Sebastian Ibarraran retweetledi

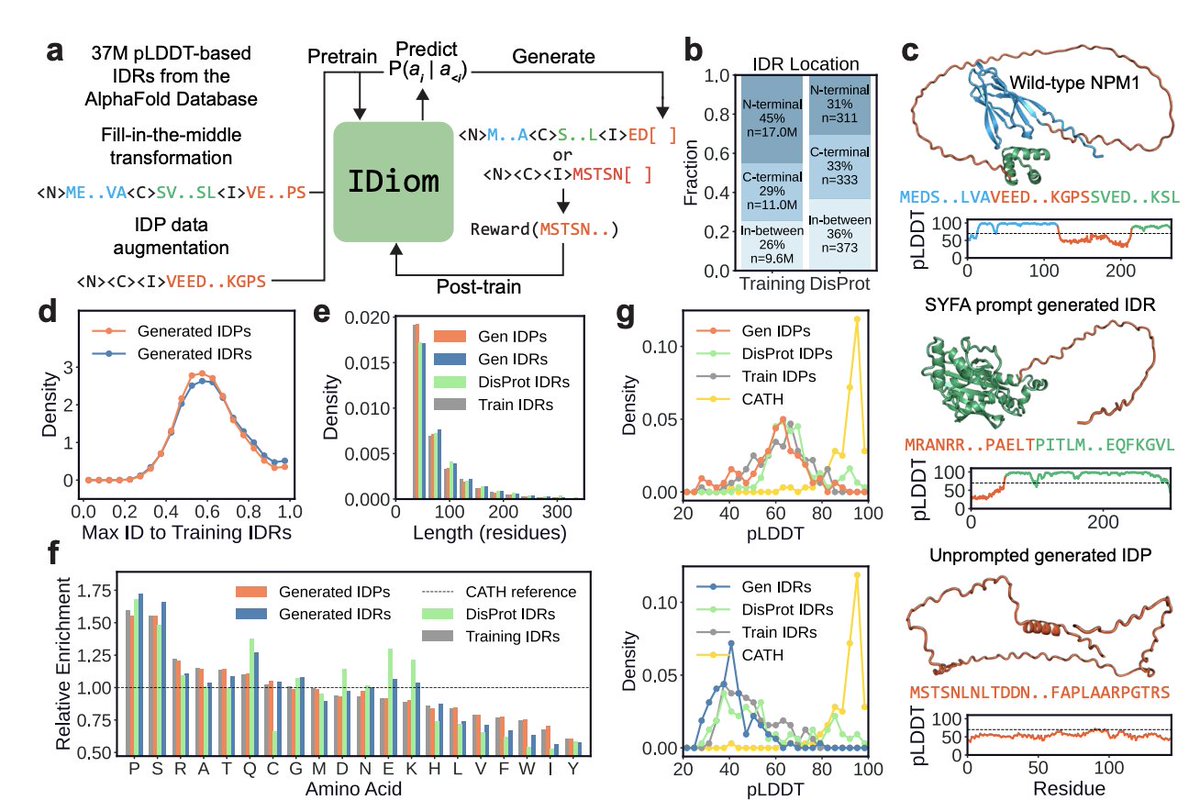
Protein design has been dominated by diffusions due to a "structure-first" perspective. What about intrinsically disordered proteins? We scale language-based design using the modern RL stack and our model IDiom.
Paper: biorxiv.org/content/10.648…
Try it: idiom-designer.vercel.app

English