Rahul Dhodapkar
109 posts

Rahul Dhodapkar
@rahuldhodapkar
physician-scientist, resident, computational biology, immunology, neuroscience, ophthalmology | MD @YaleMed | ex-software engineer @MongoDB


An exciting milestone for AI in science: Our C2S-Scale 27B foundation model, built with @Yale and based on Gemma, generated a novel hypothesis about cancer cellular behavior, which scientists experimentally validated in living cells. With more preclinical and clinical tests, this discovery may reveal a promising new pathway for developing therapies to fight cancer.

An exciting milestone for AI in science: Our C2S-Scale 27B foundation model, built with @Yale and based on Gemma, generated a novel hypothesis about cancer cellular behavior, which scientists experimentally validated in living cells. With more preclinical and clinical tests, this discovery may reveal a promising new pathway for developing therapies to fight cancer.




💡 Want to leverage the power of foundation models in graphs? 🔥 Introducing Foundation-Informed Message-Passing (FIMP), a framework for applying any pre-trained transformer-based foundation model to Graph Neural Networks! arxiv.org/abs/2210.09475

Delighted to share our latest work on #longCOVID - sex differences in symptoms and immune signatures. Led by @SilvaJ_C @taka_takehiro @wood_jamie_1 et al. with @LeyingGuan & @PutrinoLab. We find a striking inverse correlation btw testosterone levels and symptom burden👇🏼 (1/) medrxiv.org/content/10.110…




Major Cell2Sentence update 🎉🔬! We’ve been thrilled to see the attention Cell2Sentence has received from the single-cell community. Now, we’re excited to release our first update of Cell2Sentence (C2S) - a framework to leverage LLMs to train foundational single-cell models, directly in text. What’s new & out: Updated preprint with latest results biorxiv.org/content/10.110… First full cell model available on the HuggingFace hub huggingface.co/vandijklab/pyt… Updated codebase for data transformation & training github.com/vandijklab/cel… We now fine-tune language models to generate entire cells, predict combinatorial cell labels, and generate textual data insights directly from cell sentences. We train GPT-2 and Pythia models on a large multi-tissue dataset containing 36M cells from @cellxgene as well as an immune tissue dataset containing 270k cells. C2S LMs achieve SOTA performance in single-cell data generation. C2S models trained for combinatorial label prediction settings excel in low-data regimes, outperforming single-cell foundation model baselines. We also show that C2S models benefit from natural language pre-training and always outperform models trained from scratch on cell sentences. C2S provides a straightforward approach to adapting LLMs for single-cell data analysis, leveraging their natural language capabilities to generate and derive insights from single cells. We are convinced that C2S’ approach of integrating data modalities through text is the way forward for single-cell foundation models, from representing multi-omics data to generating clinical insights, all in a human readable format. We’re excited to start building a community around Cell2Sentence! If you also think that C2S will be the framework for single-cell foundation models, and are interested in contributing, reach out to us! We welcome any collaborations and discussions. Huge thanks to our collaborator @aminkarbasi and the C2S team (@danielflevine, @sachalevy3, @SyedARizvi5688, @nazreenpm, Xingyu Chen, @dzhang03, @GhadermarziSina, Ruiming Wu, Ivan Vrkic, Anna Zhong, Daphne Raskin, Insu Han, @aho_fonseca, @josueortc) for their hard work on C2S! Special thanks to @rahuldhodapkar, who co-supervises this project.

How to infer human labelling of a given dataset in a model-agnostic way? Check our new method HUME accepted at @NeurIPSConf as #spotlight!🌟 HUME provides a new view to tackle unsupervised learning. Kudos to my fantastic PhD student @artygadetsky! Paper arxiv.org/abs/2311.02940

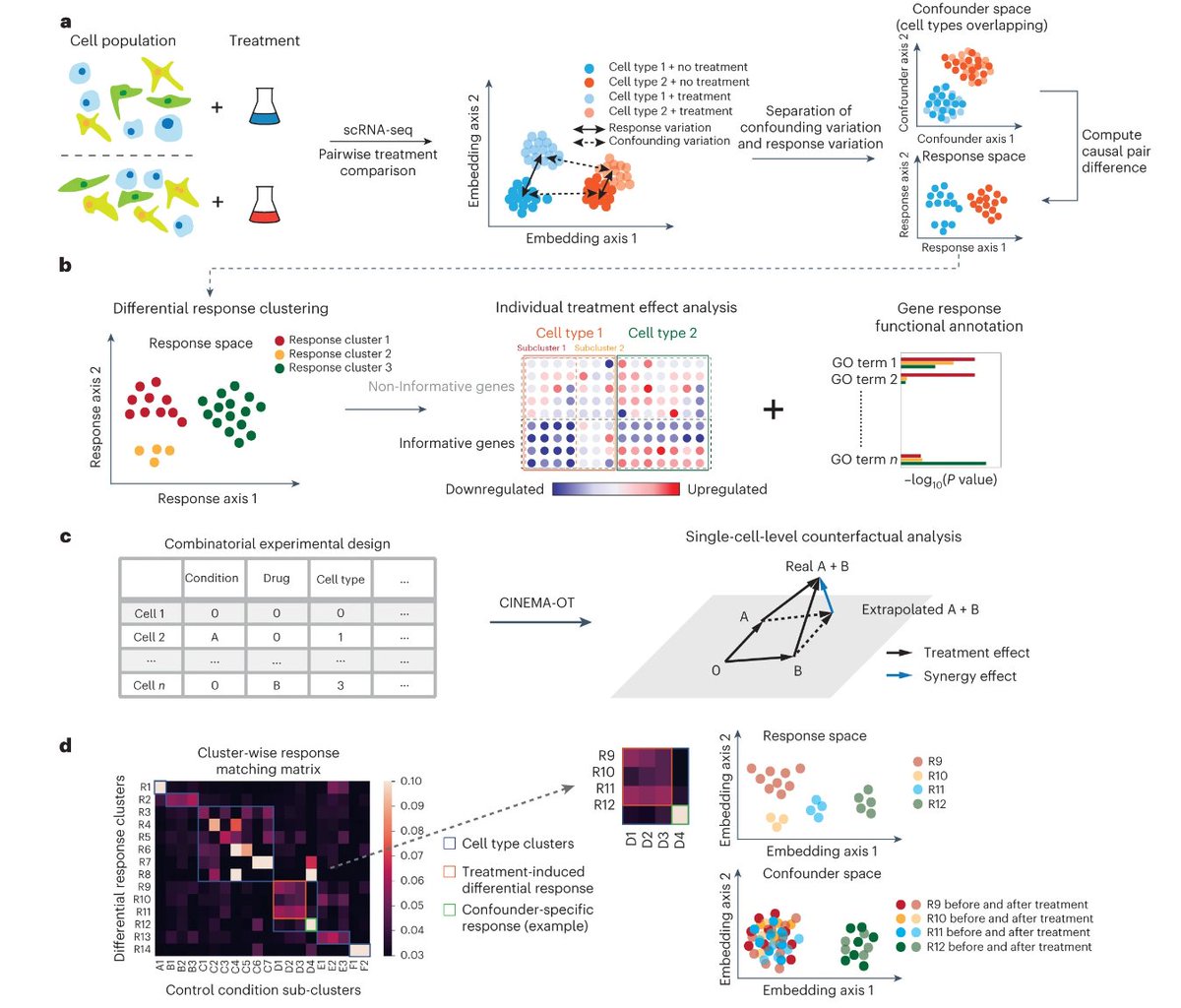

Thrilled to announce that CINEMA-OT is now published at Nature Methods! nature.com/articles/s4159…

So pleased to report that our Mount Sinai-Yale long COVID (MY-LC) paper with @putrinolab & others is now published!! Proud of the hard work of all who contributed. We found biological signatures that can distinguish people with vs. without #longCOVID (1/) nature.com/articles/s4158…

Single Cells as text? We developed Cell2Sentence, a method that allows training of Large Language Models on single-cell data! biorxiv.org/content/10.110… With @danielflevine @SyedARizvi5688 @sachalevy3 @rahuldhodapkar @YaleSEAS @YaleMed #AI #ML #NLP #genomics #CompBio #singlecell