team, intriguingly, also set up a set up a robot to automate the Gibson Assembly and transformation, but couldn't get it to work as well as a human. "We compared the robot's work to human-performed experiments at each step. The robot successfully handled the transformation process…When compared directly with human-performed transformations, the robot generated similar quality data with equivalent improvements over baseline, showing early potential for automating and accelerating biological experiment optimization." However. "while the fold-changes between the robot and human experiments were similar, absolute colony counts from the robot were approximately ten-fold lower than manual execution.")
After several rounds of iteration, the model made two notable proposals:
First, it added two additional enzymes to the normal Gibson Assembly reaction. Specifically, it added "the recombinase RecA from E. coli, and phage T4 gene 32 single-stranded DNA–binding protein (gp32)." The blog continues: "Working in tandem, gp32 smooths and detangles the loose DNA ends, and RecA then guides each strand to its correct match." This tweak improved the "cloning efficiency" metric by 14x over the standard NEB protocol.
Second, it made a subtle change to how the assembled DNA molecules were inserted into living cells. Specifically, GPT-5 told the humans to spin down cells in a centrifuge, thus forming a pellet, prior to transforming them. This is typically not recommended because competent cells are "fragile," but the OpenAI team writes that "the cells tolerated concentration well and the increased molecular collisions boosted transformation efficiency substantially (>30-fold on final validation)."
Now, recall that at the start of this little blog I said I really liked this experiment! (Do not crucify me, ye AI optimists.) But no internet commentary is truly complete without some nitpicking, so here goes.
One criticism is that the largest improvement made by the model was not related to Gibson Assembly at all! It was related to how the DNA gets delivered into cells. And, indeed, prior studies have shown something similar. (This research paper, for example, says that one of the best ways to improve transformation is to concentrate cells beforehand. Fair play to the OpenAI team for linking to this in their blog post.) And if you are a human reading this blog, and you are planning to spin down your competent cells before transformation, just be sure to aliquot everything into small tubes first; repeated spins will, over time, kill everything.
Another issue is that adding RecA and gp32 to a Gibson Assembly reaction complicates things quite a bit. For a normal Gibson reaction, everything comes in a single kit from NEB with the enzymes, and the whole experiment is done at one temperature: 50°C! But doing a Gibson Assembly this way would require one to buy purified RecA and gp32, and also change incubation temperatures to get everything working (RecA and gp32 work best at 37°C.) This is more expensive and more complicated, but maybe worthwhile in some cases.
And lastly, the selected metric — namely, how many colonies one gets from a given amount of DNA — doesn't actually seem all that useful in most scenarios. A scientist stitching together two strands of DNA doesn't actually care if they only get five colonies because, often, they only need to get ONE colony that works, and then they can grow up those cells in large beakers and extract a huge amount of the plasmid. A more useful metric might be to increase the total number of unique DNA strands that can be joined together in a single Gibson Assembly reaction, without reducing overall quality, instead.
Still, I liked this blog post as a whole. I'm glad people are optimizing the "small" things, and I don't blame OpenAI for not trying to solve cancer, in its overwhelming magnitude of manifestations, on their first attempt! Gibson Assembly is a much better starting point.

English
Patrick Lim retweetledi

Alzheimer's couldn't possibly be a long term inflammatory response to varicella viruses, which live in nerve cells to evade the immune system and which reactivate in response to stressors of failing health in later life...
(Sarcasm)
Gregory Cochran@gcochran99
Shingrix, the anti-shingles vaccine, appears to have done more good against Alzheimer's disease by _accident_ than all existing Alzheimer's research.
English




