Grant Rotskoff
53 posts

Grant Rotskoff
@grantrotskoff
Building molecular intelligence @ Stanford






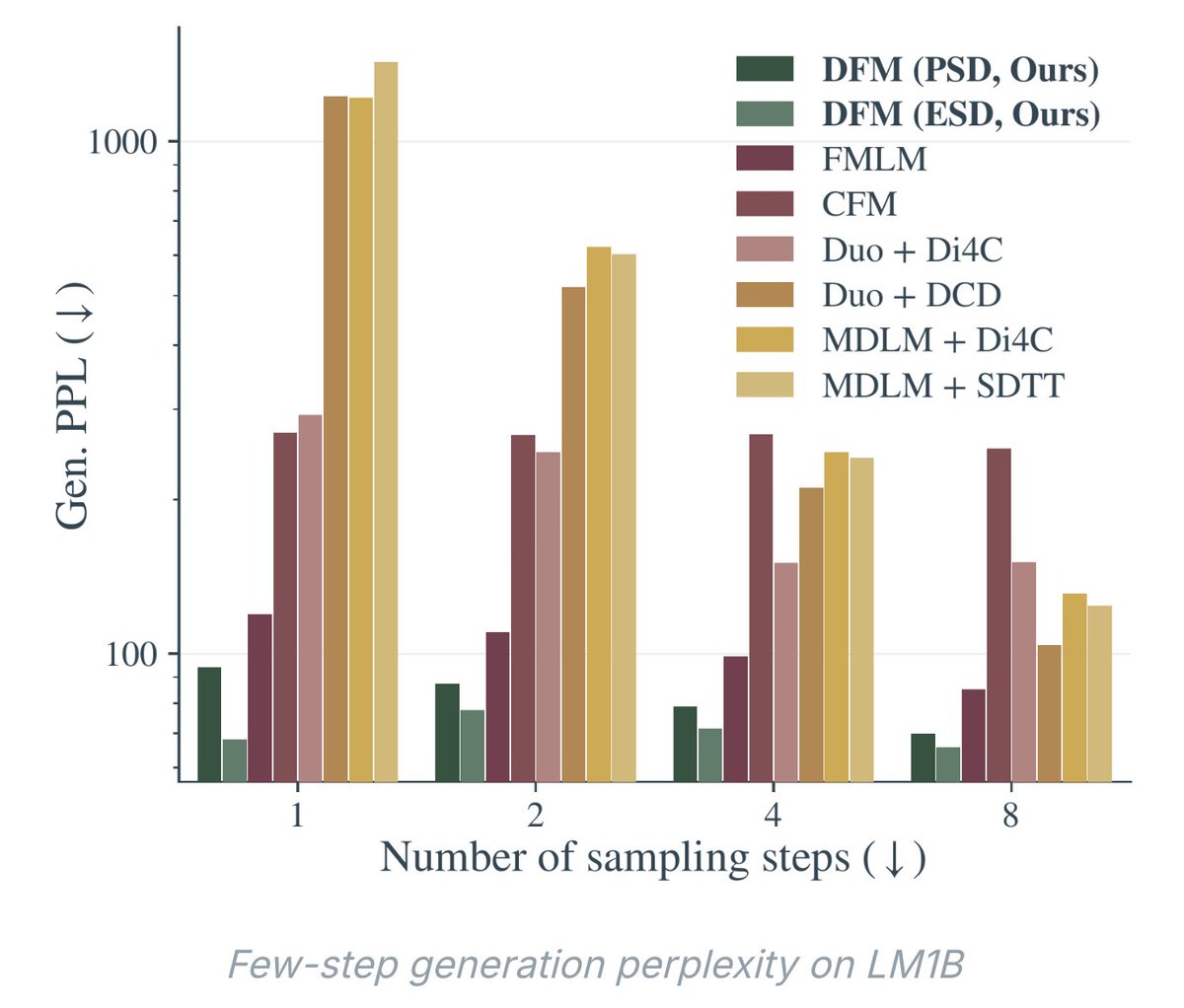

Flow matching is emerging as a unifying framework for generative biology Biology is full of mappings between states: a healthy cell turning diseased, amino acids folding into a functional protein, a ligand docking into its target. Deriving such transformations analytically is intractable—which is where generative AI steps in, and flow matching is quickly becoming its backbone. Morehead and coauthors review how flow matching (FM) is reshaping generative modeling in bioinformatics. Unlike diffusion models, FM doesn't force the source distribution to be Gaussian: it learns a time-dependent vector field that transports samples between any two distributions along straight-line, optimal-transport paths. The payoff: fewer inference steps, simulation-free training, and built-in support for geometric priors like SE(3) equivariance—essential for 3D biomolecules. What's striking is how fast FM has spread across biological scales. For molecules, FoldFlow, FrameFlow, and Multiflow generate protein backbones on SE(3)ᴺ manifolds, SemlaFlow produces valid small molecules up to 100× faster than diffusion, and Dirichlet FM handles discrete DNA/RNA sequences. FlowDock and NeuralPLexer3 predict protein–ligand complexes that match or exceed AlphaFold 3 on key benchmarks, while AlphaFlow and MDGen generate conformational ensembles and MD trajectories. At the cellular scale, CellFlow and Meta FM map unperturbed populations to perturbed states, and CryoFM and FlowSDF extend FM to cryo-EM and microscopy. The deeper point: FM subsumes diffusion models, continuous normalizing flows, and optimal transport as special cases, providing scaffolding for an AI-based virtual cell—simulating molecular, structural, and phenotypic effects of perturbations across scales. Overall, this signals a shift in what's computationally tractable. Instead of narrow, stage-specific models, FM points to unified conditional generators that design sequences, predict complexes, and model perturbation responses in one framework—shortening wet-lab cycles and making closed-loop, active-learning workflows practical. Paper: Morehead and coauthors, Nature Machine Intelligence (2026) — Journal license | doi.org/10.1038/s42256…




Soojung Yang @SoojungYang2 previously created approaches to identify rare protein conformational transitions and, in collaboration with Microsoft Research, to efficiently sample equilibrium ensembles at scale. As a FutureHouse Fellow with Grant Rotskoff @grantrotskoff, she will build machine learning models that unify protein structure, thermodynamics, and kinetics, and deploy agentic AI to search variant space and enable biochemistry-informed protein optimization.



We made Muon run up to 2x faster for free! Introducing Gram Newton-Schulz: a mathematically equivalent but computationally faster Newton-Schulz algorithm for polar decomposition. Gram Newton-Schulz rewrites Newton-Schulz such that instead of iterating on the expensive rectangular X matrix, we iterate on the small, square, symmetric XX^T Gram matrix to reduce FLOPs. This allows us to make more use of fast symmetric GEMM kernels on Hopper and Blackwell, halving the FLOPs of each of those GEMMs. Gram Newton-Schulz is a drop-in replacement of Newton-Schulz for your Muon use case: we see validation perplexity preserved within 0.01, and share our (long!) journey stabilizing this algorithm and ensuring that training quality is preserved above all else. This was a super fun project with @noahamsel, @berlinchen, and @tri_dao that spanned theory, numerical analysis, and ML systems! Blog and codebase linked below 🧵

😊We would like to invite researchers to join our 7th AI for Science workshop @icmlconf as reviewers or ACs. Thank you all for your support to AI for Science. Hope to meet more people in Seoul this summer! AC: docs.google.com/forms/d/e/1FAI… Reviewer: docs.google.com/forms/d/e/1FAI…

Andrew Huberman says Stanford had strict rules preventing faculty from talking about vaccines publicly during Covid. “Stanford had a very strict rule that we weren’t supposed to talk about vaccines publicly.”